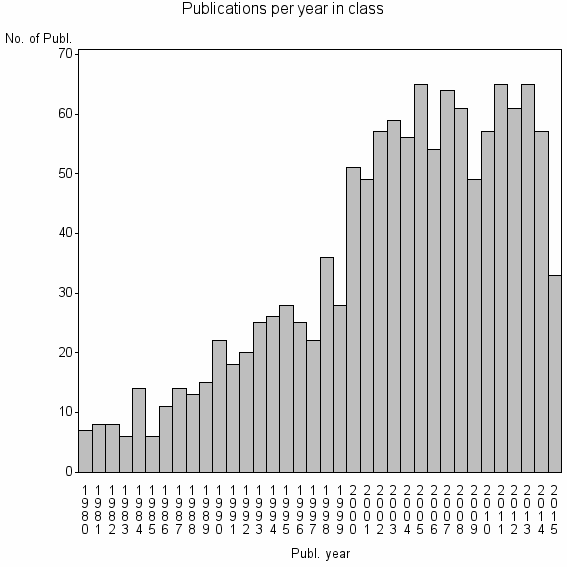
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 7737 | 1255 | 48.8 | 83% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 695 | 12311 | CHROMOSOMA//SYNAPTONEMAL COMPLEX//MEIOSIS |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | HP1 | Author keyword | 34 | 44% | 5% | 58 |
| 2 | POSITION EFFECT VARIEGATION | Author keyword | 30 | 49% | 4% | 44 |
| 3 | HETEROCHROMATIN | Author keyword | 26 | 13% | 15% | 190 |
| 4 | HETEROCHROMATIN PROTEIN 1 | Author keyword | 22 | 54% | 2% | 28 |
| 5 | JIL 1 KINASE | Author keyword | 18 | 89% | 1% | 8 |
| 6 | SUVAR3 9 | Author keyword | 13 | 80% | 1% | 8 |
| 7 | IMMUNOEPIGENET | Address | 10 | 73% | 1% | 8 |
| 8 | HETEROCHROMATIN ASSEMBLY | Author keyword | 9 | 83% | 0% | 5 |
| 9 | HP1A | Author keyword | 9 | 83% | 0% | 5 |
| 10 | CHROMO DOMAIN | Author keyword | 7 | 53% | 1% | 10 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | HP1 | 34 | 44% | 5% | 58 | Search HP1 | Search HP1 |
| 2 | POSITION EFFECT VARIEGATION | 30 | 49% | 4% | 44 | Search POSITION+EFFECT+VARIEGATION | Search POSITION+EFFECT+VARIEGATION |
| 3 | HETEROCHROMATIN | 26 | 13% | 15% | 190 | Search HETEROCHROMATIN | Search HETEROCHROMATIN |
| 4 | HETEROCHROMATIN PROTEIN 1 | 22 | 54% | 2% | 28 | Search HETEROCHROMATIN+PROTEIN+1 | Search HETEROCHROMATIN+PROTEIN+1 |
| 5 | JIL 1 KINASE | 18 | 89% | 1% | 8 | Search JIL+1+KINASE | Search JIL+1+KINASE |
| 6 | SUVAR3 9 | 13 | 80% | 1% | 8 | Search SUVAR3+9 | Search SUVAR3+9 |
| 7 | HETEROCHROMATIN ASSEMBLY | 9 | 83% | 0% | 5 | Search HETEROCHROMATIN+ASSEMBLY | Search HETEROCHROMATIN+ASSEMBLY |
| 8 | HP1A | 9 | 83% | 0% | 5 | Search HP1A | Search HP1A |
| 9 | CHROMO DOMAIN | 7 | 53% | 1% | 10 | Search CHROMO+DOMAIN | Search CHROMO+DOMAIN |
| 10 | HETEROCHROMATIC GENES | 6 | 80% | 0% | 4 | Search HETEROCHROMATIC+GENES | Search HETEROCHROMATIC+GENES |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | POSITION EFFECT VARIEGATION | 130 | 32% | 27% | 339 |
| 2 | CHROME SHADOW DOMAIN | 83 | 90% | 3% | 36 |
| 3 | NONHISTONE CHROMOSOMAL PROTEIN | 81 | 68% | 6% | 71 |
| 4 | SHADOW DOMAIN | 58 | 88% | 2% | 28 |
| 5 | HP1 PROTEINS | 55 | 46% | 7% | 90 |
| 6 | HP1 | 47 | 37% | 8% | 100 |
| 7 | CHROMODOMAIN PROTEIN | 42 | 53% | 4% | 55 |
| 8 | HETEROCHROMATIN ASSOCIATED PROTEIN | 35 | 57% | 3% | 42 |
