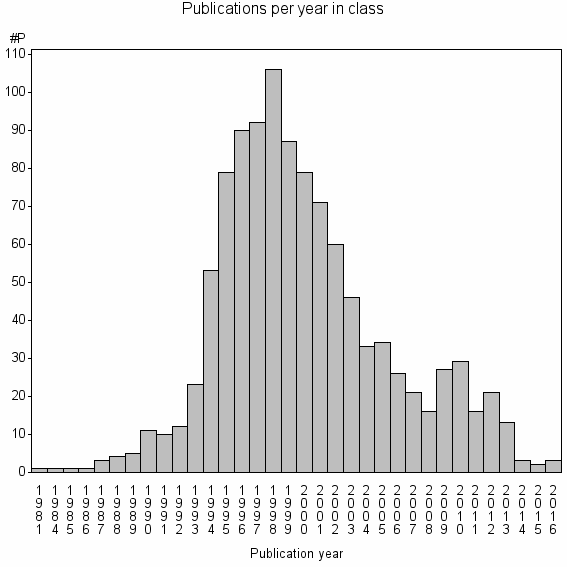
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 10241 | 1079 | 25.1 | 82% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 90 | 3 | MATHEMATICAL & COMPUTATIONAL BIOLOGY//BIOINFORMATICS//BMC BIOINFORMATICS | 77178 |
| 2924 | 2 | SAGE//SERIAL ANALYSIS OF GENE EXPRESSION//DIFFERENTIAL DISPLAY | 2737 |
| 10241 | 1 | DIFFERENTIAL DISPLAY//MRNA DIFFERENTIAL DISPLAY//ANIM NUTR FEED YUNNAN PROV | 1079 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | DIFFERENTIAL DISPLAY | authKW | 391872 | 11% | 12% | 117 |
| 2 | MRNA DIFFERENTIAL DISPLAY | authKW | 271059 | 4% | 24% | 40 |
| 3 | ANIM NUTR FEED YUNNAN PROV | address | 205374 | 1% | 48% | 15 |
| 4 | BLACK BONED SHEEP | authKW | 150918 | 1% | 67% | 8 |
| 5 | TISSUE EXPRESSION ANALYSIS | authKW | 100607 | 1% | 44% | 8 |
| 6 | CDNA AFLP | authKW | 97527 | 3% | 11% | 30 |
| 7 | SUBTRACTIVE HYBRIDIZATION | authKW | 94470 | 3% | 10% | 33 |
| 8 | MRNA DIFFERENTIAL DISPLAY PCR | authKW | 84894 | 0% | 100% | 3 |
| 9 | REPRESENTATIONAL DIFFERENCE ANALYSIS | authKW | 77124 | 2% | 16% | 17 |
| 10 | MOL GERONTOL UNIT | address | 75459 | 0% | 67% | 4 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biochemical Research Methods | 3838 | 20% | 0% | 220 |
| 2 | Biochemistry & Molecular Biology | 3518 | 48% | 0% | 517 |
| 3 | Biotechnology & Applied Microbiology | 703 | 12% | 0% | 130 |
| 4 | Genetics & Heredity | 635 | 11% | 0% | 122 |
| 5 | Biophysics | 194 | 6% | 0% | 64 |
| 6 | Cell Biology | 150 | 8% | 0% | 87 |
| 7 | Chemistry, Analytical | 95 | 6% | 0% | 60 |
| 8 | Plant Sciences | 78 | 5% | 0% | 58 |
| 9 | Agriculture, Dairy & Animal Science | 60 | 2% | 0% | 24 |
| 10 | Oncology | 58 | 6% | 0% | 65 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | ANIM NUTR FEED YUNNAN PROV | 205374 | 1% | 48% | 15 |
| 2 | MOL GERONTOL UNIT | 75459 | 0% | 67% | 4 |
| 3 | CENT HUMAN GENOME | 56596 | 0% | 100% | 2 |
| 4 | ANIM NUTR FEED YUNNAN PROVINCE | 37729 | 0% | 67% | 2 |
| 5 | SWINE GENET BREEDING | 29476 | 1% | 7% | 14 |
| 6 | BEE GRP GPA | 28298 | 0% | 100% | 1 |
| 7 | CELL BIOMOL BIOL | 28298 | 0% | 100% | 1 |
| 8 | CORANGE TECHNOL OFF | 28298 | 0% | 100% | 1 |
| 9 | DAKE SLAYTON BRAIN CANC STUDIES | 28298 | 0% | 100% | 1 |
| 10 | DEKE SLAYTON BRAIN CANC STUDIES | 28298 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BIOTECHNIQUES | 34499 | 8% | 1% | 86 |
| 2 | MOLECULAR BIOTECHNOLOGY | 8060 | 2% | 1% | 23 |
| 3 | NUCLEIC ACIDS RESEARCH | 6653 | 9% | 0% | 97 |
| 4 | GENETIC ANALYSIS-BIOMOLECULAR ENGINEERING | 3775 | 1% | 2% | 6 |
| 5 | PROGRESS IN BIOCHEMISTRY AND BIOPHYSICS | 2593 | 2% | 1% | 18 |
