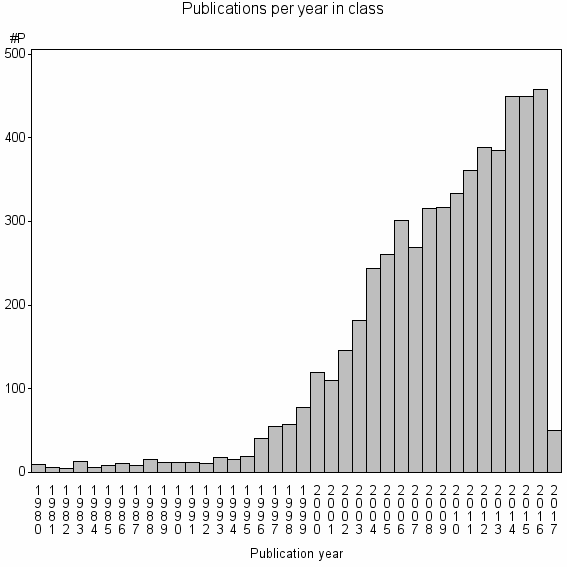
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 1995 | 5542 | 39.5 | 83% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 5 | 4 | CHEMISTRY, ORGANIC//CHEMISTRY, INORGANIC & NUCLEAR//CHEMISTRY, MULTIDISCIPLINARY | 1745167 |
| 183 | 3 | APTAMER//G QUADRUPLEX//RIBOZYME | 56307 |
| 1995 | 2 | DNA ORIGAMI//DNA NANOTECHNOLOGY//DNA NANOSTRUCTURES | 5542 |
| 2108 | 1 | DNA ORIGAMI//DNA NANOTECHNOLOGY//DNA NANOSTRUCTURES | 2354 |
| 4609 | 1 | SCI IND SANKEN//DNA ELECTRON TRANSFER//DNA MOLECULE | 1757 |
| 18071 | 1 | CHEM NANOSCI S//MOLECULAR COMBING//DNA TEMPLATE | 568 |
| 22670 | 1 | GIANT DIAMAGNETISM//SINGLE PARTICLE MAGNET//FIS MAT CFM MPC CSIC UPV EHU | 372 |
| 23710 | 1 | DNA CTMA COMPLEX//BIOMAT CHEMKITA KU//DNA LIPID COMPLEX | 337 |
| 35903 | 1 | POLYPHENAZASILINE//EFFECT OF THE SUBSTITUENT//FUNCTIONAL ADDITIVE | 90 |
| 37169 | 1 | FRONTIER SCI INNOVAT//NANOARCHITECTURE GRP//MOLECULAR BRIDGING | 64 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | DNA ORIGAMI | authKW | 697963 | 3% | 79% | 160 |
| 2 | DNA NANOTECHNOLOGY | authKW | 612538 | 3% | 72% | 155 |
| 3 | DNA NANOSTRUCTURES | authKW | 275995 | 1% | 73% | 69 |
| 4 | DNA | authKW | 163121 | 12% | 4% | 674 |
| 5 | DNA TILES | authKW | 143936 | 1% | 93% | 28 |
| 6 | TILE ASSEMBLY MODEL | authKW | 129517 | 0% | 87% | 27 |
| 7 | DNA STRUCTURES | authKW | 123459 | 2% | 25% | 89 |
| 8 | DNA SELF ASSEMBLY | authKW | 121188 | 1% | 49% | 45 |
| 9 | MOLECULAR PROGRAMMING | authKW | 114709 | 0% | 77% | 27 |
| 10 | DNA STRAND DISPLACEMENT | authKW | 88279 | 0% | 70% | 23 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Chemistry, Multidisciplinary | 19106 | 39% | 0% | 2141 |
| 2 | Nanoscience & Nanotechnology | 16180 | 19% | 0% | 1043 |
| 3 | Chemistry, Physical | 5568 | 23% | 0% | 1286 |
| 4 | Materials Science, Multidisciplinary | 4959 | 23% | 0% | 1292 |
| 5 | Physics, Applied | 2883 | 17% | 0% | 952 |
| 6 | Physics, Condensed Matter | 2652 | 13% | 0% | 740 |
| 7 | Physics, Atomic, Molecular & Chemical | 762 | 6% | 0% | 328 |
| 8 | Physics, Multidisciplinary | 260 | 5% | 0% | 274 |
| 9 | Computer Science, Theory & Methods | 226 | 2% | 0% | 136 |
