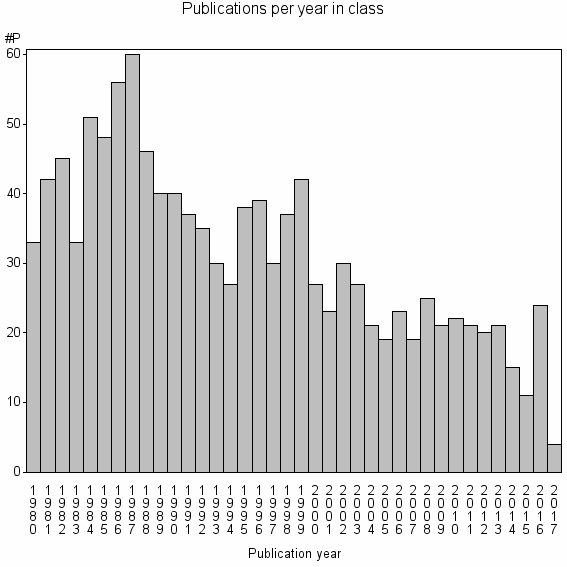
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 9156 | 1182 | 38.8 | 61% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 11 | 4 | NEUROSCIENCES//CLINICAL NEUROLOGY//NEUROL | 1112395 |
| 124 | 3 | PARKINSONS DISEASE//MOVEMENT DISORDERS//DEEP BRAIN STIMULATION | 67777 |
| 2411 | 2 | PHENYLKETONURIA//PHENYLALANINE HYDROXYLASE//PKU | 4184 |
| 9156 | 1 | PHENYLALANINE HYDROXYLASE//AROMATIC AMINO ACID HYDROXYLASE//TRYPTOPHAN HYDROXYLASE | 1182 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | PHENYLALANINE HYDROXYLASE | authKW | 435330 | 5% | 26% | 64 |
| 2 | AROMATIC AMINO ACID HYDROXYLASE | authKW | 281276 | 1% | 78% | 14 |
| 3 | TRYPTOPHAN HYDROXYLASE | authKW | 281208 | 6% | 15% | 72 |
| 4 | TYROSINE HYDROXYLASE | authKW | 238596 | 15% | 5% | 176 |
| 5 | INSECT BRAIN NEUROBIOL | address | 82661 | 0% | 80% | 4 |
| 6 | SEROTONIN BIOSYNTHESIS | authKW | 58704 | 0% | 45% | 5 |
| 7 | DOPAMINE INHIBITION | authKW | 58121 | 0% | 75% | 3 |
| 8 | NEUROCATIN | authKW | 58121 | 0% | 75% | 3 |
| 9 | TYROSINE HYDROXYLASE GENE | authKW | 55024 | 1% | 30% | 7 |
| 10 | AUTOREGULATORY SEQUENCE | authKW | 51664 | 0% | 100% | 2 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biochemistry & Molecular Biology | 7972 | 66% | 0% | 786 |
| 2 | Neurosciences | 2882 | 30% | 0% | 354 |
| 3 | Biophysics | 999 | 11% | 0% | 135 |
| 4 | Biochemical Research Methods | 139 | 5% | 0% | 54 |
| 5 | Pharmacology & Pharmacy | 93 | 8% | 0% | 93 |
| 6 | Cell Biology | 35 | 5% | 0% | 59 |
| 7 | Biology | 16 | 2% | 0% | 21 |
| 8 | Genetics & Heredity | 9 | 3% | 0% | 32 |
| 9 | Medicine, Research & Experimental | 6 | 2% | 0% | 25 |
| 10 | Chemistry, Analytical | 3 | 2% | 0% | 28 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | INSECT BRAIN NEUROBIOL | 82661 | 0% | 80% | 4 |
| 2 | LSUHSC BIOCHEM | 51664 | 0% | 100% | 2 |
| 3 | BIOL MOIL SEVERO OCHOA | 25832 | 0% | 100% | 1 |
| 4 | BIOMOL NEUROSCI | 25832 | 0% | 100% | 1 |
| 5 | CAVANILLES BIO ERAITAT BIOL EVOLUT | 25832 | 0% | 100% | 1 |
| 6 | DEPARMENT AQUACULTURE | 25832 | 0% | 100% | 1 |
| 7 | ECOL MICROBIENNEUMR5557 | 25832 | 0% | 100% | 1 |
| 8 | EQUIPE INTERACT CELLULAI MOL REPROD | 25832 | 0% | 100% | 1 |
| 9 | GENET MOL NEUROTRANSMISS PROC NEURODIGENERA | 25832 | 0% | 100% | 1 |
| 10 | GENET MOL NEUROTRANSMISS PROC NEUROGENERAT | 25832 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | JOURNAL OF NEUROCHEMISTRY | 22583 | 11% | 1% | 135 |
| 2 | PTERIDINES | 7715 | 1% | 3% | 11 |
