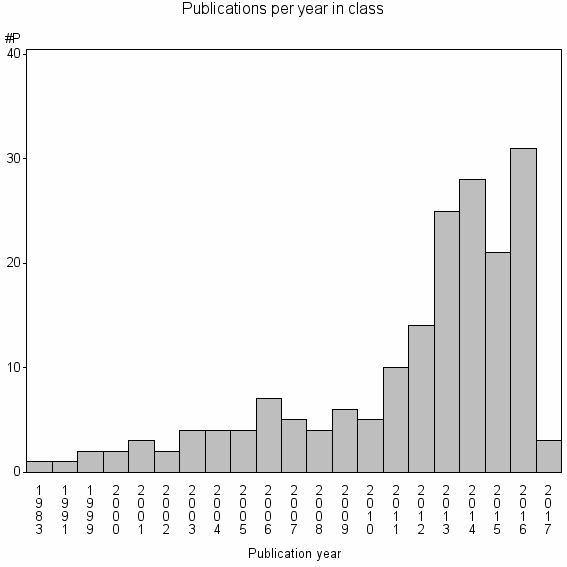
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 29672 | 182 | 44.7 | 80% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 3 | 4 | PLANT SCIENCES//AGRONOMY//FOOD SCIENCE & TECHNOLOGY | 1882812 |
| 69 | 3 | PLANT SCIENCES//EVOLUTIONARY BIOLOGY//AMERICAN JOURNAL OF BOTANY | 83717 |
| 826 | 2 | CONSERVATION GENETICS RESOURCES//MICROSATELLITES//PHYLOGEOGRAPHY | 11863 |
| 29672 | 1 | PHYLODYNAMICS//TRANSMISSION TREE//SERIAL SAMPLES | 182 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | PHYLODYNAMICS | authKW | 1255028 | 13% | 31% | 24 |
| 2 | TRANSMISSION TREE | authKW | 671105 | 3% | 67% | 6 |
| 3 | SERIAL SAMPLES | authKW | 419438 | 3% | 50% | 5 |
| 4 | ASSEMBLY BASED APPROACH | authKW | 167777 | 1% | 100% | 1 |
| 5 | BAYESIAN DIFFUSION MODELS | authKW | 167777 | 1% | 100% | 1 |
| 6 | BIOINFORMATIC TOOLS AND DATABASES | authKW | 167777 | 1% | 100% | 1 |
| 7 | BRANCH PARTITIONING | authKW | 167777 | 1% | 100% | 1 |
| 8 | CALIBRATED SKYWIS PLOT | authKW | 167777 | 1% | 100% | 1 |
| 9 | CHRONICALLY INFECTED INDIVIDUAL | authKW | 167777 | 1% | 100% | 1 |
| 10 | COMMUNICABLE DIS DYNAMICS | address | 167777 | 1% | 100% | 1 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Evolutionary Biology | 2756 | 26% | 0% | 47 |
| 2 | Mathematical & Computational Biology | 2087 | 20% | 0% | 36 |
| 3 | Genetics & Heredity | 1453 | 37% | 0% | 68 |
| 4 | Biochemical Research Methods | 572 | 19% | 0% | 35 |
| 5 | Microbiology | 307 | 18% | 0% | 33 |
| 6 | Biology | 198 | 10% | 0% | 18 |
| 7 | Infectious Diseases | 155 | 9% | 0% | 17 |
| 8 | Biotechnology & Applied Microbiology | 107 | 12% | 0% | 21 |
| 9 | Ecology | 75 | 8% | 0% | 15 |
| 10 | Biochemistry & Molecular Biology | 36 | 16% | 0% | 29 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | COMMUNICABLE DIS DYNAMICS | 167777 | 1% | 100% | 1 |
| 2 | EMOSTA | 167777 | 1% | 100% | 1 |
| 3 | EQUIPE MODELISAT STOCHAST PL EMOSTA | 167777 | 1% | 100% | 1 |
| 4 | EVOLUT BIOINFORMAT | 167777 | 1% | 100% | 1 |
| 5 | HLTH PROTECT AGCY ABORATING OXFORD | 167777 | 1% | 100% | 1 |
| 6 | HOSP UNIV MEDITERRANEE INFECTUM63CNRS7278 | 167777 | 1% | 100% | 1 |
| 7 | INFECT DIS INFORMAT UNIT | 167777 | 1% | 100% | 1 |
