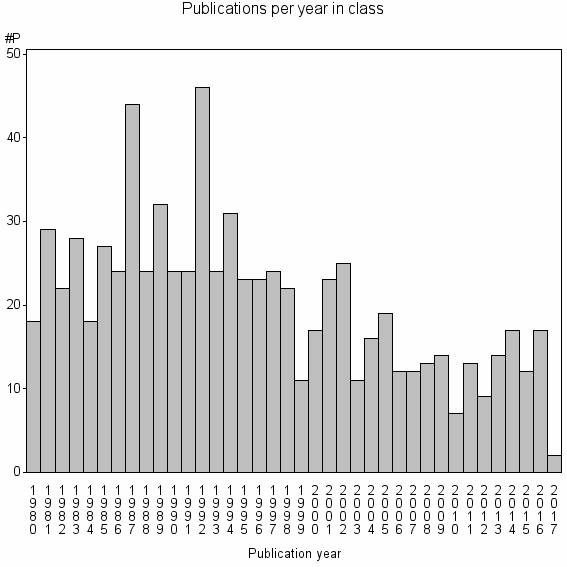
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 14399 | 771 | 30.8 | 56% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 278 | 3 | CORYNEBACTERIUM GLUTAMICUM//ACTA CRYSTALLOGRAPHICA SECTION D-BIOLOGICAL CRYSTALLOGRAPHY//BIOCHEMISTRY & MOLECULAR BIOLOGY | 43163 |
| 1967 | 2 | GLUTAMATE DEHYDROGENASE//3 ISOPROPYLMALATE DEHYDROGENASE//ISOCITRATE DEHYDROGENASE | 5642 |
| 14399 | 1 | MALATE DEHYDROGENASE//D LACTATE DEHYDROGENASE//D 2 HYDROXYACID DEHYDROGENASE | 771 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | MALATE DEHYDROGENASE | authKW | 499405 | 9% | 18% | 72 |
| 2 | D LACTATE DEHYDROGENASE | authKW | 207208 | 2% | 35% | 15 |
| 3 | D 2 HYDROXYACID DEHYDROGENASE | authKW | 123757 | 1% | 63% | 5 |
| 4 | CYTOSOLIC MALATE DEHYDROGENASE GENE | authKW | 118810 | 0% | 100% | 3 |
| 5 | L LACTATE DEHYDROGENASE | authKW | 105591 | 2% | 22% | 12 |
| 6 | LACTATE DEHYDROGENASE | authKW | 96573 | 9% | 4% | 66 |
| 7 | 2 KETOPANTOATE REDUCTASE | authKW | 79207 | 0% | 100% | 2 |
| 8 | APAD | authKW | 79207 | 0% | 100% | 2 |
| 9 | D 2 HYDROXY 4 METHYLVALERATE DEHYDROGENASE | authKW | 79207 | 0% | 100% | 2 |
| 10 | D 2 HYDROXYISOCAPROATE DEHYDROGENASE | authKW | 79207 | 0% | 100% | 2 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biochemistry & Molecular Biology | 4923 | 65% | 0% | 500 |
| 2 | Biophysics | 1336 | 16% | 0% | 122 |
| 3 | Biotechnology & Applied Microbiology | 323 | 10% | 0% | 77 |
| 4 | Microbiology | 189 | 8% | 0% | 61 |
| 5 | Parasitiology | 164 | 3% | 0% | 24 |
| 6 | Biochemical Research Methods | 96 | 5% | 0% | 36 |
| 7 | Genetics & Heredity | 22 | 4% | 0% | 29 |
| 8 | Crystallography | 17 | 2% | 0% | 15 |
| 9 | Plant Sciences | 9 | 3% | 0% | 25 |
| 10 | Cell Biology | 8 | 4% | 0% | 30 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | SOIL EROS DRY LAND FARMING LOESS P | 52803 | 0% | 67% | 2 |
| 2 | BIOCHEM MOL RECOGNIT | 39603 | 0% | 100% | 1 |
| 3 | BIOPHYS MOL BIOL STRUCT JP EBELUMR 5075 | 39603 | 0% | 100% | 1 |
| 4 | CROUCHER HUMAN GENOMICS | 39603 | 0% | 100% | 1 |
| 5 | MED BIOMED SCI BIOCHEM MOL BIOL | 39603 | 0% | 100% | 1 |
| 6 | MOL CELL BIOL BIOL | 39603 | 0% | 100% | 1 |
| 7 | MOL MED GENET PROT ENGN NETWORK EXCELLENC | 39603 | 0% | 100% | 1 |
| 8 | UR 477 BIOCHIM BACTERIENNE | 39603 | 0% | 100% | 1 |
| 9 | WINOGRAFSKY MICROBIOL | 39603 | 0% | 100% | 1 |
| 10 | MOL BIOL BIOTECHNOL GENET MOBGAM | 35639 | 0% | 30% | 3 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BIOLOGICAL CHEMISTRY HOPPE-SEYLER | 8395 | 2% | 1% | 18 |
| 2 | INTERNATIONAL JOURNAL OF BIOCHEMISTRY | 4210 | 2% | 1% | 18 |
| 3 | PROTEIN ENGINEERING | 2503 | 2% | 1% | 12 |
| 4 | EUROPEAN JOURNAL OF BIOCHEMISTRY | 2141 | 4% | 0% | 33 |
| 5 | BIOCHEMISTRY | 2103 | 7% | 0% | 54 |
| 6 | MOLECULAR AND BIOCHEMICAL PARASITOLOGY | 1157 | 2% | 0% | 13 |
| 7 | JOURNAL OF MOLECULAR BIOLOGY | 1110 | 3% | 0% | 26 |
| 8 | HOPPE-SEYLERS ZEITSCHRIFT FUR PHYSIOLOGISCHE CHEMIE | 786 | 1% | 1% | 4 |
| 9 | BIOCHIMICA ET BIOPHYSICA ACTA | 695 | 3% | 0% | 23 |
