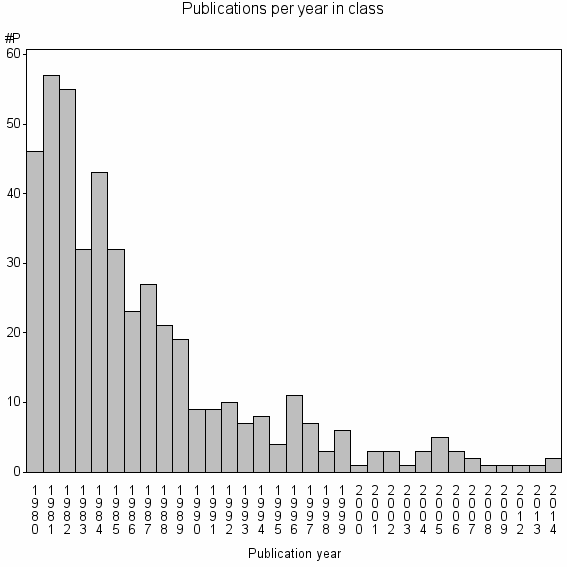
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 20513 | 456 | 38.1 | 50% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 71 | 3 | GENETICS & HEREDITY//CHROMATIN//CELL BIOLOGY | 83302 |
| 1094 | 2 | NUCLEAR MATRIX//LINKER HISTONE//HISTONE H1 | 9829 |
| 20513 | 1 | CHICKEN LYSOZYME GENE//MACROPHAGE SPECIFIC GENE EXPRESSION//ANAEROBIC RESPONSE ELEMENT | 456 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | CHICKEN LYSOZYME GENE | authKW | 150664 | 1% | 75% | 3 |
| 2 | MACROPHAGE SPECIFIC GENE EXPRESSION | authKW | 89282 | 0% | 67% | 2 |
| 3 | ANAEROBIC RESPONSE ELEMENT | authKW | 66963 | 0% | 100% | 1 |
| 4 | BENT DNA IN NUCLEOSOMES | authKW | 66963 | 0% | 100% | 1 |
| 5 | C EBP BETANF M | authKW | 66963 | 0% | 100% | 1 |
| 6 | CPG DEMETHYLATION | authKW | 66963 | 0% | 100% | 1 |
| 7 | DNA STRUCTURE IN NUCLEOSOMES | authKW | 66963 | 0% | 100% | 1 |
| 8 | DNASE I HYPERSENSITIVE SITE BETA A GLOBIN GENE | authKW | 66963 | 0% | 100% | 1 |
| 9 | DNASE SENSITIVE REGION | authKW | 66963 | 0% | 100% | 1 |
| 10 | GLOBULIN GENE | authKW | 66963 | 0% | 100% | 1 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biochemistry & Molecular Biology | 3250 | 68% | 0% | 311 |
| 2 | Cell Biology | 779 | 22% | 0% | 102 |
| 3 | Virology | 609 | 9% | 0% | 42 |
| 4 | CYTOLOGY & HISTOLOGY | 213 | 1% | 0% | 5 |
| 5 | Biophysics | 211 | 9% | 0% | 40 |
| 6 | Biology | 109 | 5% | 0% | 23 |
| 7 | Genetics & Heredity | 53 | 6% | 0% | 27 |
| 8 | Biochemical Research Methods | 18 | 3% | 0% | 14 |
| 9 | Multidisciplinary Sciences | 6 | 1% | 0% | 4 |
| 10 | Developmental Biology | 6 | 1% | 0% | 5 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | MOL MED EPIDEMIOL CANC | 22320 | 0% | 33% | 1 |
| 2 | ST JAMESS UNIV HOSP | 5172 | 1% | 1% | 6 |
| 3 | ST JAMES UNIV HOSP | 3882 | 1% | 2% | 3 |
| 4 | MOL MED UNIT | 2641 | 2% | 1% | 7 |
| 5 | MOL BIOL PROGRAM | 1509 | 0% | 1% | 2 |
| 6 | MUTAGENESE | 1173 | 0% | 2% | 1 |
| 7 | BIOMED BIOMOL SCI | 920 | 0% | 1% | 2 |
| 8 | BIOPHYS S | 846 | 0% | 1% | 1 |
| 9 | SECT EXPT HAEMATOL | 742 | 0% | 1% | 1 |
| 10 | PROGRAM PLANT MOL CELLULAR BIOL | 565 | 0% | 1% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | NUCLEIC ACIDS RESEARCH | 7594 | 15% | 0% | 67 |
| 2 | UCLA SYMPOSIA ON MOLECULAR AND CELLULAR BIOLOGY | 4934 | 1% | 2% | 3 |
| 3 | POSTEPY BIOCHEMII | 3475 | 0% | 3% | 2 |
| 4 | JOURNAL OF MOLECULAR BIOLOGY | 1913 | 6% | 0% | 26 |
| 5 | CELL | 1767 | 4% | 0% | 20 |
| 6 | COLD SPRING HARBOR SYMPOSIA ON QUANTITATIVE BIOLOGY | 1469 | 2% | 0% | 7 |