| 8 | JOHN RADCLIFFE HOSP LEVEL 5 | 167777 | 1% | 100% | 1 |
| 9 | MAT LICACOES FUNDAMENT | 167777 | 1% | 100% | 1 |
| 10 | MIVEGECUMR 5290IRD 224 | 167777 | 1% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | MOLECULAR BIOLOGY AND EVOLUTION | 12041 | 11% | 0% | 20 |
| 2 | PLOS COMPUTATIONAL BIOLOGY | 10482 | 9% | 0% | 17 |
| 3 | EPIDEMICS | 3047 | 1% | 1% | 2 |
| 4 | JOURNAL OF THE ROYAL SOCIETY INTERFACE | 1693 | 3% | 0% | 5 |
| 5 | GENETICS | 1579 | 6% | 0% | 11 |
| 6 | PHILOSOPHICAL TRANSACTIONS OF THE ROYAL SOCIETY B-BIOLOGICAL SCIENCES | 1366 | 4% | 0% | 7 |
| 7 | GENOME MEDICINE | 1022 | 1% | 0% | 2 |
| 8 | BIOINFORMATICS | 762 | 4% | 0% | 7 |
| 9 | BRITISH JOURNAL OF BIOMEDICAL SCIENCE | 755 | 1% | 0% | 2 |
| 10 | TRENDS IN ECOLOGY & EVOLUTION | 683 | 2% | 0% | 3 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | PHYLODYNAMICS | 1255028 | 13% | 31% | 24 | Search PHYLODYNAMICS | Search PHYLODYNAMICS |
| 2 | TRANSMISSION TREE | 671105 | 3% | 67% | 6 | Search TRANSMISSION+TREE | Search TRANSMISSION+TREE |
| 3 | SERIAL SAMPLES | 419438 | 3% | 50% | 5 | Search SERIAL+SAMPLES | Search SERIAL+SAMPLES |
| 4 | ASSEMBLY BASED APPROACH | 167777 | 1% | 100% | 1 | Search ASSEMBLY+BASED+APPROACH | Search ASSEMBLY+BASED+APPROACH |
| 5 | BAYESIAN DIFFUSION MODELS | 167777 | 1% | 100% | 1 | Search BAYESIAN+DIFFUSION+MODELS | Search BAYESIAN+DIFFUSION+MODELS |
| 6 | BIOINFORMATIC TOOLS AND DATABASES | 167777 | 1% | 100% | 1 | Search BIOINFORMATIC+TOOLS+AND+DATABASES | Search BIOINFORMATIC+TOOLS+AND+DATABASES |
| 7 | BRANCH PARTITIONING | 167777 | 1% | 100% | 1 | Search BRANCH+PARTITIONING | Search BRANCH+PARTITIONING |
| 8 | CALIBRATED SKYWIS PLOT | 167777 | 1% | 100% | 1 | Search CALIBRATED+SKYWIS+PLOT | Search CALIBRATED+SKYWIS+PLOT |
| 9 | CHRONICALLY INFECTED INDIVIDUAL | 167777 | 1% | 100% | 1 | Search CHRONICALLY+INFECTED+INDIVIDUAL | Search CHRONICALLY+INFECTED+INDIVIDUAL |
| 10 | CORRECTED AKAIKE CRITERION | 167777 | 1% | 100% | 1 | Search CORRECTED+AKAIKE+CRITERION | Search CORRECTED+AKAIKE+CRITERION |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | FROST, SDW , PYBUS, OG , GOG, JR , VIBOUD, C , BONHOEFFER, S , BEDFORD, T , (2015) EIGHT CHALLENGES IN PHYLODYNAMIC INFERENCE.EPIDEMICS. VOL. 10. ISSUE . P. 88 -92 | 19 | 41% | 17 |