| 10 | Polymer Science | 202 | 4% | 0% | 196 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | SUNGKYUNKWAN ADV NANOTECHNOL SAINT | 58908 | 1% | 31% | 34 |
| 2 | DNA NANOTECHNOL | 56383 | 0% | 39% | 26 |
| 3 | BIODESIGN | 40122 | 2% | 6% | 118 |
| 4 | SCI IND SANKEN | 40020 | 1% | 19% | 39 |
| 5 | ZNN WSI | 34473 | 0% | 52% | 12 |
| 6 | MOL DESIGN BIOMIMICRY | 33047 | 0% | 100% | 6 |
| 7 | DNA NANOTECHNOL CDNA | 29956 | 0% | 26% | 21 |
| 8 | LEHRSTUHL BIOELEKT | 29735 | 0% | 60% | 9 |
| 9 | FRONTIER SCI INNOVAT | 28179 | 0% | 32% | 16 |
| 10 | BIOMAT CHEMKITA KU | 27539 | 0% | 100% | 5 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | NATURAL COMPUTING | 21964 | 1% | 11% | 38 |
| 2 | NANO LETTERS | 13122 | 3% | 1% | 172 |
| 3 | JOURNAL OF THE AMERICAN CHEMICAL SOCIETY | 10670 | 8% | 0% | 433 |
| 4 | NATURE NANOTECHNOLOGY | 10456 | 1% | 4% | 50 |
| 5 | SMALL | 9681 | 2% | 2% | 95 |
| 6 | ACS NANO | 7948 | 2% | 1% | 117 |
| 7 | ANGEWANDTE CHEMIE-INTERNATIONAL EDITION | 6877 | 4% | 1% | 225 |
| 8 | JOURNAL OF PHYSICAL CHEMISTRY B | 3788 | 3% | 0% | 168 |
| 9 | NATURE CHEMISTRY | 3651 | 0% | 3% | 26 |
| 10 | CHEMICAL COMMUNICATIONS | 3299 | 3% | 0% | 166 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | DNA ORIGAMI | 697963 | 3% | 79% | 160 | Search DNA+ORIGAMI | Search DNA+ORIGAMI |
| 2 | DNA NANOTECHNOLOGY | 612538 | 3% | 72% | 155 | Search DNA+NANOTECHNOLOGY | Search DNA+NANOTECHNOLOGY |
| 3 | DNA NANOSTRUCTURES | 275995 | 1% | 73% | 69 | Search DNA+NANOSTRUCTURES | Search DNA+NANOSTRUCTURES |
| 4 | DNA | 163121 | 12% | 4% | 674 | Search DNA | Search DNA |
| 5 | DNA TILES | 143936 | 1% | 93% | 28 | Search DNA+TILES | Search DNA+TILES |
| 6 | TILE ASSEMBLY MODEL | 129517 | 0% | 87% | 27 | Search TILE+ASSEMBLY+MODEL | Search TILE+ASSEMBLY+MODEL |
| 7 | DNA STRUCTURES | 123459 | 2% | 25% | 89 | Search DNA+STRUCTURES | Search DNA+STRUCTURES |
| 8 | DNA SELF ASSEMBLY | 121188 | 1% | 49% | 45 | Search DNA+SELF+ASSEMBLY | Search DNA+SELF+ASSEMBLY |
| 9 | MOLECULAR PROGRAMMING | 114709 | 0% | 77% | 27 | Search MOLECULAR+PROGRAMMING | Search MOLECULAR+PROGRAMMING |