| 7 | VIROLOGY | 1276 | 4% | 0% | 19 |
| 8 | MOLECULAR AND CELLULAR BIOLOGY | 1178 | 4% | 0% | 20 |
| 9 | MOLECULAR BIOLOGY | 1142 | 2% | 0% | 9 |
| 10 | DNA AND CELL BIOLOGY | 880 | 1% | 0% | 6 |
Author Key Words |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | GROSS, DS , GARRARD, WT , (1988) NUCLEASE HYPERSENSITIVE SITES IN CHROMATIN.ANNUAL REVIEW OF BIOCHEMISTRY. VOL. 57. ISSUE . P. 159-197 | 82 | 20% | 936 |
| 2 | SIPPEL, AE , SAUERESSIG, H , HUBER, MC , HOEFER, HC , STIEF, A , BORGMEYER, U , BONIFER, C , (1996) IDENTIFICATION OF CIS-ACTING ELEMENTS AS DNASE I HYPERSENSITIVE SITES IN LYSOZYME GENE CHROMATIN.RNA POLYMERASE AND ASSOCIATED FACTORS, PT B. VOL. 274. ISSUE . P. 233-246 | 17 | 68% | 13 |
| 3 | EISSENBERG, JC , CARTWRIGHT, IL , THOMAS, GH , ELGIN, SCR , (1985) SELECTED TOPICS IN CHROMATIN STRUCTURE.ANNUAL REVIEW OF GENETICS. VOL. 19. ISSUE . P. 485-536 | 62 | 22% | 220 |
| 4 | KUBE, D , MILAVETZ, B , (1996) DIFFERENTIAL REGULATION BY SV40 T-ANTIGEN BINDING AT SITE I DEFINES TWO DISTINCT CLASSES OF NUCLEOSOME-FREE PROMOTER.ANATOMICAL RECORD. VOL. 244. ISSUE 1. P. 28-32 | 13 | 76% | 0 |
| 5 | KONDOLEON, SK , KURKINEN, NA , HALLICK, LM , (1989) THE SV40 NUCLEOSOME-FREE REGION IS DETECTED THROUGHOUT THE VIRUS LIFE-CYCLE.VIROLOGY. VOL. 173. ISSUE 1. P. 129-135 | 15 | 83% | 5 |
| 6 | KUBE, D , MILAVETZ, B , (1989) GENERATION OF A NUCLEOSOME-FREE PROMOTER REGION IN SV40 DOES NOT REQUIRE T-ANTIGEN BINDING TO SITE-I.VIROLOGY. VOL. 172. ISSUE 1. P. 100-105 | 17 | 68% | 3 |
| 7 | HERMANSEN, R , SIERRA, MA , JOHNSON, J , FRIEZ, M , MILAVETZ, B , (1996) IDENTIFICATION OF SIMIAN VIRUS 40 PROMOTER DNA SEQUENCES CAPABLE OF CONFERRING RESTRICTION ENDONUCLEASE HYPERSENSITIVITY.JOURNAL OF VIROLOGY. VOL. 70. ISSUE 6. P. 3416-3422 | 17 | 49% | 9 |
| 8 | FRIEZ, M , HERMANSEN, R , MILAVETZ, B , (1999) CHROMATIN STRUCTURE OF THE SIMIAN VIRUS 40 LATE PROMOTER: A DELETIONAL ANALYSIS.JOURNAL OF VIROLOGY. VOL. 73. ISSUE 3. P. 1990-1997 | 15 | 47% | 11 |
| 9 | LEFEVRE, P , MELNIK, S , WILSON, N , RIGGS, AD , BONIFER, C , (2003) DEVELOPMENTALLY REGULATED RECRUITMENT OF TRANSCRIPTION FACTORS AND CHROMATIN MODIFICATION ACTIVITIES TO CHICKEN LYSOZYME CIS-REGULATORY ELEMENTS IN VIVO.MOLECULAR AND CELLULAR BIOLOGY. VOL. 23. ISSUE 12. P. 4386-4400 | 19 | 30% | 36 |
| 10 | CHU, Y , HUANG, TS , HSU, MT , (1990) P1 NUCLEASE DEFINES A SUBPOPULATION OF ACTIVE SV40 CHROMATIN - A NEW NUCLEASE HYPERSENSITIVITY ASSAY.NUCLEIC ACIDS RESEARCH. VOL. 18. ISSUE 13. P. 3705-3711 | 14 | 70% | 0 |
Classes with closest relation at Level 1 |