| 2 | DE MAIO, N , WU, CH , WILSON, DJ , (2016) SCOTTI: EFFICIENT RECONSTRUCTION OF TRANSMISSION WITHIN OUTBREAKS WITH THE STRUCTURED COALESCENT.PLOS COMPUTATIONAL BIOLOGY. VOL. 12. ISSUE 9. P. - | 19 | 46% | 0 |
| 3 | KENAH, E , BRITTON, T , HALLORAN, ME , LONGINI, IM , (2016) MOLECULAR INFECTIOUS DISEASE EPIDEMIOLOGY: SURVIVAL ANALYSIS AND ALGORITHMS LINKING PHYLOGENIES TO TRANSMISSION TREES.PLOS COMPUTATIONAL BIOLOGY. VOL. 12. ISSUE 4. P. - | 21 | 41% | 0 |
| 4 | HALL, MD , WOOLHOUSE, MEJ , RAMBAUT, A , (2016) USING GENOMICS DATA TO RECONSTRUCT TRANSMISSION TREES DURING DISEASE OUTBREAKS.REVUE SCIENTIFIQUE ET TECHNIQUE-OFFICE INTERNATIONAL DES EPIZOOTIES. VOL. 35. ISSUE 1. P. 287 -296 | 15 | 56% | 0 |
| 5 | VOLZ, EM , FROST, SDW , (2014) SAMPLING THROUGH TIME AND PHYLODYNAMIC INFERENCE WITH COALESCENT AND BIRTH-DEATH MODELS.JOURNAL OF THE ROYAL SOCIETY INTERFACE. VOL. 11. ISSUE 101. P. - | 14 | 54% | 4 |
| 6 | WORBY, CJ , CHANG, HH , HANAGE, WP , LIPSITCH, M , (2014) THE DISTRIBUTION OF PAIRWISE GENETIC DISTANCES: A TOOL FOR INVESTIGATING DISEASE TRANSMISSION.GENETICS. VOL. 198. ISSUE 4. P. 1395 -+ | 12 | 57% | 5 |
| 7 | HALL, M , WOOLHOUSE, M , RAMBAUT, A , (2015) EPIDEMIC RECONSTRUCTION IN A PHYLOGENETICS FRAMEWORK: TRANSMISSION TREES AS PARTITIONS OF THE NODE SET.PLOS COMPUTATIONAL BIOLOGY. VOL. 11. ISSUE 12. P. - | 17 | 35% | 6 |
| 8 | WORBY, CJ , O'NEILL, PD , KYPRAIOS, T , ROBOTHAM, JV , DE ANGELIS, D , CARTWRIGHT, EJP , PEACOCK, SJ , (2016) RECONSTRUCTING TRANSMISSION TREES FOR COMMUNICABLE DISEASES USING DENSELY SAMPLED GENETIC DATA.ANNALS OF APPLIED STATISTICS. VOL. 10. ISSUE 1. P. 395 -417 | 12 | 46% | 1 |
| 9 | LI, LM , GRASSLY, NC , FRASER, C , (2014) GENOMIC ANALYSIS OF EMERGING PATHOGENS: METHODS, APPLICATION AND FUTURE TRENDS.GENOME BIOLOGY. VOL. 15. ISSUE 11. P. - | 18 | 33% | 5 |
| 10 | STADLER, T , VAUGHAN, TG , GAVRYUSHKIN, A , GUINDON, S , KUHNERT, D , LEVENTHAL, GE , DRUMMOND, AJ , (2015) HOW WELL CAN THE EXPONENTIAL-GROWTH COALESCENT APPROXIMATE CONSTANT-RATE BIRTH-DEATH POPULATION DYNAMICS?.PROCEEDINGS OF THE ROYAL SOCIETY B-BIOLOGICAL SCIENCES. VOL. 282. ISSUE 1806. P. - | 15 | 39% | 1 |
Classes with closest relation at Level 1 |