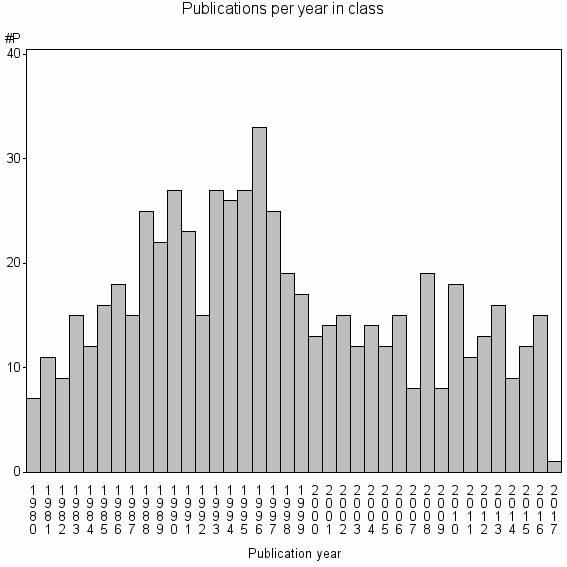
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 17163 | 614 | 32.3 | 63% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 478 | 3 | PHAGE DISPLAY//CELL FREE PROTEIN SYNTHESIS//ANTIBODY ENGINEERING | 23687 |
| 3617 | 2 | Q BETA REPLICASE//NICOTINE VACCINE//IMMUNOPHARMACOTHERAPY | 1239 |
| 17163 | 1 | Q BETA REPLICASE//RNA PHAGE//RNA BACTERIOPHAGES | 614 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | Q BETA REPLICASE | authKW | 914152 | 4% | 74% | 25 |
| 2 | RNA PHAGE | authKW | 501449 | 2% | 92% | 11 |
| 3 | RNA BACTERIOPHAGES | authKW | 414418 | 2% | 83% | 10 |
| 4 | LEVIVIRIDAE | authKW | 255756 | 1% | 86% | 6 |
| 5 | POLONIES | authKW | 255756 | 1% | 86% | 6 |
| 6 | MOLECULAR COLONIES | authKW | 248653 | 1% | 100% | 5 |
| 7 | SPONTANEOUS RNA SYNTHESIS | authKW | 248653 | 1% | 100% | 5 |
| 8 | BACTERIOPHAGE Q BETA | authKW | 248643 | 2% | 50% | 10 |
| 9 | Q BETA PHAGE | authKW | 223785 | 1% | 75% | 6 |
| 10 | MS2 VIRUS LIKE PARTICLES | authKW | 207209 | 1% | 83% | 5 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biochemistry & Molecular Biology | 2697 | 55% | 0% | 336 |
| 2 | Virology | 714 | 9% | 0% | 53 |
| 3 | Biophysics | 311 | 9% | 0% | 56 |
| 4 | Physics, Atomic, Molecular & Chemical | 82 | 6% | 0% | 36 |
| 5 | Biochemical Research Methods | 79 | 5% | 0% | 29 |
| 6 | Biotechnology & Applied Microbiology | 71 | 6% | 0% | 37 |
| 7 | Cell Biology | 61 | 7% | 0% | 44 |
| 8 | Chemistry, Physical | 59 | 10% | 0% | 60 |
| 9 | Mathematical & Computational Biology | 40 | 2% | 0% | 11 |
| 10 | Microbiology | 33 | 5% | 0% | 28 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | NETHERLANDS ELE ON NANOSCOPY | 99461 | 0% | 100% | 2 |
| 2 | IMMUNOASSAY MOL DIAG | 66306 | 0% | 67% | 2 |
| 3 | ASTBURY STRUCT MOL BIOL | 50659 | 5% | 3% | 32 |
| 4 | AGCY TECHNOL TRANSFER | 49731 | 0% | 100% | 1 |
| 5 | BIOTECHNOL BIOTHER IES TEAM | 49731 | 0% | 100% | 1 |
| 6 | CAMPUS TECHNOL FREIBURG CTF GMBH | 49731 | 0% | 100% | 1 |
| 7 | CLIN DIAGNOSTAFFILIATED HOSP | 49731 | 0% | 100% | 1 |
| 8 | EMERGING DIS INNOVAT THER IES IMETICEA F | 49731 | 0% | 100% | 1 |
| 9 | IFR 140 GENET FONCT AGRON SANTE MED | 49731 | 0% | 100% | 1 |
| 10 | KEY DISCIPLINE CLIN MED SHANDONG PROV LA | 49731 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | JOURNAL OF MOLECULAR BIOLOGY | 10137 | 11% | 0% | 69 |
| 2 | BERICHTE DER BUNSEN-GESELLSCHAFT-PHYSICAL CHEMISTRY CHEMICAL PHYSICS | 3510 | 3% | 0% | 18 |
| 3 | NUCLEIC ACIDS RESEARCH | 2068 | 7% | 0% | 41 |
| 4 | BIOPHYSICAL CHEMISTRY | 1672 | 2% | 0% | 12 |
| 5 | HERALD OF THE RUSSIAN ACADEMY OF SCIENCES | 1277 | 0% | 1% | 3 |
| 6 | TOXICOLOGY METHODS | 1006 | 0% | 1% | 2 |
| 7 | RNA | 949 | 1% | 0% | 7 |
| 8 | SEMINARS IN VIROLOGY | 880 | 0% | 1% | 2 |
| 9 | MOLECULAR BIOLOGY | 844 | 1% | 0% | 9 |
| 10 | PROGRESS IN NUCLEIC ACID RESEARCH AND MOLECULAR BIOLOGY | 835 | 0% | 1% | 3 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | Q BETA REPLICASE | 914152 | 4% | 74% | 25 | Search Q+BETA+REPLICASE | Search Q+BETA+REPLICASE |
