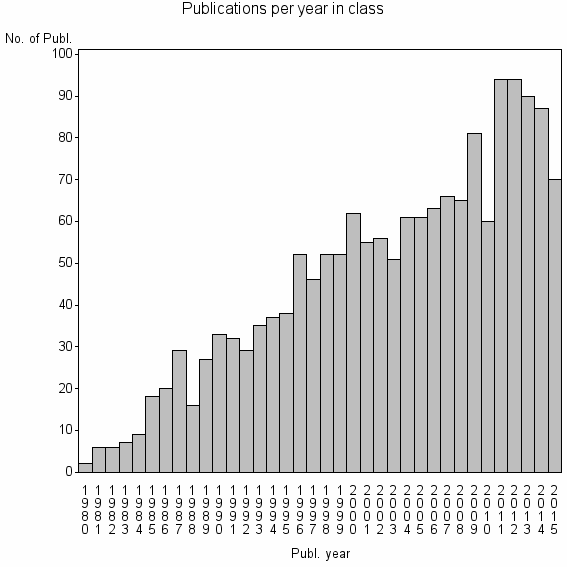
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 4731 | 1662 | 46.0 | 87% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | CENTROMERE | Author keyword | 104 | 33% | 16% | 262 |
| 2 | CENP A | Author keyword | 102 | 74% | 5% | 76 |
| 3 | NEOCENTROMERE | Author keyword | 68 | 79% | 3% | 44 |
| 4 | CENH3 | Author keyword | 58 | 83% | 2% | 33 |
| 5 | HUMAN ARTIFICIAL CHROMOSOME | Author keyword | 39 | 80% | 1% | 24 |
| 6 | CENP C | Author keyword | 35 | 81% | 1% | 21 |
| 7 | KINETOCHORE | Author keyword | 30 | 22% | 7% | 123 |
| 8 | CENP B | Author keyword | 27 | 62% | 2% | 28 |
| 9 | ARTIFICIAL CHROMOSOMES | Author keyword | 16 | 56% | 1% | 19 |
| 10 | HUMAN ARTIFICIAL CHROMOSOME HAC | Author keyword | 14 | 100% | 0% | 7 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | CENTROMERE | 104 | 33% | 16% | 262 | Search CENTROMERE | Search CENTROMERE |
| 2 | CENP A | 102 | 74% | 5% | 76 | Search CENP+A | Search CENP+A |
| 3 | NEOCENTROMERE | 68 | 79% | 3% | 44 | Search NEOCENTROMERE | Search NEOCENTROMERE |
| 4 | CENH3 | 58 | 83% | 2% | 33 | Search CENH3 | Search CENH3 |
| 5 | HUMAN ARTIFICIAL CHROMOSOME | 39 | 80% | 1% | 24 | Search HUMAN+ARTIFICIAL+CHROMOSOME | Search HUMAN+ARTIFICIAL+CHROMOSOME |
| 6 | CENP C | 35 | 81% | 1% | 21 | Search CENP+C | Search CENP+C |
| 7 | KINETOCHORE | 30 | 22% | 7% | 123 | Search KINETOCHORE | Search KINETOCHORE |
| 8 | CENP B | 27 | 62% | 2% | 28 | Search CENP+B | Search CENP+B |
| 9 | ARTIFICIAL CHROMOSOMES | 16 | 56% | 1% | 19 | Search ARTIFICIAL+CHROMOSOMES | Search ARTIFICIAL+CHROMOSOMES |
| 10 | HUMAN ARTIFICIAL CHROMOSOME HAC | 14 | 100% | 0% | 7 | Search HUMAN+ARTIFICIAL+CHROMOSOME+HAC | Search HUMAN+ARTIFICIAL+CHROMOSOME+HAC |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | HISTONE H3 VARIANT | 142 | 83% | 5% | 81 |
| 2 | INNER KINETOCHORE PLATE | 134 | 87% | 4% | 66 |
| 3 | CENP A | 98 | 45% | 10% | 163 |
| 4 | ALPHA SATELLITE DNA | 93 | 37% | 12% | 205 |
| 5 | FISSION YEAST SCM3 | 90 | 87% | 3% | 45 |
| 6 | HISTONE FOLD DOMAIN | 85 | 92% | 2% | 34 |
| 7 | KINETOCHORE | 78 | 32% | 12% | 205 |
| 8 | HUMAN MINICHROMOSOMES | 61 | 95% | 1% | 20 |
| 9 | FUNCTIONAL HUMAN CENTROMERE | 58 | 86% | 2% | 30 |
