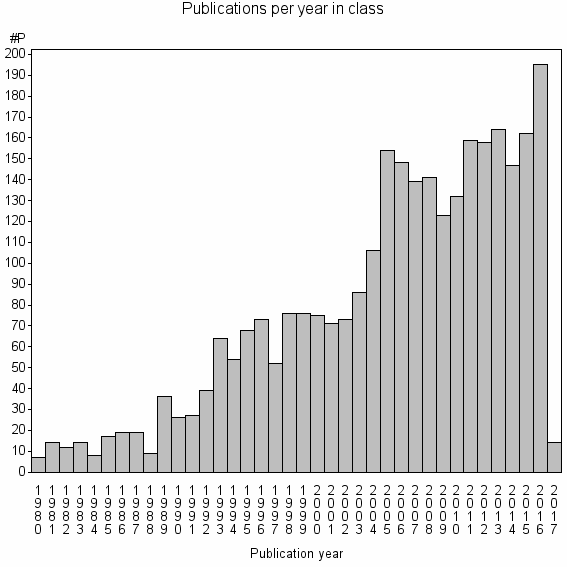
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 922 | 2957 | 37.1 | 65% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 90 | 3 | MATHEMATICAL & COMPUTATIONAL BIOLOGY//BIOINFORMATICS//BMC BIOINFORMATICS | 77178 |
| 126 | 2 | MOLECULAR BIOLOGY AND EVOLUTION//BIOINFORMATICS//MATHEMATICAL & COMPUTATIONAL BIOLOGY | 23963 |
| 922 | 1 | SYSTEMATIC BIOLOGY//PHYLOGENETIC NETWORK//PHYLOGENETIC TREE | 2957 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | SYSTEMATIC BIOLOGY | journal | 963182 | 12% | 26% | 353 |
| 2 | PHYLOGENETIC NETWORK | authKW | 775522 | 4% | 66% | 114 |
| 3 | PHYLOGENETIC TREE | authKW | 303065 | 7% | 14% | 206 |
| 4 | GENE TREE | authKW | 279796 | 3% | 37% | 74 |
| 5 | SPECIES TREE | authKW | 271139 | 3% | 31% | 85 |
| 6 | SUPERTREE | authKW | 270460 | 2% | 46% | 57 |
| 7 | NEIGHBOR JOINING | authKW | 256072 | 2% | 48% | 52 |
| 8 | EVOLUTIONARY TREE | authKW | 221878 | 2% | 42% | 51 |
| 9 | PHYLOGENETIC INFERENCE | authKW | 217001 | 2% | 36% | 58 |
| 10 | PHYLOGENETICS | authKW | 215716 | 8% | 9% | 237 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Evolutionary Biology | 135685 | 45% | 1% | 1320 |
| 2 | Mathematical & Computational Biology | 38796 | 21% | 1% | 625 |
| 3 | Genetics & Heredity | 11803 | 27% | 0% | 795 |
| 4 | Biochemical Research Methods | 6254 | 16% | 0% | 472 |
| 5 | Statistics & Probability | 2919 | 8% | 0% | 240 |
| 6 | Biotechnology & Applied Microbiology | 2289 | 13% | 0% | 384 |
| 7 | Computer Science, Interdisciplinary Applications | 2200 | 8% | 0% | 234 |
| 8 | Biology | 1934 | 8% | 0% | 232 |
| 9 | Biochemistry & Molecular Biology | 1337 | 21% | 0% | 630 |
| 10 | Computer Science, Theory & Methods | 1329 | 6% | 0% | 191 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | ALGORITHMS BIOINFORMAT COMPLEX FORMAL METHODS R | 91973 | 0% | 64% | 14 |
| 2 | BIOINFORMAT ZBIT | 57801 | 0% | 40% | 14 |
| 3 | BIOMATH | 44528 | 4% | 4% | 108 |
| 4 | ALLAN WILSON MOL ECOL EVOLUT | 42330 | 2% | 8% | 51 |
| 5 | DKE | 38909 | 0% | 54% | 7 |
| 6 | INFORMAT FONDAMENTALE PLICAT | 38909 | 0% | 54% | 7 |
| 7 | UNIV VIENNA | 33706 | 0% | 23% | 14 |
| 8 | GK STRUKTURBILDUNGSPROZESSE | 33037 | 0% | 80% | 4 |
| 9 | ROBERT CEDERGREN | 23243 | 0% | 17% | 13 |
| 10 | EXELIXIS | 23217 | 0% | 25% | 9 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | SYSTEMATIC BIOLOGY | 963182 | 12% | 26% | 353 |
| 2 | MOLECULAR BIOLOGY AND EVOLUTION | 125147 | 9% | 5% | 260 |
| 3 | CLADISTICS | 123269 | 3% | 13% | 92 |
| 4 | IEEE-ACM TRANSACTIONS ON COMPUTATIONAL BIOLOGY AND BIOINFORMATICS | 85791 | 3% | 9% | 97 |
| 5 | JOURNAL OF COMPUTATIONAL BIOLOGY | 50027 | 3% | 6% | 87 |
| 6 | CLADISTICS-THE INTERNATIONAL JOURNAL OF THE WILLI HENNIG SOCIETY | 49520 | 1% | 20% | 24 |
| 7 | ALGORITHMS FOR MOLECULAR BIOLOGY | 47219 | 1% | 13% | 36 |
| 8 | MOLECULAR PHYLOGENETICS AND EVOLUTION | 45954 | 5% | 3% | 154 |
| 9 | EVOLUTIONARY BIOINFORMATICS | 33506 | 1% | 10% | 32 |
| 10 | JOURNAL OF CLASSIFICATION | 30726 | 1% | 8% | 37 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | PHYLOGENETIC NETWORK | 775522 | 4% | 66% | 114 | Search PHYLOGENETIC+NETWORK | Search PHYLOGENETIC+NETWORK |