| 6 | ANALYTICAL BIOCHEMISTRY | 2281 | 4% | 0% | 39 |
| 7 | TRENDS IN GENETICS | 2017 | 1% | 1% | 14 |
| 8 | MOLECULAR BIOLOGY REPORTS | 1209 | 1% | 0% | 16 |
| 9 | ANIMAL SCIENCE PAPERS AND REPORTS | 1050 | 0% | 1% | 4 |
| 10 | PLANT MOLECULAR BIOLOGY REPORTER | 989 | 1% | 1% | 7 |
Author Key Words |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | STURTEVANT, J , (2000) APPLICATIONS OF DIFFERENTIAL-DISPLAY REVERSE TRANSCRIPTION-PCR TO MOLECULAR PATHOGENESIS AND MEDICAL MYCOLOGY.CLINICAL MICROBIOLOGY REVIEWS. VOL. 13. ISSUE 3. P. 408 -+ | 86 | 61% | 27 |
| 2 | SHIUE, L , (1997) IDENTIFICATION OF CANDIDATE GENES FOR DRUG DISCOVERY BY DIFFERENTIAL DISPLAY.DRUG DEVELOPMENT RESEARCH. VOL. 41. ISSUE 3-4. P. 142-159 | 63 | 65% | 5 |
| 3 | ZHANG, JS , DUNCAN, EL , CHANG, ACM , REDDEL, RR , (1998) DIFFERENTIAL DISPLAY OF MRNA.MOLECULAR BIOTECHNOLOGY. VOL. 10. ISSUE 2. P. 155 -165 | 41 | 71% | 36 |
| 4 | LIEVENS, S , GOORMACHTIG, S , HOLSTERS, M , (2001) A CRITICAL EVALUATION OF DIFFERENTIAL DISPLAY AS A TOOL TO IDENTIFY GENES INVOLVED IN LEGUME NODULATION: LOOKING BACK AND LOOKING FORWARD.NUCLEIC ACIDS RESEARCH. VOL. 29. ISSUE 17. P. 3459 -3468 | 48 | 48% | 39 |
| 5 | LORKOWSKI, S , CULLEN, P , (2004) HIGH-THROUGHPUT ANALYSIS OF MRNA EXPRESSION: MICROARRAYS ARE NOT THE WHOLE STORY.EXPERT OPINION ON THERAPEUTIC PATENTS. VOL. 14. ISSUE 3. P. 377-403 | 47 | 47% | 4 |
| 6 | BURGER, AL , BOTHA, FC , (2004) CLONING OF A SPECIFIC RIPENING-RELATED GENE FROM THE MULTIPLE OF RIPENING-RELATED GENES IDENTIFIED FROM A SINGLE BAND EXCISED FROM A CDNA-AFLP GEL.PLANT MOLECULAR BIOLOGY REPORTER. VOL. 22. ISSUE 3. P. 225-236 | 27 | 79% | 5 |
| 7 | GROHME, M , FROHME, M , MALI, B , (2009) OPEN ARCHITECTURE PCR-BASED METHODS FOR DIFFERENTIAL GENE EXPRESSION ANALYSIS.CURRENT PHARMACEUTICAL ANALYSIS. VOL. 5. ISSUE 1. P. 1-9 | 29 | 60% | 1 |
| 8 | MATZ, MV , LUKYANOV, SA , (1998) DIFFERENT STRATEGIES OF DIFFERENTIAL DISPLAY: AREAS OF APPLICATION.NUCLEIC ACIDS RESEARCH. VOL. 26. ISSUE 24. P. 5537-5543 | 26 | 84% | 58 |
| 9 | ALI, M , MARKHAM, AF , ISAACS, JD , (2001) APPLICATION OF DIFFERENTIAL DISPLAY TO IMMUNOLOGICAL RESEARCH.JOURNAL OF IMMUNOLOGICAL METHODS. VOL. 250. ISSUE 1-2. P. 29-43 | 45 | 45% | 13 |
| 10 | DIATCHENKO, L , LAU, YFC , CAMPBELL, AP , CHENCHIK, A , MOQADAM, F , HUANG, B , LUKYANOV, S , LUKYANOV, K , GURSKAYA, N , SVERDLOV, ED , ET AL (1996) SUPPRESSION SUBTRACTIVE HYBRIDIZATION: A METHOD FOR GENERATING DIFFERENTIALLY REGULATED OR TISSUE-SPECIFIC CDNA PROBES AND LIBRARIES.PROCEEDINGS OF THE NATIONAL ACADEMY OF SCIENCES OF THE UNITED STATES OF AMERICA. VOL. 93. ISSUE 12. P. 6025-6030 | 10 | 77% | 2335 |
Classes with closest relation at Level 1 |