| 9 | INTERCALARY HETEROCHROMATIN | 34 | 65% | 3% | 33 |
| 10 | RITS | 33 | 66% | 2% | 31 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Mechanisms of functional promiscuity by HP1 proteins | 2014 | 13 | 86 | 77% |
| Heterochromatin revisited | 2007 | 524 | 140 | 52% |
| RNAi-dependent formation of heterochromatin and its diverse functions | 2010 | 118 | 79 | 63% |
| HP1a: a structural chromosomal protein regulating transcription | 2014 | 10 | 73 | 62% |
| Small RNAs in transcriptional gene silencing and genome defence | 2009 | 314 | 99 | 30% |
| Transcription and RNA interference in the formation of heterochromatin | 2007 | 229 | 79 | 68% |
| The changing faces of HP1: From heterochromatin formation and gene silencing to euchromatic gene expression | 2011 | 55 | 106 | 64% |
| HP1: a functionally multifaceted protein | 2008 | 69 | 40 | 90% |
| Different means, same end - heterochromatin formation by RNAi and RNAi-independent RNA processing factors in fission yeast | 2012 | 26 | 70 | 67% |
| Heterochromatin and epigenetic control of gene expression | 2003 | 564 | 60 | 37% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | IMMUNOEPIGENET | 10 | 73% | 0.6% | 8 |
| 2 | CHROMATIN FUNCT | 6 | 80% | 0.3% | 4 |
| 3 | NUCL REPROGRAMMING | 4 | 39% | 0.7% | 9 |
| 4 | DIPARTIMENTO GENET BIOL MOL CHARLES DARWIN | 4 | 46% | 0.5% | 6 |
| 5 | CHROMATIN DYNAM | 3 | 22% | 1.0% | 13 |
| 6 | CHROMATIN BIOCHEM | 2 | 22% | 0.7% | 9 |
| 7 | STEM CELL CHROMATIN GRP | 2 | 50% | 0.2% | 3 |
| 8 | PROTE SUPPORT UNIT | 2 | 33% | 0.3% | 4 |
| 9 | BASIC SCIGENE REGULAT CHROMOSOME BIOL | 1 | 100% | 0.2% | 2 |
| 10 | GENOM FUNZ PROTE SISTEMI MODELLO | 1 | 50% | 0.2% | 2 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000243396 | DNA PUFF//DNA PUFFS//RHYNCHOSCIARA |
| 2 | 0.0000177234 | POKEY//LAMPBRUSH LOOPS//FERTILITY GENES |
| 3 | 0.0000148618 | IMMORTAL DNA//ADULT STEM CELL TECHNOL//IMMORTAL STRAND HYPOTHESIS |
| 4 | 0.0000146007 | SIR COMPLEX//SIR3//SIR PROTEINS |
| 5 | 0.0000126743 | SMALL ACTIVATING RNA//SARNA//RNA ACTIVATION |
| 6 | 0.0000120146 | CENTROMERE//CENP A//NEOCENTROMERE |
| 7 | 0.0000119723 | PIRNA//PIWI//PIRNAS |
| 8 | 0.0000114601 | HISTONE DEMETHYLASE//HISTONE DEMETHYLATION//LSD1 |
| 9 | 0.0000113115 | CTCF//CONTROL GENET PROC//ENHANCER BLOCKING |
| 10 | 0.0000100882 | POLYTENE CHROMOSOMES//CHIRONOMUS//BALBIANI RING GENES |