| 10 | HJURP | 58 | 75% | 3% | 42 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Epigenetic regulation of centromeric chromatin: old dogs, new tricks? | 2008 | 201 | 118 | 75% |
| Neocentromeres: New insights into centromere structure, disease development, and karyotype evolution | 2008 | 139 | 140 | 70% |
| The centromere paradox: Stable inheritance with rapidly evolving DNA | 2001 | 467 | 78 | 63% |
| The Centromere: Chromatin Foundation for the Kinetochore Machinery | 2014 | 9 | 154 | 75% |
| A network of players in H3 histone variant deposition and maintenance at centromeres | 2014 | 8 | 130 | 77% |
| CENP-A: the key player behind centromere identity, propagation, and kinetochore assembly | 2012 | 28 | 89 | 85% |
| Steven Henikoff and Yamini Dalai | 2005 | 93 | 49 | 86% |
| The ABCs of CENPs | 2011 | 60 | 178 | 69% |
| Molecular architecture of vertebrate kinetochores | 2012 | 26 | 63 | 75% |
| Neocentromeres: a place for everything and everything in its place | 2014 | 5 | 77 | 86% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | CHROMOSOME ENGN | 8 | 37% | 1.1% | 18 |
| 2 | REGENERAT MED BIOFUNCT | 8 | 20% | 2.2% | 36 |
| 3 | CHROMATIN STRUCT EPIGENET MECH UNIT | 6 | 100% | 0.2% | 4 |
| 4 | S MOL PHARMACOL | 4 | 75% | 0.2% | 3 |
| 5 | HUMAN GENOME SCI | 4 | 28% | 0.7% | 11 |
| 6 | CELL INNOVAT PROJECT | 3 | 100% | 0.2% | 3 |
| 7 | CHROMOSOME 2 | 3 | 100% | 0.2% | 3 |
| 8 | CHROMOSOME FUNCT REGULAT | 3 | 100% | 0.2% | 3 |
| 9 | MARGINAL BIOSCI | 3 | 100% | 0.2% | 3 |
| 10 | ARIZONA GENOME | 2 | 67% | 0.1% | 2 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000186062 | SATELLITE DNA//HIGHER ORDER REPEATS//SATELLITE DNA EVOLUTION |
| 2 | 0.0000173491 | KINETOCHORE//SPINDLE ASSEMBLY CHECKPOINT//MAD2 |
| 3 | 0.0000132119 | BVES//STAHLMAN CARDIOVASC S//LEK1 |
| 4 | 0.0000120146 | HP1//POSITION EFFECT VARIEGATION//HETEROCHROMATIN |
| 5 | 0.0000090432 | PACHYTENE CHROMOSOME//FLUOROCHROME BANDING//PACHYTENE CHROMOSOMES |
| 6 | 0.0000078987 | HISTONE CHAPERONE//ASF1//CAF 1 |
| 7 | 0.0000078876 | CAT EYE SYNDROME//SUPERNUMERARY MARKER CHROMOSOMES//SMALL SUPERNUMERARY MARKER CHROMOSOME |
| 8 | 0.0000066467 | JUMPING TRANSLOCATION//INTERSTITIAL TELOMERIC SEQUENCES//TELOMERE ASSOCIATION |
| 9 | 0.0000064301 | ROBERTS SYNDROME//TAR SYNDROME//SC PHOCOMELIA |
| 10 | 0.0000055375 | 2 MU M PLASMID//SUC2 PROMOTER//PLASMID STABILITY |