| 2 | RNA PHAGE | 501449 | 2% | 92% | 11 | Search RNA+PHAGE | Search RNA+PHAGE |
| 3 | RNA BACTERIOPHAGES | 414418 | 2% | 83% | 10 | Search RNA+BACTERIOPHAGES | Search RNA+BACTERIOPHAGES |
| 4 | LEVIVIRIDAE | 255756 | 1% | 86% | 6 | Search LEVIVIRIDAE | Search LEVIVIRIDAE |
| 5 | POLONIES | 255756 | 1% | 86% | 6 | Search POLONIES | Search POLONIES |
| 6 | MOLECULAR COLONIES | 248653 | 1% | 100% | 5 | Search MOLECULAR+COLONIES | Search MOLECULAR+COLONIES |
| 7 | SPONTANEOUS RNA SYNTHESIS | 248653 | 1% | 100% | 5 | Search SPONTANEOUS+RNA+SYNTHESIS | Search SPONTANEOUS+RNA+SYNTHESIS |
| 8 | BACTERIOPHAGE Q BETA | 248643 | 2% | 50% | 10 | Search BACTERIOPHAGE+Q+BETA | Search BACTERIOPHAGE+Q+BETA |
| 9 | Q BETA PHAGE | 223785 | 1% | 75% | 6 | Search Q+BETA+PHAGE | Search Q+BETA+PHAGE |
| 10 | MS2 VIRUS LIKE PARTICLES | 207209 | 1% | 83% | 5 | Search MS2+VIRUS+LIKE+PARTICLES | Search MS2+VIRUS+LIKE+PARTICLES |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | PUMPENS, P , RENHOFA, R , DISHLERS, A , KOZLOVSKA, T , OSE, V , PUSHKO, P , TARS, K , GRENS, E , BACHMANN, MF , (2016) THE TRUE STORY AND ADVANTAGES OF RNA PHAGE CAPSIDS AS NANOTOOLS.INTERVIROLOGY. VOL. 59. ISSUE 2. P. 74 -110 | 105 | 34% | 0 |
| 2 | FU, Y , LI, JM , (2016) A NOVEL DELIVERY PLATFORM BASED ON BACTERIOPHAGE MS2 VIRUS-LIKE PARTICLES.VIRUS RESEARCH. VOL. 211. ISSUE . P. 9 -16 | 44 | 49% | 2 |
| 3 | BLECKLEY, S , SCHROEDER, SJ , (2012) INCORPORATING GLOBAL FEATURES OF RNA MOTIFS IN PREDICTIONS FOR AN ENSEMBLE OF SECONDARY STRUCTURES FOR ENCAPSIDATED MS2 BACTERIOPHAGE RNA.RNA. VOL. 18. ISSUE 7. P. 1309-1318 | 36 | 65% | 4 |
| 4 | ROLFSSON, O , MIDDLETON, S , MANFIELD, IW , WHITE, SJ , FAN, BC , VAUGHAN, R , RANSON, NA , DYKEMAN, E , TWAROCK, R , FORD, J , ET AL (2016) DIRECT EVIDENCE FOR PACKAGING SIGNAL-MEDIATED ASSEMBLY OF BACTERIOPHAGE MS2.JOURNAL OF MOLECULAR BIOLOGY. VOL. 428. ISSUE 2. P. 431 -448 | 34 | 47% | 5 |
| 5 | CHETVERIN, AB , (2011) PARADOXES OF REPLICATION OF RNA OF A BACTERIAL VIRUS.MOLECULAR BIOLOGY. VOL. 45. ISSUE 1. P. 139-149 | 26 | 90% | 2 |
| 6 | RUMNIEKS, J , TARS, K , (2014) CRYSTAL STRUCTURE OF THE BACTERIOPHAGE Q BETA COAT PROTEIN IN COMPLEX WITH THE RNA OPERATOR OF THE REPLICASE GENE.JOURNAL OF MOLECULAR BIOLOGY. VOL. 426. ISSUE 5. P. 1039-1049 | 24 | 80% | 5 |
| 7 | STOCKLEY, PG , RANSON, NA , TWAROCK, R , (2013) A NEW PARADIGM FOR THE ROLES OF THE GENOME IN SSRNA VIRUSES.FUTURE VIROLOGY. VOL. 8. ISSUE 6. P. 531 -543 | 40 | 50% | 2 |
| 8 | PERSSON, M , TARS, K , LILJAS, L , (2013) PRR1 COAT PROTEIN BINDING TO ITS RNA TRANSLATIONAL OPERATOR.ACTA CRYSTALLOGRAPHICA SECTION D-BIOLOGICAL CRYSTALLOGRAPHY. VOL. 69. ISSUE . P. 367-372 | 22 | 88% | 2 |
| 9 | RUMNIEKS, J , OSE, V , TARS, K , DISLERS, A , STRODS, A , CIELENS, I , RENHOFA, R , (2009) ASSEMBLY OF MIXED ROD-LIKE AND SPHERICAL PARTICLES FROM GROUP I AND II RNA BACTERIOPHAGE COAT PROTEINS.VIROLOGY. VOL. 391. ISSUE 2. P. 187-194 | 24 | 86% | 3 |
| 10 | CHETVERIN, AB , (2004) REPLICABLE AND RECOMBINOGENIC RNAS.FEBS LETTERS. VOL. 567. ISSUE 1. P. 35-41 | 32 | 73% | 10 |
Classes with closest relation at Level 1 |