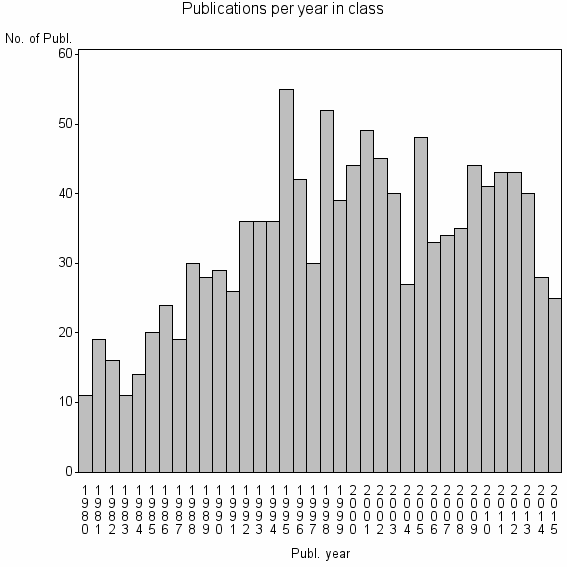
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 8358 | 1192 | 35.6 | 62% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 587 | 13504 | NEW ZEALAND FLORA//NEW ZEALAND JOURNAL OF BOTANY//GENOME SIZE |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | PACHYTENE CHROMOSOME | Author keyword | 18 | 89% | 1% | 8 |
| 2 | FLUOROCHROME BANDING | Author keyword | 15 | 50% | 2% | 22 |
| 3 | PACHYTENE CHROMOSOMES | Author keyword | 15 | 88% | 1% | 7 |
| 4 | 5S AND 45S RDNA | Author keyword | 13 | 80% | 1% | 8 |
| 5 | 5S RDNA | Author keyword | 12 | 24% | 4% | 45 |
| 6 | RDNA SITES | Author keyword | 9 | 55% | 1% | 11 |
| 7 | 45S RDNA | Author keyword | 7 | 34% | 2% | 18 |
| 8 | PLANT CYTOGENET | Address | 7 | 53% | 1% | 9 |
| 9 | 5S RRNA GENES | Author keyword | 7 | 67% | 1% | 6 |
| 10 | RDNAS | Author keyword | 7 | 67% | 1% | 6 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | PACHYTENE CHROMOSOME | 18 | 89% | 1% | 8 | Search PACHYTENE+CHROMOSOME | Search PACHYTENE+CHROMOSOME |
| 2 | FLUOROCHROME BANDING | 15 | 50% | 2% | 22 | Search FLUOROCHROME+BANDING | Search FLUOROCHROME+BANDING |
| 3 | PACHYTENE CHROMOSOMES | 15 | 88% | 1% | 7 | Search PACHYTENE+CHROMOSOMES | Search PACHYTENE+CHROMOSOMES |
| 4 | 5S AND 45S RDNA | 13 | 80% | 1% | 8 | Search 5S+AND+45S+RDNA | Search 5S+AND+45S+RDNA |
| 5 | 5S RDNA | 12 | 24% | 4% | 45 | Search 5S+RDNA | Search 5S+RDNA |
| 6 | RDNA SITES | 9 | 55% | 1% | 11 | Search RDNA+SITES | Search RDNA+SITES |
| 7 | 45S RDNA | 7 | 34% | 2% | 18 | Search 45S+RDNA | Search 45S+RDNA |
| 8 | 5S RRNA GENES | 7 | 67% | 1% | 6 | Search 5S+RRNA+GENES | Search 5S+RRNA+GENES |
| 9 | RDNAS | 7 | 67% | 1% | 6 | Search RDNAS | Search RDNAS |
| 10 | QUANTITATIVE CHROMOSOME MAP | 6 | 80% | 0% | 4 | Search QUANTITATIVE+CHROMOSOME+MAP | Search QUANTITATIVE+CHROMOSOME+MAP |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | PACHYTENE CHROMOSOMES | 41 | 69% | 3% | 35 |
| 2 | MEIOTIC PACHYTENE | 33 | 100% | 1% | 13 |
| 3 | RDNA LOCI | 32 | 51% | 4% | 45 |
| 4 | EXTENDED DNA FIBERS | 25 | 66% | 2% | 23 |
| 5 | 5S | 22 | 31% | 5% | 59 |
| 6 | PLANT CHROMOSOMES | 19 | 32% | 4% | 49 |
| 7 | 18S 25S RDNA | 12 | 44% | 2% | 21 |
| 8 | ADENOLINUM | 12 | 86% | 1% | 6 |
| 9 | 45S RDNA | 11 | 65% | 1% | 11 |
