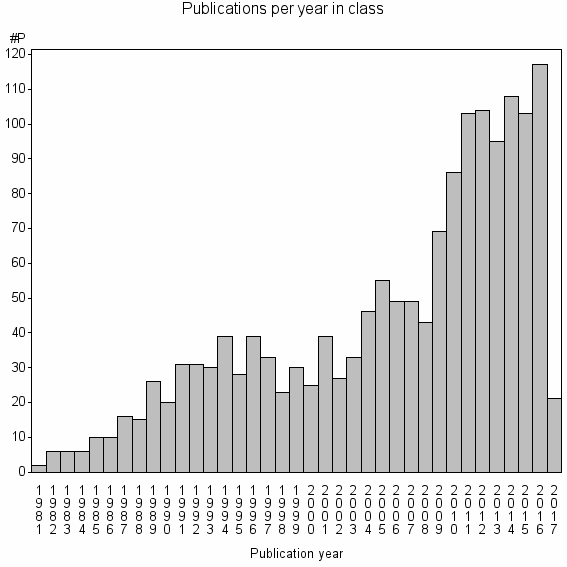
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 5802 | 1573 | 51.0 | 77% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 219 | 3 | PROTEINS-STRUCTURE FUNCTION AND BIOINFORMATICS//PROTEIN FOLDING//PROTEIN STRUCTURE PREDICTION | 50409 |
| 260 | 2 | PROTEIN FOLDING//JOURNAL OF CHEMICAL THEORY AND COMPUTATION//JOURNAL OF COMPUTATIONAL CHEMISTRY | 19582 |
| 5802 | 1 | THERMODYNAMIC INTEGRATION//FREE ENERGY CALCULATIONS//FREE ENERGY PERTURBATION | 1573 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | THERMODYNAMIC INTEGRATION | authKW | 337651 | 3% | 36% | 49 |
| 2 | FREE ENERGY CALCULATIONS | authKW | 328667 | 4% | 25% | 68 |
| 3 | FREE ENERGY PERTURBATION | authKW | 267396 | 3% | 34% | 41 |
| 4 | ONE STEP PERTURBATION | authKW | 213515 | 1% | 100% | 11 |
| 5 | SAMPL4 | authKW | 194105 | 1% | 100% | 10 |
| 6 | SAMPL5 | authKW | 186336 | 1% | 80% | 12 |
| 7 | LINEAR INTERACTION ENERGY | authKW | 145564 | 1% | 50% | 15 |
| 8 | BINDING FREE ENERGY | authKW | 132414 | 3% | 15% | 46 |
| 9 | JOURNAL OF COMPUTER-AIDED MOLECULAR DESIGN | journal | 123298 | 7% | 6% | 106 |
| 10 | ALCHEMICAL | authKW | 118887 | 0% | 88% | 7 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Physics, Atomic, Molecular & Chemical | 8800 | 30% | 0% | 471 |
| 2 | Chemistry, Physical | 6681 | 44% | 0% | 688 |
| 3 | Computer Science, Interdisciplinary Applications | 4161 | 14% | 0% | 225 |
| 4 | Biophysics | 2451 | 15% | 0% | 237 |
| 5 | Chemistry, Multidisciplinary | 1847 | 24% | 0% | 378 |
| 6 | Chemistry, Medicinal | 1259 | 9% | 0% | 142 |
| 7 | Biochemistry & Molecular Biology | 805 | 22% | 0% | 351 |
| 8 | Computer Science, Information Systems | 384 | 5% | 0% | 73 |
| 9 | Mathematical & Computational Biology | 92 | 2% | 0% | 27 |
| 10 | Biochemical Research Methods | 31 | 2% | 0% | 39 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | MOL MODELING SIMULAT | 82408 | 1% | 22% | 19 |
| 2 | COMPUTAT BIOL BIOMOL DYNAM | 38821 | 0% | 100% | 2 |
| 3 | PHARMACEUT CHEM 513 PARNASSUS AVE | 38821 | 0% | 100% | 2 |
| 4 | COMPUTAT BIOL CHEM | 26213 | 1% | 14% | 10 |
| 5 | CHEM PHYS NIH | 25879 | 0% | 67% | 2 |
| 6 | CHEM PL BIOSCI COMPUTAT SCI | 25879 | 0% | 67% | 2 |
