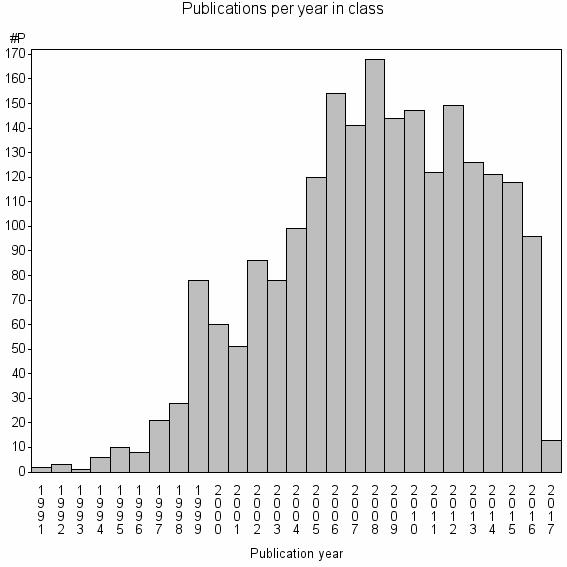
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 2775 | 2150 | 43.0 | 87% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 5 | 4 | CHEMISTRY, ORGANIC//CHEMISTRY, INORGANIC & NUCLEAR//CHEMISTRY, MULTIDISCIPLINARY | 1745167 |
| 183 | 3 | APTAMER//G QUADRUPLEX//RIBOZYME | 56307 |
| 740 | 2 | JOURNAL OF BIOMOLECULAR STRUCTURE & DYNAMICS//SINGLE MOLECULE FORCE SPECTROSCOPY//Z DNA | 12558 |
| 2775 | 1 | SINGLE MOLECULE FORCE SPECTROSCOPY//FORCE SPECTROSCOPY//MECHANICAL UNFOLDING | 2150 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | SINGLE MOLECULE FORCE SPECTROSCOPY | authKW | 761831 | 5% | 55% | 98 |
| 2 | FORCE SPECTROSCOPY | authKW | 583715 | 6% | 34% | 122 |
| 3 | MECHANICAL UNFOLDING | authKW | 324910 | 1% | 74% | 31 |
| 4 | LEHRSTUHL ANGEW PHYS | address | 303193 | 3% | 36% | 59 |
| 5 | DYNAMIC FORCE SPECTROSCOPY | authKW | 206146 | 1% | 52% | 28 |
| 6 | PHYS E22 | address | 159299 | 2% | 30% | 37 |
| 7 | PROTEIN STRETCHING | authKW | 146061 | 1% | 86% | 12 |
| 8 | UNBINDING FORCE | authKW | 122730 | 1% | 79% | 11 |
| 9 | IMETUM | address | 102506 | 1% | 38% | 19 |
| 10 | STATE PRECIS MEASUREMENTS TECHNOL R | address | 100981 | 0% | 89% | 8 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biophysics | 9983 | 25% | 0% | 540 |
| 2 | Chemistry, Physical | 2110 | 23% | 0% | 494 |
| 3 | Chemistry, Multidisciplinary | 1555 | 20% | 0% | 422 |
| 4 | Biochemistry & Molecular Biology | 1078 | 22% | 0% | 476 |
| 5 | Microscopy | 940 | 2% | 0% | 51 |
| 6 | Nanoscience & Nanotechnology | 831 | 8% | 0% | 162 |
| 7 | Materials Science, Multidisciplinary | 642 | 15% | 0% | 327 |
| 8 | Physics, Atomic, Molecular & Chemical | 524 | 7% | 0% | 159 |
| 9 | Polymer Science | 324 | 6% | 0% | 125 |
| 10 | Cell Biology | 208 | 7% | 0% | 153 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | LEHRSTUHL ANGEW PHYS | 303193 | 3% | 36% | 59 |
| 2 | PHYS E22 | 159299 | 2% | 30% | 37 |
| 3 | IMETUM | 102506 | 1% | 38% | 19 |
| 4 | STATE PRECIS MEASUREMENTS TECHNOL R | 100981 | 0% | 89% | 8 |
| 5 | NANONEUROBIOPHYS GRP | 63900 | 0% | 75% | 6 |
| 6 | CHRISTIAN DOPPLER NANOSCOP METHODS BIOPHYS | 61354 | 1% | 39% | 11 |
| 7 | ERNST BERL MAKROMOL CHEM | 56803 | 0% | 100% | 4 |
| 8 | SINGLE MOL STUDY | 44363 | 0% | 31% | 10 |
| 9 | CHAIR PL PHYS | 43615 | 1% | 24% | 13 |
| 10 | BIOL INSPIRED MAT MAT SYST | 38311 | 1% | 15% | 18 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BIOPHYSICAL JOURNAL | 35831 | 10% | 1% | 222 |
| 2 | JOURNAL OF MOLECULAR RECOGNITION | 13412 | 2% | 3% | 34 |
| 3 | LANGMUIR | 5704 | 6% | 0% | 128 |
| 4 | ULTRAMICROSCOPY | 3237 | 2% | 1% | 36 |
| 5 | SCIENTIFIC MODELING AND SIMULATIONS | 3152 | 0% | 11% | 2 |
| 6 | CHEMPHYSCHEM | 2645 | 2% | 1% | 33 |
