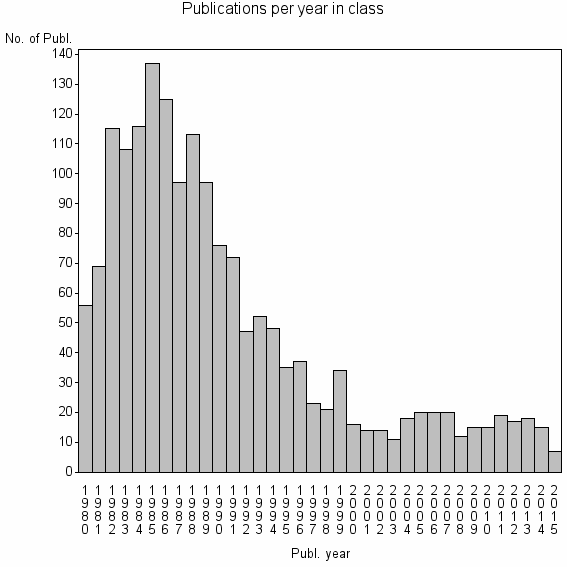
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 4333 | 1729 | 34.4 | 60% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 1635 | 6457 | Z DNA//JOURNAL OF BIOMOLECULAR STRUCTURE & DYNAMICS//B Z TRANSITION |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | Z DNA | Author keyword | 58 | 54% | 4% | 75 |
| 2 | B Z TRANSITION | Author keyword | 38 | 86% | 1% | 19 |
| 3 | Z DNA BINDING PROTEIN | Author keyword | 26 | 100% | 1% | 11 |
| 4 | Z ALPHA | Author keyword | 14 | 100% | 0% | 7 |
| 5 | PKZ | Author keyword | 9 | 83% | 0% | 5 |
| 6 | LEFT HANDED DNA | Author keyword | 5 | 63% | 0% | 5 |
| 7 | B Z JUNCTION | Author keyword | 4 | 67% | 0% | 4 |
| 8 | LEFT HANDED Z DNA | Author keyword | 4 | 67% | 0% | 4 |
| 9 | BIOTECH RD | Address | 4 | 75% | 0% | 3 |
| 10 | Z ALPHA DOMAIN | Author keyword | 3 | 100% | 0% | 3 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | Z DNA | 58 | 54% | 4% | 75 | Search Z+DNA | Search Z+DNA |
| 2 | B Z TRANSITION | 38 | 86% | 1% | 19 | Search B+Z+TRANSITION | Search B+Z+TRANSITION |
| 3 | Z DNA BINDING PROTEIN | 26 | 100% | 1% | 11 | Search Z+DNA+BINDING+PROTEIN | Search Z+DNA+BINDING+PROTEIN |
| 4 | Z ALPHA | 14 | 100% | 0% | 7 | Search Z+ALPHA | Search Z+ALPHA |
| 5 | PKZ | 9 | 83% | 0% | 5 | Search PKZ | Search PKZ |
| 6 | LEFT HANDED DNA | 5 | 63% | 0% | 5 | Search LEFT+HANDED+DNA | Search LEFT+HANDED+DNA |
| 7 | B Z JUNCTION | 4 | 67% | 0% | 4 | Search B+Z+JUNCTION | Search B+Z+JUNCTION |
| 8 | LEFT HANDED Z DNA | 4 | 67% | 0% | 4 | Search LEFT+HANDED+Z+DNA | Search LEFT+HANDED+Z+DNA |
| 9 | Z ALPHA DOMAIN | 3 | 100% | 0% | 3 | Search Z+ALPHA+DOMAIN | Search Z+ALPHA+DOMAIN |
| 10 | ALLOSTERIC DNA | 2 | 67% | 0% | 2 | Search ALLOSTERIC+DNA | Search ALLOSTERIC+DNA |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | HANDED Z DNA | 139 | 55% | 10% | 175 |
| 2 | Z ALPHA DOMAIN | 41 | 60% | 3% | 44 |
| 3 | HUMAN EDITING ENZYME | 40 | 70% | 2% | 33 |
| 4 | Z FORM | 32 | 69% | 2% | 27 |
| 5 | B Z TRANSITION | 32 | 48% | 3% | 49 |
| 6 | POLYDG DC | 30 | 55% | 2% | 37 |
| 7 | MAMMALIAN CELL NUCLEI | 27 | 78% | 1% | 18 |
| 8 | Z TRANSITION | 25 | 49% | 2% | 37 |
| 9 | POLYDA TPOLYDA T | 24 | 91% | 1% | 10 |
