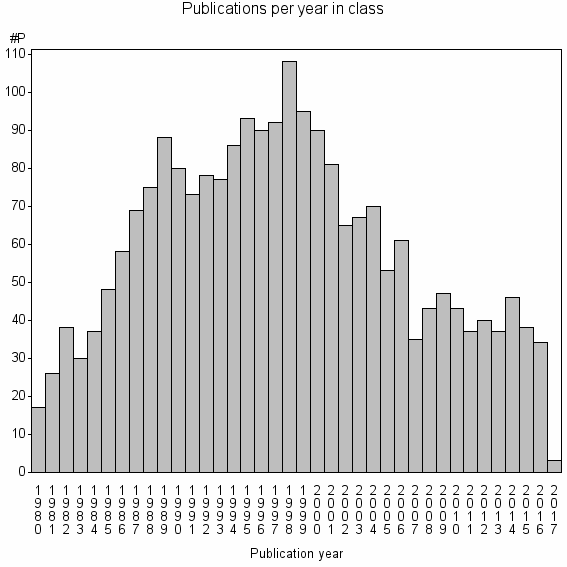
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 2425 | 2248 | 45.0 | 75% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 5 | 4 | CHEMISTRY, ORGANIC//CHEMISTRY, INORGANIC & NUCLEAR//CHEMISTRY, MULTIDISCIPLINARY | 1745167 |
| 183 | 3 | APTAMER//G QUADRUPLEX//RIBOZYME | 56307 |
| 740 | 2 | JOURNAL OF BIOMOLECULAR STRUCTURE & DYNAMICS//SINGLE MOLECULE FORCE SPECTROSCOPY//Z DNA | 12558 |
| 2425 | 1 | DNA CURVATURE//A TRACT//JOURNAL OF BIOMOLECULAR STRUCTURE & DYNAMICS | 2248 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | DNA CURVATURE | authKW | 273847 | 2% | 46% | 44 |
| 2 | A TRACT | authKW | 187156 | 1% | 66% | 21 |
| 3 | JOURNAL OF BIOMOLECULAR STRUCTURE & DYNAMICS | journal | 141951 | 8% | 6% | 181 |
| 4 | DNA FLEXIBILITY | authKW | 137000 | 1% | 39% | 26 |
| 5 | B DNA | authKW | 110206 | 1% | 32% | 25 |
| 6 | BIOCHIM THEOR | address | 103020 | 2% | 16% | 47 |
| 7 | SEQUENCE DEPENDENT STRUCTURE | authKW | 96578 | 0% | 89% | 8 |
| 8 | DNA BENDING | authKW | 96360 | 2% | 15% | 46 |
| 9 | DNA HYDRATION | authKW | 88267 | 1% | 50% | 13 |
| 10 | UPR 9080 | address | 85797 | 1% | 20% | 32 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biochemistry & Molecular Biology | 16086 | 68% | 0% | 1536 |
| 2 | Biophysics | 14339 | 29% | 0% | 657 |
| 3 | Chemistry, Physical | 356 | 11% | 0% | 258 |
| 4 | Biochemical Research Methods | 210 | 4% | 0% | 94 |
| 5 | Chemistry, Multidisciplinary | 188 | 9% | 0% | 203 |
| 6 | Biology | 174 | 3% | 0% | 72 |
| 7 | Mathematical & Computational Biology | 143 | 2% | 0% | 40 |
| 8 | Crystallography | 136 | 3% | 0% | 62 |
| 9 | Physics, Atomic, Molecular & Chemical | 119 | 4% | 0% | 95 |
| 10 | Cell Biology | 44 | 4% | 0% | 101 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BIOCHIM THEOR | 103020 | 2% | 16% | 47 |
| 2 | UPR 9080 | 85797 | 1% | 20% | 32 |
| 3 | MOL BIOPHYS PROGRAM | 83547 | 2% | 18% | 35 |
| 4 | CNRS UPR 9080 | 39139 | 0% | 41% | 7 |
| 5 | INFORMAT CHEM | 28925 | 0% | 30% | 7 |
| 6 | CHEM BIOMOLEC STEREODYNAM | 27163 | 0% | 100% | 2 |
| 7 | SWISS FED POLYTECH | 27163 | 0% | 100% | 2 |
| 8 | JOINT IRB BSC PROGRAM COMPUTAT BIOL | 15054 | 0% | 12% | 9 |
| 9 | MOL MODELING BIOINFORMAT UNIT | 14371 | 0% | 18% | 6 |
| 10 | ADV BIOMED COMP SIL EXCELLENCE | 13582 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | JOURNAL OF BIOMOLECULAR STRUCTURE & DYNAMICS | 141951 | 8% | 6% | 181 |
| 2 | BIOPOLYMERS | 41484 | 6% | 2% | 141 |
| 3 | JOURNAL OF MOLECULAR BIOLOGY | 20679 | 8% | 1% | 189 |
| 4 | NUCLEIC ACIDS RESEARCH | 18817 | 10% | 1% | 235 |
| 5 | BIOPHYSICAL CHEMISTRY | 7010 | 2% | 1% | 47 |
| 6 | BIOPHYSICAL JOURNAL | 6297 | 4% | 0% | 96 |
| 7 | BIOCHEMISTRY | 5706 | 7% | 0% | 152 |
| 8 | MOLECULAR BIOLOGY | 3489 | 2% | 1% | 35 |
| 9 | CURRENT OPINION IN STRUCTURAL BIOLOGY | 3117 | 1% | 1% | 23 |
| 10 | ACTA CRYSTALLOGRAPHICA SECTION D-BIOLOGICAL CRYSTALLOGRAPHY | 2204 | 1% | 1% | 27 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | DNA CURVATURE | 273847 | 2% | 46% | 44 | Search DNA+CURVATURE | Search DNA+CURVATURE |