| 10 | ACTA CRYSTALLOGRAPHICA SECTION F-STRUCTURAL BIOLOGY COMMUNICATIONS | 456 | 1% | 0% | 6 |
Author Key Words |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | ARAI, K , ICHIKAWA, J , NONAKA, S , MIYANAGA, A , UCHIKOBA, H , FUSHINOBU, S , TAGUCHI, H , (2011) A MOLECULAR DESIGN THAT STABILIZES ACTIVE STATE IN BACTERIAL ALLOSTERIC L-LACTATE DEHYDROGENASES.JOURNAL OF BIOCHEMISTRY. VOL. 150. ISSUE 5. P. 579 -591 | 46 | 87% | 1 |
| 2 | TAKAHASHI-INIGUEZ, T , ABURTO-RODRIGUEZ, N , VILCHIS-GONZALEZ, AL , FLORES, ME , (2016) FUNCTION, KINETIC PROPERTIES, CRYSTALLIZATION, AND REGULATION OF MICROBIAL MALATE DEHYDROGENASE.JOURNAL OF ZHEJIANG UNIVERSITY-SCIENCE B. VOL. 17. ISSUE 4. P. 247 -261 | 51 | 59% | 0 |
| 3 | IKEHARA, Y , ARAI, K , FURUKAWA, N , OHNO, T , MIYAKE, T , FUSHINOBU, S , NAKAJIMA, M , MIYANAGA, A , TAGUCHI, H , (2014) THE CORE OF ALLOSTERIC MOTION IN THERMUS CALDOPHILUS L-LACTATE DEHYDROGENASE.JOURNAL OF BIOLOGICAL CHEMISTRY. VOL. 289. ISSUE 45. P. 31550 -31564 | 32 | 80% | 1 |
| 4 | ARAI, K , ISHIMITSU, T , FUSHINOBU, S , UCHIKOBA, H , MATSUZAWA, H , TAGUCHI, H , (2010) ACTIVE AND INACTIVE STATE STRUCTURES OF UNLIGANDED LACTABACILLUS CASEI ALLOSTERIC L-LACTATE DEHYDROGENASE.PROTEINS-STRUCTURE FUNCTION AND BIOINFORMATICS. VOL. 78. ISSUE 3. P. 681-694 | 29 | 85% | 7 |
| 5 | ARAI, K , HISHIDA, A , ISHIYAMA, M , KAMATA, T , UCHIKOBA, H , FUSHINOBU, S , MATSUZAWA, H , TAGUCHI, H , (2002) AN ABSOLUTE REQUIREMENT OF FRUCTOSE 1,6-BISPHOSPHATE FOR THE LACTOBACILLUS CASEI L-LACTATE DEHYDROGENASE ACTIVITY INDUCED BY A SINGLE AMINO ACID SUBSTITUTION.PROTEIN ENGINEERING. VOL. 15. ISSUE 1. P. 35 -41 | 32 | 94% | 12 |
| 6 | GOWARD, CR , NICHOLLS, DJ , (1994) MALATE-DEHYDROGENASE - A MODEL FOR STRUCTURE, EVOLUTION, AND CATALYSIS.PROTEIN SCIENCE. VOL. 3. ISSUE 10. P. 1883-1888 | 32 | 82% | 122 |
| 7 | TOKUDA, C , ISHIKURA, Y , SHIGEMATSU, M , MUTOH, H , TSUZUKI, S , NAKAHIRA, Y , TAMURA, Y , SHINODA, T , ARAI, K , TAKAHASHI, O , ET AL (2003) CONVERSION OF LACTOBACILLUS PENTOSUS D-LACTATE DEHYDROGENASE TO A D-HYDROXYISOCAPROATE DEHYDROGENASE THROUGH A SINGLE AMINO ACID REPLACEMENT.JOURNAL OF BACTERIOLOGY. VOL. 185. ISSUE 16. P. 5023-5026 | 26 | 84% | 23 |
| 8 | PENG, HL , EGAWA, T , CHANG, E , DENG, H , CALLENDER, R , (2015) MECHANISM OF THERMAL ADAPTATION IN THE LACTATE DEHYDROGENASES.JOURNAL OF PHYSICAL CHEMISTRY B. VOL. 119. ISSUE 49. P. 15256 -15262 | 19 | 76% | 1 |
| 9 | WANG, ZD , WANG, BJ , GE, YD , PAN, W , WANG, J , XU, L , LIU, AM , ZHU, GP , (2011) EXPRESSION AND IDENTIFICATION OF A THERMOSTABLE MALATE DEHYDROGENASE FROM MULTICELLULAR PROKARYOTE STREPTOMYCES AVERMITILIS MA-4680.MOLECULAR BIOLOGY REPORTS. VOL. 38. ISSUE 3. P. 1629 -1636 | 25 | 61% | 8 |
| 10 | ARAI, K , KAMATA, T , UCHIKOBA, H , FUSHINOBU, S , MATSUZAWA, H , TAGUCHI, H , (2001) SOME LACTOBACILLUS L-LACTATE DEHYDROGENASES EXHIBIT COMPARABLE CATALYTIC ACTIVITIES FOR PYRUVATE AND OXALOACETATE.JOURNAL OF BACTERIOLOGY. VOL. 183. ISSUE 1. P. 397 -400 | 23 | 85% | 14 |
Classes with closest relation at Level 1 |