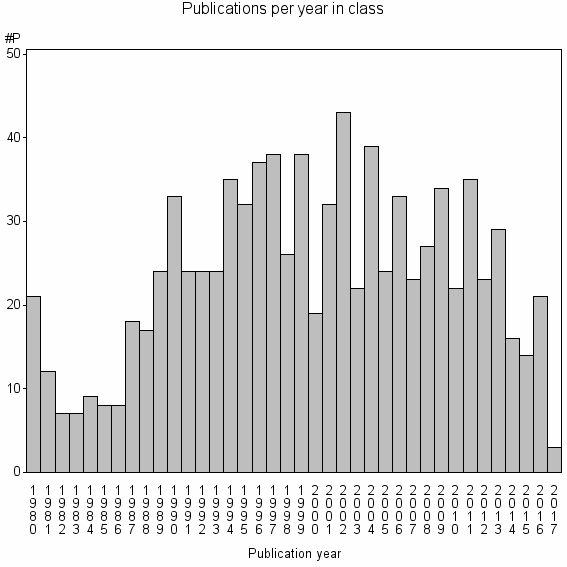
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 12501 | 901 | 50.9 | 85% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 71 | 3 | GENETICS & HEREDITY//CHROMATIN//CELL BIOLOGY | 83302 |
| 225 | 2 | YEAST//SACCHAROMYCES CEREVISIAE//SCHIZOSACCHAROMYCES POMBE | 20517 |
| 12501 | 1 | SIR COMPLEX//SIR PROTEINS//SIR3 | 901 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | SIR COMPLEX | authKW | 372779 | 1% | 100% | 11 |
| 2 | SIR PROTEINS | authKW | 361472 | 2% | 67% | 16 |
| 3 | SIR3 | authKW | 332104 | 2% | 70% | 14 |
| 4 | ABF1 | authKW | 228747 | 1% | 75% | 9 |
| 5 | SILENCING | authKW | 180534 | 6% | 10% | 51 |
| 6 | DONOR PREFERENCE | authKW | 169445 | 1% | 100% | 5 |
| 7 | SIR4 | authKW | 150955 | 1% | 64% | 7 |
| 8 | UNIT CHROMATIN TRANSCRIPT | address | 144586 | 1% | 53% | 8 |
| 9 | RECOMBINATION ENHANCER | authKW | 135556 | 0% | 100% | 4 |
| 10 | SIR2 | authKW | 122389 | 3% | 14% | 25 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Cell Biology | 7335 | 46% | 0% | 415 |
| 2 | Biochemistry & Molecular Biology | 5429 | 63% | 0% | 569 |
| 3 | Genetics & Heredity | 4633 | 30% | 0% | 273 |
| 4 | Mycology | 726 | 4% | 0% | 34 |
| 5 | Developmental Biology | 632 | 6% | 0% | 51 |
| 6 | Biology | 159 | 4% | 0% | 40 |
| 7 | Microbiology | 129 | 6% | 0% | 58 |
| 8 | Biophysics | 81 | 5% | 0% | 41 |
| 9 | Biotechnology & Applied Microbiology | 53 | 5% | 0% | 43 |
| 10 | Biochemical Research Methods | 14 | 2% | 0% | 21 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | UNIT CHROMATIN TRANSCRIPT | 144586 | 1% | 53% | 8 |
| 2 | FDN IST PASTEUR | 35293 | 1% | 21% | 5 |
| 3 | BIOL ROSENSTIEL | 33889 | 0% | 100% | 1 |
| 4 | BIOL SCIMOO CANC | 33889 | 0% | 100% | 1 |
| 5 | BIOL STRUCT RADIOBIOLCNRSURA2096 | 33889 | 0% | 100% | 1 |
| 6 | BIOTECHNOL UNIL BIOTECHNOL MOL UNIL | 33889 | 0% | 100% | 1 |
| 7 | BIOTECHNOLISPFSB MOL BIOTECHNOL | 33889 | 0% | 100% | 1 |
| 8 | CNRS ENSSL 5665 | 33889 | 0% | 100% | 1 |
| 9 | CNRS UNITE MIXTE RECH BIOL MOL CELLULAIRE | 33889 | 0% | 100% | 1 |
| 10 | DSV DBCM SERV BIOCHEM GENET MOL | 33889 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | GENETICS | 30962 | 12% | 1% | 108 |
| 2 | MOLECULAR AND CELLULAR BIOLOGY | 23814 | 14% | 1% | 125 |
| 3 | YEAST | 8044 | 3% | 1% | 26 |
| 4 | GENES & DEVELOPMENT | 7991 | 5% | 1% | 42 |
| 5 | COLD SPRING HARBOR SYMPOSIA ON QUANTITATIVE BIOLOGY | 4394 | 2% | 1% | 17 |
| 6 | CELL | 3400 | 4% | 0% | 39 |
| 7 | MOLECULAR CELL | 1985 | 2% | 0% | 18 |
| 8 | EMBO JOURNAL | 1861 | 3% | 0% | 31 |
| 9 | CURRENT GENETICS | 1783 | 2% | 0% | 14 |
| 10 | NUCLEIC ACIDS RESEARCH | 1384 | 5% | 0% | 41 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | SIR COMPLEX | 372779 | 1% | 100% | 11 | Search SIR+COMPLEX | Search SIR+COMPLEX |
| 2 | SIR PROTEINS | 361472 | 2% | 67% | 16 | Search SIR+PROTEINS | Search SIR+PROTEINS |
| 3 | SIR3 | 332104 | 2% | 70% | 14 | Search SIR3 | Search SIR3 |
| 4 | ABF1 | 228747 | 1% | 75% | 9 | Search ABF1 | Search ABF1 |
| 5 | SILENCING | 180534 | 6% | 10% | 51 | Search SILENCING | Search SILENCING |
| 6 | DONOR PREFERENCE | 169445 | 1% | 100% | 5 | Search DONOR+PREFERENCE | Search DONOR+PREFERENCE |
