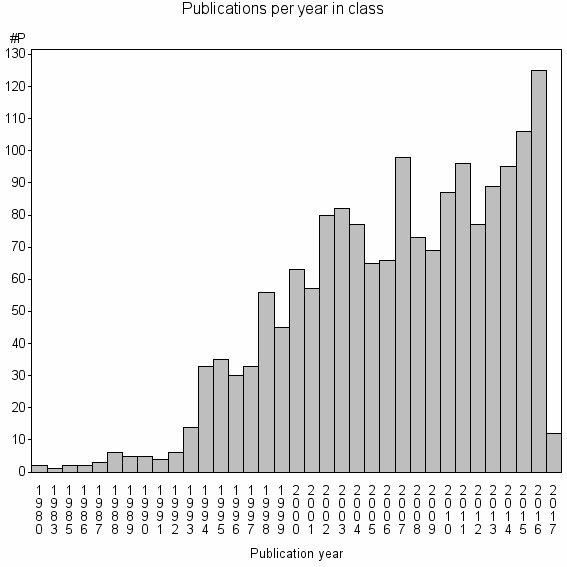
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 4945 | 1699 | 52.5 | 87% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 3 | 4 | PLANT SCIENCES//AGRONOMY//FOOD SCIENCE & TECHNOLOGY | 1882812 |
| 7 | 3 | PLANT SCIENCES//PLANT PHYSIOLOGY//ARABIDOPSIS | 149859 |
| 88 | 2 | PLANT SCIENCES//SALICYLIC ACID//JASMONIC ACID | 26878 |
| 4945 | 1 | NBS LRR//CLADOSPORIUM FULVUM//RESISTANCE GENE ANALOGS | 1699 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | NBS LRR | authKW | 712004 | 3% | 70% | 57 |
| 2 | CLADOSPORIUM FULVUM | authKW | 439801 | 2% | 64% | 38 |
| 3 | RESISTANCE GENE ANALOGS | authKW | 386188 | 2% | 61% | 35 |
| 4 | DISEASE RESISTANCE GENES | authKW | 354388 | 3% | 42% | 47 |
| 5 | SAINSBURY | address | 337869 | 6% | 18% | 104 |
| 6 | SGT1 | authKW | 311477 | 2% | 67% | 26 |
| 7 | R GENE | authKW | 308179 | 3% | 32% | 54 |
| 8 | NBS LRR GENES | authKW | 253146 | 1% | 78% | 18 |
| 9 | NUCLEOTIDE BINDING SITE NBS | authKW | 224630 | 1% | 83% | 15 |
| 10 | NUCLEOTIDE BINDING SITE | authKW | 204157 | 2% | 41% | 28 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Plant Sciences | 34003 | 61% | 0% | 1044 |
| 2 | Genetics & Heredity | 4169 | 21% | 0% | 364 |
| 3 | Horticulture | 3620 | 8% | 0% | 132 |
| 4 | Biochemistry & Molecular Biology | 3484 | 39% | 0% | 665 |
| 5 | Cell Biology | 1544 | 17% | 0% | 288 |
| 6 | Agronomy | 1520 | 8% | 0% | 144 |
| 7 | Biotechnology & Applied Microbiology | 1461 | 14% | 0% | 231 |
| 8 | Evolutionary Biology | 148 | 2% | 0% | 40 |
| 9 | Parasitiology | 138 | 2% | 0% | 35 |
| 10 | Virology | 132 | 3% | 0% | 45 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | SAINSBURY | 337869 | 6% | 18% | 104 |
| 2 | PLANT MICROBE INTERACT | 97454 | 3% | 12% | 47 |
| 3 | CURRICULUM GENET | 95426 | 1% | 30% | 18 |
| 4 | PHYTOPATHOL | 71219 | 5% | 5% | 78 |
| 5 | HORT BIOL | 49915 | 0% | 56% | 5 |
| 6 | SPICE GENOM GRP | 35942 | 0% | 100% | 2 |
| 7 | SUSTAINABLE DIS ISTANCE TEAM | 35942 | 0% | 100% | 2 |
| 8 | UR11ES10 | 35942 | 0% | 100% | 2 |
| 9 | PROGRAM PLANT MOL BIOL BIOTECHNOL | 26953 | 0% | 50% | 3 |
| 10 | PLANT PATHOL | 23773 | 9% | 1% | 156 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | MOLECULAR PLANT-MICROBE INTERACTIONS | 77312 | 7% | 4% | 121 |