| 3 | NEUROCHEMISTRY INTERNATIONAL | 5565 | 3% | 1% | 34 |
| 4 | BIOCHEMISTRY | 2854 | 7% | 0% | 78 |
| 5 | BIOGENIC AMINES | 2675 | 1% | 1% | 9 |
| 6 | JOURNAL OF BIOLOGICAL CHEMISTRY | 1716 | 9% | 0% | 104 |
| 7 | BIOCHEMICAL JOURNAL | 1289 | 3% | 0% | 39 |
| 8 | NEUROCHEMICAL RESEARCH | 1120 | 2% | 0% | 18 |
| 9 | EUROPEAN JOURNAL OF BIOCHEMISTRY | 981 | 2% | 0% | 28 |
| 10 | ARCHIVES OF BIOCHEMISTRY AND BIOPHYSICS | 836 | 2% | 0% | 24 |
Author Key Words |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | TEKIN, I , ROSKOSKI, R , CARKACI-SALLI, N , VRANA, KE , (2014) COMPLEX MOLECULAR REGULATION OF TYROSINE HYDROXYLASE.JOURNAL OF NEURAL TRANSMISSION. VOL. 121. ISSUE 12. P. 1451 -1481 | 160 | 59% | 8 |
| 2 | FITZPATRICK, PF , (1999) TETRAHYDROPTERIN-DEPENDENT AMINO ACID HYDROXYLASES.ANNUAL REVIEW OF BIOCHEMISTRY. VOL. 68. ISSUE . P. 355 -381 | 112 | 90% | 282 |
| 3 | KAPPOCK, TJ , CARADONNA, JP , (1996) PTERIN-DEPENDENT AMINO ACID HYDROXYLASES.CHEMICAL REVIEWS. VOL. 96. ISSUE 7. P. 2659 -2756 | 233 | 47% | 229 |
| 4 | DICKSON, PW , VON NAGY-FELSOBUKI, EI , GRAHAM, ME , BOBROVSKAYA, L , DUNKLEY, PR , (2004) TYROSINE HYDROXYLASE PHOSPHORYLATION: REGULATION AND CONSEQUENCES.JOURNAL OF NEUROCHEMISTRY. VOL. 91. ISSUE 5. P. 1025-1043 | 94 | 87% | 249 |
| 5 | FITZPATRICK, PF , (2000) THE AROMATIC AMINO ACID HYDROXYLASES.ADVANCES IN ENZYMOLOGY, VOL 74. VOL. 74. ISSUE . P. 235 -+ | 125 | 79% | 59 |
| 6 | FITZPATRICK, PF , (2012) ALLOSTERIC REGULATION OF PHENYLALANINE HYDROXYLASE.ARCHIVES OF BIOCHEMISTRY AND BIOPHYSICS. VOL. 519. ISSUE 2. P. 194-201 | 76 | 90% | 23 |
| 7 | HUFTON, SE , JENNINGS, IG , COTTON, RGH , (1995) STRUCTURE AND FUNCTION OF THE AROMATIC AMINO-ACID HYDROXYLASES.BIOCHEMICAL JOURNAL. VOL. 311. ISSUE . P. 353 -366 | 127 | 80% | 151 |
| 8 | KUMER, SC , VRANA, KE , (1996) INTRICATE REGULATION OF TYROSINE HYDROXYLASE ACTIVITY AND GENE EXPRESSION.JOURNAL OF NEUROCHEMISTRY. VOL. 67. ISSUE 2. P. 443 -462 | 106 | 58% | 514 |
| 9 | WANG, SZ , SURA, GR , DANGOTT, LJ , FITZPATRICK, PF , (2009) IDENTIFICATION BY HYDROGEN/DEUTERIUM EXCHANGE OF STRUCTURAL CHANGES IN TYROSINE HYDROXYLASE ASSOCIATED WITH REGULATION.BIOCHEMISTRY. VOL. 48. ISSUE 22. P. 4972-4979 | 51 | 91% | 6 |
| 10 | OLSSON, E , TEIGEN, K , MARTINEZ, A , JENSEN, VR , (2010) THE AROMATIC AMINO ACID HYDROXYLASE MECHANISM: A PERSPECTIVE FROM COMPUTATIONAL CHEMISTRY.ADVANCES IN INORGANIC CHEMISTRY: THEORETICAL AND COMPUTATIONAL INORGANIC CHEMISTRY, VOL 62. VOL. 62. ISSUE . P. 437-500 | 72 | 55% | 5 |
Classes with closest relation at Level 1 |