| 7 | PROCEEDINGS OF THE NATIONAL ACADEMY OF SCIENCES OF THE UNITED STATES OF AMERICA | 1295 | 5% | 0% | 98 |
| 8 | JOURNAL OF MOLECULAR BIOLOGY | 1214 | 2% | 0% | 46 |
| 9 | CURRENT OPINION IN STRUCTURAL BIOLOGY | 1197 | 1% | 1% | 14 |
| 10 | STRUCTURE | 929 | 1% | 0% | 16 |
Author Key Words |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | OTT, W , JOBST, MA , SCHOELER, C , GAUB, HE , NASH, MA , (2017) SINGLE-MOLECULE FORCE SPECTROSCOPY ON POLYPROTEINS AND RECEPTOR-LIGAND COMPLEXES: THE CURRENT TOOLBOX.JOURNAL OF STRUCTURAL BIOLOGY. VOL. 197. ISSUE 1. P. 3 -12 | 85 | 71% | 1 |
| 2 | HUGHES, ML , DOUGAN, L , (2016) THE PHYSICS OF PULLING POLYPROTEINS: A REVIEW OF SINGLE MOLECULE FORCE SPECTROSCOPY USING THE AFM TO STUDY PROTEIN UNFOLDING.REPORTS ON PROGRESS IN PHYSICS. VOL. 79. ISSUE 7. P. - | 129 | 58% | 0 |
| 3 | GIANNOTTI, MI , VANCSO, GJ , (2007) INTERROGATION OF SINGLE SYNTHETIC POLYMER CHAINS AND POLYSACCHARIDES BY AFM-BASED FORCE SPECTROSCOPY.CHEMPHYSCHEM. VOL. 8. ISSUE 16. P. 2290 -2307 | 97 | 60% | 53 |
| 4 | SCHOLL, ZN , LI, Q , MARSZALEK, PE , (2014) SINGLE MOLECULE MECHANICAL MANIPULATION FOR STUDYING BIOLOGICAL PROPERTIES OF PROTEINS, DNA, AND SUGARS.WILEY INTERDISCIPLINARY REVIEWS-NANOMEDICINE AND NANOBIOTECHNOLOGY. VOL. 6. ISSUE 3. P. 211-229 | 75 | 67% | 9 |
| 5 | HOFFMANN, T , TYCH, KM , HUGHES, ML , BROCKWELL, DJ , DOUGAN, L , (2013) TOWARDS DESIGN PRINCIPLES FOR DETERMINING THE MECHANICAL STABILITY OF PROTEINS.PHYSICAL CHEMISTRY CHEMICAL PHYSICS. VOL. 15. ISSUE 38. P. 15767-15780 | 69 | 73% | 9 |
| 6 | LI, HB , (2008) 'MECHANICAL ENGINEERING' OF ELASTOMERIC PROTEINS: TOWARD DESIGNING NEW PROTEIN BUILDING BLOCKS FOR BIOMATERIALS.ADVANCED FUNCTIONAL MATERIALS. VOL. 18. ISSUE 18. P. 2643 -2657 | 74 | 76% | 16 |
| 7 | GUO, S , RAY, C , KIRKPATRICK, A , LAD, N , AKHREMITCHEV, BB , (2008) EFFECTS OF MULTIPLE-BOND RUPTURES ON KINETIC PARAMETERS EXTRACTED FROM FORCE SPECTROSCOPY MEASUREMENTS: REVISITING BIOTIN-STREPTAVIDIN INTERACTIONS.BIOPHYSICAL JOURNAL. VOL. 95. ISSUE 8. P. 3964 -3976 | 62 | 86% | 36 |
| 8 | KIM, Y , KIM, W , PARK, JW , (2016) PRINCIPLES AND APPLICATIONS OF FORCE SPECTROSCOPY USING ATOMIC FORCE MICROSCOPY.BULLETIN OF THE KOREAN CHEMICAL SOCIETY. VOL. 37. ISSUE 12. P. 1895 -1907 | 84 | 57% | 0 |
| 9 | BIZZARRI, AR , CANNISTRARO, S , (2010) THE APPLICATION OF ATOMIC FORCE SPECTROSCOPY TO THE STUDY OF BIOLOGICAL COMPLEXES UNDERGOING A BIORECOGNITION PROCESS.CHEMICAL SOCIETY REVIEWS. VOL. 39. ISSUE 2. P. 734 -749 | 79 | 59% | 54 |
| 10 | CRAMPTON, N , BROCKWELL, DJ , (2010) UNRAVELLING THE DESIGN PRINCIPLES FOR SINGLE PROTEIN MECHANICAL STRENGTH.CURRENT OPINION IN STRUCTURAL BIOLOGY. VOL. 20. ISSUE 4. P. 508 -517 | 65 | 74% | 35 |
Classes with closest relation at Level 1 |