| 2 | PLANT CELL | 39101 | 7% | 2% | 114 |
| 3 | MOLECULAR PLANT PATHOLOGY | 26558 | 2% | 4% | 42 |
| 4 | PLANT JOURNAL | 24783 | 6% | 1% | 100 |
| 5 | CURRENT OPINION IN PLANT BIOLOGY | 11106 | 2% | 2% | 33 |
| 6 | PHYSIOLOGICAL AND MOLECULAR PLANT PATHOLOGY | 9289 | 2% | 2% | 31 |
| 7 | THEORETICAL AND APPLIED GENETICS | 7995 | 4% | 1% | 67 |
| 8 | MOLECULAR GENETICS AND GENOMICS | 7599 | 2% | 1% | 29 |
| 9 | ANNUAL REVIEW OF PHYTOPATHOLOGY | 4442 | 1% | 2% | 14 |
| 10 | GENOME | 2544 | 1% | 1% | 24 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | NBS LRR | 712004 | 3% | 70% | 57 | Search NBS+LRR | Search NBS+LRR |
| 2 | CLADOSPORIUM FULVUM | 439801 | 2% | 64% | 38 | Search CLADOSPORIUM+FULVUM | Search CLADOSPORIUM+FULVUM |
| 3 | RESISTANCE GENE ANALOGS | 386188 | 2% | 61% | 35 | Search RESISTANCE+GENE+ANALOGS | Search RESISTANCE+GENE+ANALOGS |
| 4 | DISEASE RESISTANCE GENES | 354388 | 3% | 42% | 47 | Search DISEASE+RESISTANCE+GENES | Search DISEASE+RESISTANCE+GENES |
| 5 | SGT1 | 311477 | 2% | 67% | 26 | Search SGT1 | Search SGT1 |
| 6 | R GENE | 308179 | 3% | 32% | 54 | Search R+GENE | Search R+GENE |
| 7 | NBS LRR GENES | 253146 | 1% | 78% | 18 | Search NBS+LRR+GENES | Search NBS+LRR+GENES |
| 8 | NUCLEOTIDE BINDING SITE NBS | 224630 | 1% | 83% | 15 | Search NUCLEOTIDE+BINDING+SITE+NBS | Search NUCLEOTIDE+BINDING+SITE+NBS |
| 9 | NUCLEOTIDE BINDING SITE | 204157 | 2% | 41% | 28 | Search NUCLEOTIDE+BINDING+SITE | Search NUCLEOTIDE+BINDING+SITE |
| 10 | TOMATO LEAF MOLD | 199060 | 1% | 92% | 12 | Search TOMATO+LEAF+MOLD | Search TOMATO+LEAF+MOLD |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | SUKARTA, OCA , SLOOTWEG, EJ , GOVERSE, A , (2016) STRUCTURE-INFORMED INSIGHTS FOR NLR FUNCTIONING IN PLANT IMMUNITY.SEMINARS IN CELL & DEVELOPMENTAL BIOLOGY. VOL. 56. ISSUE . P. 134 -149 | 107 | 74% | 2 |
| 2 | JACOB, F , VERNALDI, S , MAEKAWA, T , (2013) EVOLUTION AND CONSERVATION OF PLANT NLR FUNCTIONS.FRONTIERS IN IMMUNOLOGY. VOL. 4. ISSUE . P. - | 104 | 65% | 46 |
| 3 | CUI, HT , TSUDA, K , PARKER, JE , (2015) EFFECTOR-TRIGGERED IMMUNITY: FROM PATHOGEN PERCEPTION TO ROBUST DEFENSE.ANNUAL REVIEW OF PLANT BIOLOGY, VOL 66. VOL. 66. ISSUE . P. 487 -511 | 86 | 56% | 44 |