| 2 | PHYLOGENETIC TREE | 303065 | 7% | 14% | 206 | Search PHYLOGENETIC+TREE | Search PHYLOGENETIC+TREE |
| 3 | GENE TREE | 279796 | 3% | 37% | 74 | Search GENE+TREE | Search GENE+TREE |
| 4 | SPECIES TREE | 271139 | 3% | 31% | 85 | Search SPECIES+TREE | Search SPECIES+TREE |
| 5 | SUPERTREE | 270460 | 2% | 46% | 57 | Search SUPERTREE | Search SUPERTREE |
| 6 | NEIGHBOR JOINING | 256072 | 2% | 48% | 52 | Search NEIGHBOR+JOINING | Search NEIGHBOR+JOINING |
| 7 | EVOLUTIONARY TREE | 221878 | 2% | 42% | 51 | Search EVOLUTIONARY+TREE | Search EVOLUTIONARY+TREE |
| 8 | PHYLOGENETIC INFERENCE | 217001 | 2% | 36% | 58 | Search PHYLOGENETIC+INFERENCE | Search PHYLOGENETIC+INFERENCE |
| 9 | PHYLOGENETICS | 215716 | 8% | 9% | 237 | Search PHYLOGENETICS | Search PHYLOGENETICS |
| 10 | PHYLOGENETIC INVARIANTS | 177482 | 1% | 90% | 19 | Search PHYLOGENETIC+INVARIANTS | Search PHYLOGENETIC+INVARIANTS |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | MORRISON, DA , (2006) PHYLOGENETIC ANALYSES OF PARASITES IN THE NEW MILLENNIUM.ADVANCES IN PARASITOLOGY, VOL 63. VOL. 63. ISSUE . P. 1 -124 | 211 | 63% | 7 |
| 2 | GUINDON, S , GASCUEL, O , (2003) A SIMPLE, FAST, AND ACCURATE ALGORITHM TO ESTIMATE LARGE PHYLOGENIES BY MAXIMUM LIKELIHOOD.SYSTEMATIC BIOLOGY. VOL. 52. ISSUE 5. P. 696-704 | 27 | 77% | 9889 |
| 3 | EDWARDS, SV , XI, ZX , JANKE, A , FAIRCLOTH, BC , MCCORMACK, JE , GLENN, TC , ZHONG, BJ , WU, SY , LEMMON, EM , LEMMON, AR , ET AL (2016) IMPLEMENTING AND TESTING THE MULTISPECIES COALESCENT MODEL: A VALUABLE PARADIGM FOR PHYLOGENOMICS.MOLECULAR PHYLOGENETICS AND EVOLUTION. VOL. 94. ISSUE . P. 447 -462 | 66 | 68% | 12 |
| 4 | MIRARAB, S , BAYZID, MS , WARNOW, T , (2016) EVALUATING SUMMARY METHODS FOR MULTILOCUS SPECIES TREE ESTIMATION IN THE PRESENCE OF INCOMPLETE LINEAGE SORTING.SYSTEMATIC BIOLOGY. VOL. 65. ISSUE 3. P. 366 -380 | 44 | 83% | 17 |
| 5 | LIU, L , XI, ZX , WU, SY , DAVIS, CC , EDWARDS, SV , (2015) ESTIMATING PHYLOGENETIC TREES FROM GENOME-SCALE DATA.YEAR IN EVOLUTIONARY BIOLOGY. VOL. 1360. ISSUE . P. 36 -53 | 74 | 77% | 16 |
| 6 | XI, ZX , LIU, L , DAVIS, CC , (2016) THE IMPACT OF MISSING DATA ON SPECIES TREE ESTIMATION.MOLECULAR BIOLOGY AND EVOLUTION. VOL. 33. ISSUE 3. P. 838 -860 | 60 | 65% | 7 |
| 7 | SIMMONS, MP , SLOAN, DB , GATESY, J , (2016) THE EFFECTS OF SUBSAMPLING GENE TREES ON COALESCENT METHODS APPLIED TO ANCIENT DIVERGENCES.MOLECULAR PHYLOGENETICS AND EVOLUTION. VOL. 97. ISSUE . P. 76 -89 | 68 | 69% | 3 |
| 8 | LIU, L , YU, LL , KUBATKO, L , PEARL, DK , EDWARDS, SV , (2009) COALESCENT METHODS FOR ESTIMATING PHYLOGENETIC TREES.MOLECULAR PHYLOGENETICS AND EVOLUTION. VOL. 53. ISSUE 1. P. 320 -328 | 58 | 85% | 146 |
| 9 | GUINDON, S , DUFAYARD, JF , LEFORT, V , ANISIMOVA, M , HORDIJK, W , GASCUEL, O , (2010) NEW ALGORITHMS AND METHODS TO ESTIMATE MAXIMUM-LIKELIHOOD PHYLOGENIES: ASSESSING THE PERFORMANCE OF PHYML 3.0.SYSTEMATIC BIOLOGY. VOL. 59. ISSUE 3. P. 307 -321 | 19 | 66% | 3880 |
| 10 | GATESY, J , SPRINGER, MS , (2014) PHYLOGENETIC ANALYSIS AT DEEP TIMESCALES: UNRELIABLE GENE TREES, BYPASSED HIDDEN SUPPORT, AND THE COALESCENCE/CONCATALESCENCE CONUNDRUM.MOLECULAR PHYLOGENETICS AND EVOLUTION. VOL. 80. ISSUE . P. 231 -266 | 76 | 55% | 44 |
Classes with closest relation at Level 1 |