| 7 | COMPUTAT CHEM LEAD IDENTIFICAT OPTIMIZAT SUPPOR | 25879 | 0% | 67% | 2 |
| 8 | LAUFER PHYS QUANTITAT BIOL | 23946 | 1% | 11% | 11 |
| 9 | ANTI CANC DRUG DISCOVERY DEV PROGRAM | 19410 | 0% | 100% | 1 |
| 10 | BEIDA YANGSHENG TANG JOINT NAT PROD | 19410 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | JOURNAL OF COMPUTER-AIDED MOLECULAR DESIGN | 123298 | 7% | 6% | 106 |
| 2 | JOURNAL OF CHEMICAL THEORY AND COMPUTATION | 117926 | 11% | 4% | 167 |
| 3 | JOURNAL OF COMPUTATIONAL CHEMISTRY | 61229 | 9% | 2% | 139 |
| 4 | JOURNAL OF CHEMICAL INFORMATION AND MODELING | 27968 | 4% | 2% | 67 |
| 5 | JOURNAL OF PHYSICAL CHEMISTRY B | 9795 | 9% | 0% | 140 |
| 6 | MOLECULAR SIMULATION | 5746 | 2% | 1% | 28 |
| 7 | JOURNAL OF CHEMICAL PHYSICS | 4640 | 9% | 0% | 146 |
| 8 | PROTEINS-STRUCTURE FUNCTION AND BIOINFORMATICS | 3671 | 2% | 1% | 35 |
| 9 | PERSPECTIVES IN DRUG DISCOVERY AND DESIGN | 2134 | 0% | 3% | 4 |
| 10 | JOURNAL OF MOLECULAR GRAPHICS & MODELLING | 1734 | 1% | 1% | 13 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | THERMODYNAMIC INTEGRATION | 337651 | 3% | 36% | 49 | Search THERMODYNAMIC+INTEGRATION | Search THERMODYNAMIC+INTEGRATION |
| 2 | FREE ENERGY CALCULATIONS | 328667 | 4% | 25% | 68 | Search FREE+ENERGY+CALCULATIONS | Search FREE+ENERGY+CALCULATIONS |
| 3 | FREE ENERGY PERTURBATION | 267396 | 3% | 34% | 41 | Search FREE+ENERGY+PERTURBATION | Search FREE+ENERGY+PERTURBATION |
| 4 | ONE STEP PERTURBATION | 213515 | 1% | 100% | 11 | Search ONE+STEP+PERTURBATION | Search ONE+STEP+PERTURBATION |
| 5 | SAMPL4 | 194105 | 1% | 100% | 10 | Search SAMPL4 | Search SAMPL4 |
| 6 | SAMPL5 | 186336 | 1% | 80% | 12 | Search SAMPL5 | Search SAMPL5 |
| 7 | LINEAR INTERACTION ENERGY | 145564 | 1% | 50% | 15 | Search LINEAR+INTERACTION+ENERGY | Search LINEAR+INTERACTION+ENERGY |
| 8 | BINDING FREE ENERGY | 132414 | 3% | 15% | 46 | Search BINDING+FREE+ENERGY | Search BINDING+FREE+ENERGY |
| 9 | ALCHEMICAL | 118887 | 0% | 88% | 7 | Search ALCHEMICAL | Search ALCHEMICAL |
| 10 | HYDRATION FREE ENERGY | 105819 | 1% | 32% | 17 | Search HYDRATION+FREE+ENERGY | Search HYDRATION+FREE+ENERGY |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | HANSEN, N , VAN GUNSTEREN, WF , (2014) PRACTICAL ASPECTS OF FREE-ENERGY CALCULATIONS: A REVIEW.JOURNAL OF CHEMICAL THEORY AND COMPUTATION. VOL. 10. ISSUE 7. P. 2632 -2647 | 143 | 49% | 63 |