| 7 | SIR4 | 150955 | 1% | 64% | 7 | Search SIR4 | Search SIR4 |
| 8 | RECOMBINATION ENHANCER | 135556 | 0% | 100% | 4 | Search RECOMBINATION+ENHANCER | Search RECOMBINATION+ENHANCER |
| 9 | SIR2 | 122389 | 3% | 14% | 25 | Search SIR2 | Search SIR2 |
| 10 | YEAST MATING TYPE | 76249 | 0% | 75% | 3 | Search YEAST+MATING+TYPE | Search YEAST+MATING+TYPE |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | GARTENBERG, MR , SMITH, JS , (2016) THE NUTS AND BOLTS OF TRANSCRIPTIONALLY SILENT CHROMATIN IN SACCHAROMYCES CEREVISIAE.GENETICS. VOL. 203. ISSUE 4. P. 1563 -+ | 247 | 59% | 0 |
| 2 | KUENG, S , OPPIKOFER, M , GASSER, SM , (2013) SIR PROTEINS AND THE ASSEMBLY OF SILENT CHROMATIN IN BUDDING YEAST.ANNUAL REVIEW OF GENETICS, VOL 47. VOL. 47. ISSUE . P. 275-+ | 169 | 69% | 30 |
| 3 | GRUNSTEIN, M , GASSER, SM , (2013) EPIGENETICS IN SACCHAROMYCES CEREVISIAE.COLD SPRING HARBOR PERSPECTIVES IN BIOLOGY. VOL. 5. ISSUE 7. P. - | 97 | 71% | 15 |
| 4 | RUSCHE, LN , KIRCHMAIER, AL , RINE, J , (2003) THE ESTABLISHMENT, INHERITANCE, AND FUNCTION OF SILENCED CHROMATIN IN SACCHAROMYCES CEREVISIAE.ANNUAL REVIEW OF BIOCHEMISTRY. VOL. 72. ISSUE . P. 481-516 | 117 | 50% | 480 |
| 5 | HABER, JE , (2012) MATING-TYPE GENES AND MAT SWITCHING IN SACCHAROMYCES CEREVISIAE.GENETICS. VOL. 191. ISSUE 1. P. 33-64 | 125 | 41% | 105 |
| 6 | ARMACHE, KJ , CANZIO, D , NARLIKAR, GJ , KINGSTON, RE , GARLICK, JD , (2011) STRUCTURAL BASIS OF SILENCING: SIR3 BAH DOMAIN IN COMPLEX WITH A NUCLEOSOME AT 3.0 ANGSTROM RESOLUTION.SCIENCE. VOL. 334. ISSUE 6058. P. 977 -982 | 48 | 92% | 89 |
| 7 | OPPIKOFER, M , KUENG, S , GASSER, SM , (2013) SIR-NUCLEOSOME INTERACTIONS: STRUCTURE-FUNCTION RELATIONSHIPS IN YEAST SILENT CHROMATIN.GENE. VOL. 527. ISSUE 1. P. 10-25 | 115 | 43% | 10 |
| 8 | KUENG, S , TSAI-PFLUGFELDER, M , OPPIKOFER, M , FERREIRA, HC , ROBERTS, E , TSAI, C , ROLOFF, TC , SACK, R , GASSER, SM , (2012) REGULATING REPRESSION: ROLES FOR THE SIR4 N-TERMINUS IN LINKER DNA PROTECTION AND STABILIZATION OF EPIGENETIC STATES.PLOS GENETICS. VOL. 8. ISSUE 5. P. - | 71 | 72% | 4 |
| 9 | LARIN, ML , HARDING, K , WILLIAMS, EC , LIANGA, N , DORE, C , PILON, S , LANGIS, E , YANOFSKY, C , RUDNER, AD , (2015) COMPETITION BETWEEN HETEROCHROMATIC LOCI ALLOWS THE ABUNDANCE OF THE SILENCING PROTEIN, SIR4, TO REGULATE DE NOVO ASSEMBLY OF HETEROCHROMATIN.PLOS GENETICS. VOL. 11. ISSUE 11. P. - | 66 | 65% | 2 |
| 10 | YOUNG, TJ , KIRCHMAIER, AL , (2012) CELL CYCLE REGULATION OF SILENT CHROMATIN FORMATION.BIOCHIMICA ET BIOPHYSICA ACTA-GENE REGULATORY MECHANISMS. VOL. 1819. ISSUE 3-4. P. 303-312 | 72 | 57% | 4 |
Classes with closest relation at Level 1 |
| Rank | Class id | link |
|---|---|---|
| 1 | 1312 | TELOMERE//TELOMERASE//TRF2 |
| 2 | 14125 | USP22//SET1//UBIQUITIN SPECIFIC PROTEASE 22 |
| 3 | 11310 | GAL80//GAL4P//GAL4 |
| 4 | 17572 | TY1//SECT EUKARYOT TRANSPOSABLE ELEMENTS//TY ELEMENTS |
| 5 | 5093 | HISTONE CHAPERONE//HISTONE VARIANTS//ASF1 |
| 6 | 16690 | YEAST//CHROMOSOME XIV//CHROMOSOME XV |
| 7 | 13617 | 2 MU M PLASMID//2 MU M CIRCLE//YEP VECTOR |
| 8 | 18452 | IME1//IME2//PROSPORE MEMBRANE |
| 9 | 3273 | NUCLEOSOME POSITIONING//ISWI//NUCLEOSOME |
| 10 | 7575 | HP1//POSITION EFFECT VARIEGATION//HETEROCHROMATIN |