| 4 | MARONE, D , RUSSO, MA , LAIDO, G , DE LEONARDIS, AM , MASTRANGELO, AM , (2013) PLANT NUCLEOTIDE BINDING SITE-LEUCINE-RICH REPEAT (NBS-LRR) GENES: ACTIVE GUARDIANS IN HOST DEFENSE RESPONSES.INTERNATIONAL JOURNAL OF MOLECULAR SCIENCES. VOL. 14. ISSUE 4. P. 7302-7326 | 84 | 69% | 66 |
| 5 | MARTIN, GB , BOGDANOVE, AJ , SESSA, G , (2003) UNDERSTANDING THE FUNCTIONS OF PLANT DISEASE RESISTANCE PROTEINS.ANNUAL REVIEW OF PLANT BIOLOGY. VOL. 54. ISSUE . P. 23 -61 | 111 | 55% | 492 |
| 6 | SEKHWAL, MK , LI, PC , LAM, I , WANG, XE , CLOUTIER, S , YOU, FM , (2015) DISEASE RESISTANCE GENE ANALOGS (RGAS) IN PLANTS.INTERNATIONAL JOURNAL OF MOLECULAR SCIENCES. VOL. 16. ISSUE 8. P. 19248 -19290 | 130 | 48% | 8 |
| 7 | WANG, GF , JI, JB , EI-KASMI, F , DANGL, JL , JOHAL, G , BALINT-KURTI, PJ , (2015) MOLECULAR AND FUNCTIONAL ANALYSES OF A MAIZE AUTOACTIVE NB-LRR PROTEIN IDENTIFY PRECISE STRUCTURAL REQUIREMENTS FOR ACTIVITY.PLOS PATHOGENS. VOL. 11. ISSUE 2. P. - | 60 | 91% | 12 |
| 8 | HULBERT, SH , WEBB, CA , SMITH, SM , SUN, Q , (2001) RESISTANCE GENE COMPLEXES: EVOLUTION AND UTILIZATION.ANNUAL REVIEW OF PHYTOPATHOLOGY. VOL. 39. ISSUE . P. 285 -312 | 93 | 64% | 430 |
| 9 | QI, D , INNES, RW , (2013) RECENT ADVANCES IN PLANT NLR STRUCTURE, FUNCTION, LOCALIZATION, AND SIGNALING.FRONTIERS IN IMMUNOLOGY. VOL. 4. ISSUE . P. - | 72 | 77% | 32 |
| 10 | LEE, HA , YEOM, SI , (2015) PLANT NB-LRR PROTEINS: TIGHTLY REGULATED SENSORS IN A COMPLEX MANNER.BRIEFINGS IN FUNCTIONAL GENOMICS. VOL. 14. ISSUE 4. P. 233 -242 | 86 | 62% | 7 |
Classes with closest relation at Level 1 |
| Rank | Class id | link |
|---|---|---|
| 1 | 12560 | RECEPTOR LIKE KINASE//FLG22//RLK |
| 2 | 6203 | NPR1//SALICYLIC ACID//SYSTEMIC ACQUIRED RESISTANCE |
| 3 | 11167 | ELICITIN//OOMYCETES//CRYPTOGEIN |
| 4 | 3976 | MOLECULAR PLANT-MICROBE INTERACTIONS//HARPIN//XANTHOMONAS |
| 5 | 7729 | RICE BLAST//BACTERIAL BLIGHT//MAGNAPORTHE GRISEA |
| 6 | 11560 | POWDERY MILDEW//MLO//BLUMERIA GRAMINIS F SP HORDEI |
| 7 | 18980 | WRKY TRANSCRIPTION FACTOR//WRKY//WRKY PROTEIN |
| 8 | 26060 | NON HOST RESISTANCE//POST PENETRATION//NONHOST RESISTANCE |
| 9 | 35133 | COPINE//N COPINE//BON1 |
| 10 | 19896 | U BOX//RING H2//SAUL1 |