| 2 | GENHEDEN, S , RYDE, U , (2015) THE MM/PBSA AND MM/GBSA METHODS TO ESTIMATE LIGAND-BINDING AFFINITIES.EXPERT OPINION ON DRUG DISCOVERY. VOL. 10. ISSUE 5. P. 449 -461 | 64 | 65% | 62 |
| 3 | BARON, R , MCCAMMON, JA , (2013) MOLECULAR RECOGNITION AND LIGAND ASSOCIATION.ANNUAL REVIEW OF PHYSICAL CHEMISTRY, VOL 64. VOL. 64. ISSUE . P. 151 -175 | 78 | 60% | 55 |
| 4 | MOBLEY, DL , KLIMOVICH, PV , (2012) PERSPECTIVE: ALCHEMICAL FREE ENERGY CALCULATIONS FOR DRUG DISCOVERY.JOURNAL OF CHEMICAL PHYSICS. VOL. 137. ISSUE 23. P. - | 76 | 66% | 35 |
| 5 | BOSISIO, S , MEY, ASJS , MICHEL, J , (2017) BLINDED PREDICTIONS OF HOST-GUEST STANDARD FREE ENERGIES OF BINDING IN THE SAMPL5 CHALLENGE.JOURNAL OF COMPUTER-AIDED MOLECULAR DESIGN. VOL. 31. ISSUE 1. P. 61 -70 | 30 | 70% | 3 |
| 6 | DENG, YQ , ROUX, B , (2009) COMPUTATIONS OF STANDARD BINDING FREE ENERGIES WITH MOLECULAR DYNAMICS SIMULATIONS.JOURNAL OF PHYSICAL CHEMISTRY B. VOL. 113. ISSUE 8. P. 2234-2246 | 61 | 53% | 225 |
| 7 | SUAREZ, D , DIAZ, N , (2015) DIRECT METHODS FOR COMPUTING SINGLE-MOLECULE ENTROPIES FROM MOLECULAR SIMULATIONS.WILEY INTERDISCIPLINARY REVIEWS-COMPUTATIONAL MOLECULAR SCIENCE. VOL. 5. ISSUE 1. P. 1 -26 | 56 | 62% | 6 |
| 8 | LEE, HC , HSU, WC , LIU, AL , HSU, CJ , SUN, YC , (2014) USING THERMODYNAMIC INTEGRATION MD SIMULATION TO COMPUTE RELATIVE PROTEIN-LIGAND BINDING FREE ENERGY OF A GSK3 BETA KINASE INHIBITOR AND ITS ANALOGS.JOURNAL OF MOLECULAR GRAPHICS & MODELLING. VOL. 51. ISSUE . P. 37 -49 | 49 | 73% | 6 |
| 9 | BIDER, NS , TSCHOPP, JP , HUNENBERGER, PH , (2015) MULTISTATE LAMBDA-LOCAL-ELEVATION UMBRELLA-SAMPLING (MS-LAMBDA-LEUS): METHOD AND APPLICATION TO THE COMPLEXATION OF CATIONS BY CROWN ETHERS.JOURNAL OF CHEMICAL THEORY AND COMPUTATION. VOL. 11. ISSUE 6. P. 2575 -2588 | 63 | 57% | 0 |
| 10 | MICHEL, J , ESSEX, JW , (2010) PREDICTION OF PROTEIN-LIGAND BINDING AFFINITY BY FREE ENERGY SIMULATIONS: ASSUMPTIONS, PITFALLS AND EXPECTATIONS.JOURNAL OF COMPUTER-AIDED MOLECULAR DESIGN. VOL. 24. ISSUE 8. P. 639 -658 | 67 | 47% | 91 |
Classes with closest relation at Level 1 |