| 2 | A TRACT | 187156 | 1% | 66% | 21 | Search A+TRACT | Search A+TRACT |
| 3 | DNA FLEXIBILITY | 137000 | 1% | 39% | 26 | Search DNA+FLEXIBILITY | Search DNA+FLEXIBILITY |
| 4 | B DNA | 110206 | 1% | 32% | 25 | Search B+DNA | Search B+DNA |
| 5 | SEQUENCE DEPENDENT STRUCTURE | 96578 | 0% | 89% | 8 | Search SEQUENCE+DEPENDENT+STRUCTURE | Search SEQUENCE+DEPENDENT+STRUCTURE |
| 6 | DNA BENDING | 96360 | 2% | 15% | 46 | Search DNA+BENDING | Search DNA+BENDING |
| 7 | DNA HYDRATION | 88267 | 1% | 50% | 13 | Search DNA+HYDRATION | Search DNA+HYDRATION |
| 8 | A DNA | 67425 | 1% | 41% | 12 | Search A+DNA | Search A+DNA |
| 9 | BASE STACKING INTERACTIONS | 56588 | 0% | 83% | 5 | Search BASE+STACKING+INTERACTIONS | Search BASE+STACKING+INTERACTIONS |
| 10 | DINUCLEOTIDE STEP | 56588 | 0% | 83% | 5 | Search DINUCLEOTIDE+STEP | Search DINUCLEOTIDE+STEP |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | BEVERIDGE, DL , DIXIT, SB , BARREIRO, G , THAYER, KM , (2004) MOLECULAR DYNAMICS SIMULATIONS OF DNA CURVATURE AND FLEXIBILITY: HELIX PHASING AND PREMELTING.BIOPOLYMERS. VOL. 73. ISSUE 3. P. 380 -403 | 117 | 86% | 42 |
| 2 | HARAN, TE , MOHANTY, U , (2009) THE UNIQUE STRUCTURE OF A-TRACTS AND INTRINSIC DNA BENDING.QUARTERLY REVIEWS OF BIOPHYSICS. VOL. 42. ISSUE 1. P. 41-81 | 129 | 53% | 65 |
| 3 | BEVERIDGE, DL , CHEATHAM, TE , MEZEI, M , (2012) THE ABCS OF MOLECULAR DYNAMICS SIMULATIONS ON B-DNA, CIRCA 2012.JOURNAL OF BIOSCIENCES. VOL. 37. ISSUE 3. P. 379 -397 | 69 | 81% | 18 |
| 4 | BEVERIDGE, DL , DIXIT, SB , BYUN, KS , BARREIRO, G , THAYER, KM , PONOMAREV, S , (2004) MOLECULAR DYNAMICS OF DNA AND PROTEIN-DNA COMPLEXES: PROGRESS ON SEQUENCE EFFECTS, CONFORMATIONAL STABILITY, AXIS CURVATURE, AND STRUCTURAL BIOINFORMATICS.NUCLEIC ACIDS: CURVATURE AND DEFORMATION. VOL. 884. ISSUE . P. 13 -64 | 109 | 67% | 5 |
| 5 | HUD, NV , PLAVEC, J , (2003) A UNIFIED MODEL FOR THE ORIGIN OF DNA SEQUENCE-DIRECTED CURVATURE.BIOPOLYMERS. VOL. 69. ISSUE 1. P. 144-159 | 76 | 86% | 88 |
| 6 | MCCONNELL, KJ , BEVERIDGE, DL , (2001) MOLECULAR DYNAMICS SIMULATIONS OF B '-DNA: SEQUENCE EFFECTS ON A-TRACT-INDUCED BENDING AND FLEXIBILITY.JOURNAL OF MOLECULAR BIOLOGY. VOL. 314. ISSUE 1. P. 23 -40 | 76 | 87% | 48 |
| 7 | MA, N , VAN DER VAART, A , (2016) ANISOTROPY OF B-DNA GROOVE BENDING.JOURNAL OF THE AMERICAN CHEMICAL SOCIETY. VOL. 138. ISSUE 31. P. 9951 -9958 | 66 | 66% | 1 |
| 8 | DIXIT, SB , MEZEI, M , BEVERIDGE, DL , (2012) STUDIES OF BASE PAIR SEQUENCE EFFECTS ON DNA SOLVATION BASED ON ALL-ATOM MOLECULAR DYNAMICS SIMULATIONS.JOURNAL OF BIOSCIENCES. VOL. 37. ISSUE 3. P. 399-421 | 66 | 72% | 13 |
| 9 | HARTMANN, B , LAVERY, R , (1996) DNA STRUCTURAL FORMS.QUARTERLY REVIEWS OF BIOPHYSICS. VOL. 29. ISSUE 4. P. 309-368 | 149 | 46% | 74 |
| 10 | YOUNG, MA , RAVISHANKER, G , BEVERIDGE, DL , BERMAN, HM , (1995) ANALYSIS OF LOCAL HELIX BENDING IN CRYSTAL-STRUCTURES OF DNA OLIGONUCLEOTIDES AND DNA-PROTEIN COMPLEXES.BIOPHYSICAL JOURNAL. VOL. 68. ISSUE 6. P. 2454-2468 | 98 | 72% | 78 |
Classes with closest relation at Level 1 |