| 10 | GENUS PINUS | 10 | 52% | 1% | 14 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Current status and the future of fluorescence in situ hybridization (FISH) in plant genome research | 2006 | 101 | 121 | 46% |
| Organisation of the plant genome in chromosomes | 2011 | 53 | 130 | 13% |
| High resolution FISH in plants - techniques and applications | 1999 | 92 | 33 | 45% |
| Different chromatin fractions of tomato (Solanum lycopersicum L.) and related species | 2009 | 8 | 33 | 58% |
| Role of Fluorescence in situ Hybridization in Sequencing the Tomato Genome | 2009 | 16 | 38 | 37% |
| Organization and evolution of highly repeated satellite DNA sequences in plant chromosomes | 2005 | 51 | 116 | 30% |
| Preparation of samples for comparative studies of plant chromosomes using in situ hybridization methods | 2005 | 18 | 28 | 50% |
| Advances in plant chromosome identification and cytogenetic techniques | 2005 | 25 | 51 | 39% |
| NONISOTOPIC IN-SITU HYBRIDIZATION AND PLANT GENOME MAPPING - THE FIRST 10 YEARS | 1994 | 184 | 77 | 21% |
| Plant highly repeated satellite DNA: molecular evolution, distribution and use for identification of hybrids | 2007 | 18 | 140 | 33% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | PLANT CYTOGENET | 7 | 53% | 0.8% | 9 |
| 2 | PLANT ANAT CYTOL | 5 | 17% | 2.3% | 28 |
| 3 | RICE GENET ENGN | 4 | 50% | 0.5% | 6 |
| 4 | STATE COTTON BIOL CHINA | 3 | 100% | 0.3% | 3 |
| 5 | CITOGENET VEGETAL | 3 | 28% | 0.8% | 9 |
| 6 | BIOSSISTEMAT | 2 | 67% | 0.2% | 2 |
| 7 | PROTECT ENDEM OURCES | 2 | 33% | 0.3% | 4 |
| 8 | KARYOBIOL GRP | 2 | 26% | 0.4% | 5 |
| 9 | ALL RUSSIAN WILLIAMS FODDER | 1 | 100% | 0.2% | 2 |
| 10 | FO T TREE MOL CYTOGENET | 1 | 100% | 0.2% | 2 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000296993 | RDNA INTERGENIC SPACER//NON CONCERTED EVOLUTION//INTERGENIC SPACER |
| 2 | 0.0000172833 | BRACHYPODIUM DISTACHYON//BRACHYPODIUM//BRACHYPODIUM HYBRIDUM |
| 3 | 0.0000146636 | GAGEA//INCA LILY//BOMAREA |
| 4 | 0.0000144784 | TRITICEAE//ELYMUS//LEYMUS |
| 5 | 0.0000132390 | ARTEMISIINAE//ANTHEMIDEAE//TRIPLEUROSPERMUM |
| 6 | 0.0000131384 | NARCISSUS SPP//MOSQUITO CELL LINES//PLANT CELL NUCLEUS |
| 7 | 0.0000120681 | GENOME SIZE//C VALUE//NUCLEAR DNA CONTENT |
| 8 | 0.0000105005 | RICE GENOME PROGRAM//CROP DESIGN//GENOME IL |
| 9 | 0.0000098418 | RETROTRANSPOSONS//TY1 COPIA//TRANSPOSABLE ELEMENTS |
| 10 | 0.0000090902 | POLYPLOIDY//AUTOPOLYPLOIDY//ALLOPOLYPLOIDY |