| 10 | SYNTHETIC DNA | 24 | 49% | 2% | 35 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Z-DNA: the long road to biological function | 2003 | 221 | 59 | 75% |
| INFRARED-SPECTROSCOPY OF DNA | 1992 | 239 | 30 | 73% |
| Z-DNA, an active element in the genome | 2007 | 44 | 156 | 52% |
| THE CHEMISTRY AND BIOLOGY OF LEFT-HANDED Z-DNA | 1984 | 925 | 145 | 81% |
| THE TRANSITION BETWEEN B-DNA AND Z-DNA | 1987 | 176 | 222 | 78% |
| Advancements in Z-DNA: Development of Inducers and Stabilizers for B to Z Transition | 2012 | 4 | 118 | 79% |
| Molecular mechanisms for the B-Z transition in the example of poly[d(G-C)center dot d(G-C)] polymers. A critical review | 2006 | 34 | 134 | 70% |
| Z-DNA Binding Proteins as Targets for Structure-Based Virtual Screening | 2010 | 13 | 74 | 50% |
| LOCAL SUPERCOIL-STABILIZED DNA STRUCTURES | 1991 | 181 | 493 | 33% |
| INTERPRETATION OF DNA VIBRATIONAL-SPECTRA BY NORMAL COORDINATE ANALYSIS | 1990 | 42 | 23 | 96% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | BIOTECH RD | 4 | 75% | 0.2% | 3 |
| 2 | ADULT DI IYODA KU | 1 | 50% | 0.1% | 1 |
| 3 | CHANGCHUN PL CHEMSTATE RARE EARTH | 1 | 50% | 0.1% | 1 |
| 4 | FREDERICK CANC DEV ADV BIOSCI S | 1 | 50% | 0.1% | 1 |
| 5 | GENOM SCI S GRP 3 | 1 | 50% | 0.1% | 1 |
| 6 | LPCB CNRS UA 198 | 1 | 50% | 0.1% | 1 |
| 7 | SCI BRANCH CHEM | 0 | 33% | 0.1% | 1 |
| 8 | BIOP | 0 | 25% | 0.1% | 1 |
| 9 | EDIFICI M | 0 | 25% | 0.1% | 1 |
| 10 | MED DANA FARBER CANC MED | 0 | 17% | 0.1% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000179011 | DNA CURVATURE//A TRACT//DNA FLEXIBILITY |
| 2 | 0.0000157158 | SHEARED G CENTER DOT A PAIR//DGGTATACC2//SHEARED A CENTER DOT C PAIR |
| 3 | 0.0000108317 | 5 PHOSPHODIESTERASE//THREE STRANDED POLYNUCLEOTIDES//CU ATP COMPLEX |
| 4 | 0.0000102111 | L DNA//HETEROCHIRAL DNA//UMR CNRS UMII 5625 |
| 5 | 0.0000094450 | TRIPLEX DNA//DNA TRIPLEX//TRIPLEX |
| 6 | 0.0000085833 | HYDROXYMETHYLPHOSPHONATE DNA//PHOSPHATEMETHYLATED DNA//3 3 DIPHENYLINDAN 1 2 DIONE |
| 7 | 0.0000082647 | DNA DYNAMICS//PEYRARD BISHOP MODEL//DNA DENATURATION |
| 8 | 0.0000071023 | EMANUEL SYNDROME//PARTIAL TRISOMY 11Q//DUPLICATION 11Q |
| 9 | 0.0000067899 | REVERSING PULSE ELECTRIC BIREFRINGENCE//LOW FREQUENCY ANOMALIES IN COLLOIDAL ELECTRO OPTICS//ELECTRIC DICHROISM |
| 10 | 0.0000064670 | WARWICK ANALYT SCI//GELRED//DNA TOPOISOMERS |