| 10 | DNA STRAND DISPLACEMENT | 88279 | 0% | 70% | 23 | Search DNA+STRAND+DISPLACEMENT | Search DNA+STRAND+DISPLACEMENT |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | GENEREUX, JC , BARTON, JK , (2010) MECHANISMS FOR DNA CHARGE TRANSPORT.CHEMICAL REVIEWS. VOL. 110. ISSUE 3. P. 1642 -1662 | 187 | 75% | 315 |
| 2 | SUN, LF , YU, L , SHEN, WQ , (2014) DNA NANOTECHNOLOGY AND ITS APPLICATIONS IN BIOMEDICAL RESEARCH.JOURNAL OF BIOMEDICAL NANOTECHNOLOGY. VOL. 10. ISSUE 9. P. 2350 -2370 | 198 | 93% | 5 |
| 3 | PINHEIRO, AV , HAN, DR , SHIH, WM , YAN, H , (2011) CHALLENGES AND OPPORTUNITIES FOR STRUCTURAL DNA NANOTECHNOLOGY.NATURE NANOTECHNOLOGY. VOL. 6. ISSUE 12. P. 763 -772 | 114 | 84% | 396 |
| 4 | ZHANG, F , NANGREAVE, J , LIU, Y , YAN, H , (2014) STRUCTURAL DNA NANOTECHNOLOGY: STATE OF THE ART AND FUTURE PERSPECTIVE.JOURNAL OF THE AMERICAN CHEMICAL SOCIETY. VOL. 136. ISSUE 32. P. 11198 -11211 | 117 | 80% | 99 |
| 5 | TINTORE, M , ERITJA, R , FABREGA, C , (2014) DNA NANOARCHITECTURES: STEPS TOWARDS BIOLOGICAL APPLICATIONS.CHEMBIOCHEM. VOL. 15. ISSUE 10. P. 1374 -1390 | 167 | 86% | 12 |
| 6 | WAGENKNECHT, HA , (2006) ELECTRON TRANSFER PROCESSES IN DNA: MECHANISMS, BIOLOGICAL RELEVANCE AND APPLICATIONS IN DNA ANALYTICS.NATURAL PRODUCT REPORTS. VOL. 23. ISSUE 6. P. 973-1006 | 254 | 64% | 68 |
| 7 | GATES, EP , DEARDEN, AM , WOOLLEY, AT , (2014) DNA-TEMPLATED LITHOGRAPHY AND NANOFABRICATION FOR THE FABRICATION OF NANOSCALE ELECTRONIC CIRCUITRY.CRITICAL REVIEWS IN ANALYTICAL CHEMISTRY. VOL. 44. ISSUE 4. P. 354 -370 | 145 | 94% | 4 |
| 8 | ZHANG, Z , FU, YM , LI, BJ , FENG, GY , LI, C , FAN, CH , HE, L , (2011) SELF-ASSEMBLY-BASED STRUCTURAL DNA NANOTECHNOLOGY.CURRENT ORGANIC CHEMISTRY. VOL. 15. ISSUE 4. P. 534 -547 | 144 | 96% | 2 |
| 9 | LI, HY , LABEAN, TH , LEONG, KW , (2011) NUCLEIC ACID-BASED NANOENGINEERING: NOVEL STRUCTURES FOR BIOMEDICAL APPLICATIONS.INTERFACE FOCUS. VOL. 1. ISSUE 5. P. 702 -724 | 166 | 72% | 17 |
| 10 | WANG, F , LU, CH , WILLNER, I , (2014) FROM CASCADED CATALYTIC NUCLEIC ACIDS TO ENZYME-DNA NANOSTRUCTURES: CONTROLLING REACTIVITY, SENSING, LOGIC OPERATIONS, AND ASSEMBLY OF COMPLEX STRUCTURES.CHEMICAL REVIEWS. VOL. 114. ISSUE 5. P. 2881 -2941 | 218 | 25% | 153 |
Classes with closest relation at Level 2 |