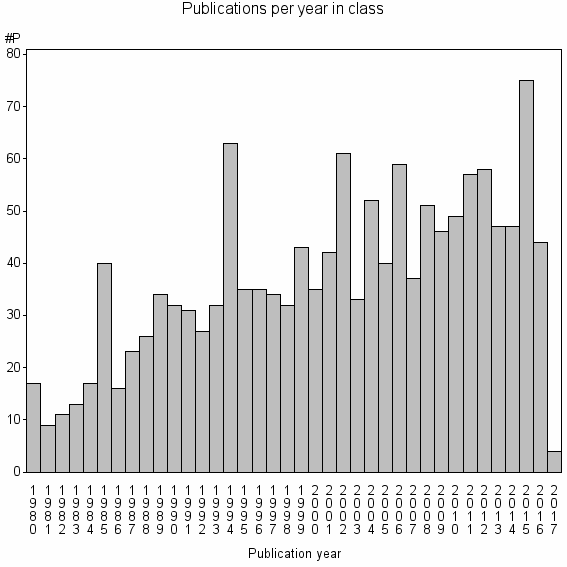
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 7082 | 1407 | 33.7 | 63% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 142 | 3 | DNA REPAIR//INTERNATIONAL JOURNAL OF RADIATION BIOLOGY//RADIATION RESEARCH | 64602 |
| 271 | 2 | INTERNATIONAL JOURNAL OF RADIATION BIOLOGY//RADIATION RESEARCH//ATAXIA TELANGIECTASIA | 19288 |
| 7082 | 1 | CLUSTERED DNA DAMAGE//NANODOSIMETRY//GEANT4 DNA | 1407 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | CLUSTERED DNA DAMAGE | authKW | 451984 | 2% | 77% | 27 |
| 2 | NANODOSIMETRY | authKW | 431014 | 2% | 83% | 24 |
| 3 | GEANT4 DNA | authKW | 407478 | 2% | 72% | 26 |
| 4 | RADIAT BIOPHYS GRP | address | 393455 | 2% | 58% | 31 |
| 5 | RADIAT GENOME STABIL UNIT | address | 287061 | 4% | 24% | 56 |
| 6 | TRACK STRUCTURE | authKW | 225965 | 2% | 39% | 27 |
| 7 | ULTRASOFT X RAYS | authKW | 223205 | 1% | 86% | 12 |
| 8 | CLUSTERED DAMAGE | authKW | 156236 | 1% | 60% | 12 |
| 9 | ADV SPACE STUDIES | address | 138882 | 1% | 80% | 8 |
| 10 | RADIATION RESEARCH | journal | 136345 | 14% | 3% | 203 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Radiology, Nuclear Medicine & Medical Imaging | 33586 | 59% | 0% | 834 |
| 2 | Nuclear Science & Technology | 30202 | 43% | 0% | 604 |
| 3 | Biology | 19218 | 33% | 0% | 466 |
| 4 | Biophysics | 4460 | 21% | 0% | 295 |
| 5 | Environmental Sciences | 2698 | 22% | 0% | 313 |
| 6 | Public, Environmental & Occupational Health | 2143 | 17% | 0% | 238 |
| 7 | Physics, Atomic, Molecular & Chemical | 1155 | 12% | 0% | 174 |
| 8 | Instruments & Instrumentation | 757 | 8% | 0% | 113 |
| 9 | Engineering, Biomedical | 719 | 6% | 0% | 87 |
| 10 | Physics, Nuclear | 703 | 7% | 0% | 104 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | RADIAT BIOPHYS GRP | 393455 | 2% | 58% | 31 |
| 2 | RADIAT GENOME STABIL UNIT | 287061 | 4% | 24% | 56 |
| 3 | ADV SPACE STUDIES | 138882 | 1% | 80% | 8 |
| 4 | THOMAS HARRIOT ARTS SCI | 114938 | 1% | 38% | 14 |
| 5 | PL NUCL ENERGY | 99197 | 1% | 57% | 8 |
| 6 | PHYS RAYONNEMENTS PLICAT | 90418 | 0% | 83% | 5 |
| 7 | FUNDAMENTALS DOSIMETRY | 86803 | 0% | 100% | 4 |
| 8 | CENBG | 61748 | 2% | 11% | 27 |
| 9 | RADIAT RISK ANAL | 45204 | 0% | 42% | 5 |
| 10 | DRPH SDE | 43402 | 0% | 100% | 2 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | RADIATION RESEARCH | 136345 | 14% | 3% | 203 |
| 2 | RADIATION PROTECTION DOSIMETRY | 104739 | 16% | 2% | 220 |
| 3 | INTERNATIONAL JOURNAL OF RADIATION BIOLOGY | 96703 | 11% | 3% | 150 |
| 4 | RADIATION AND ENVIRONMENTAL BIOPHYSICS | 78052 | 5% | 5% | 73 |
| 5 | PHYSICS IN MEDICINE AND BIOLOGY | 13985 | 6% | 1% | 84 |
| 6 | JOURNAL OF RADIATION RESEARCH | 6026 | 2% | 1% | 24 |
| 7 | ADVANCES IN RADIATION BIOLOGY | 5418 | 0% | 6% | 4 |
| 8 | RADIATION PHYSICS AND CHEMISTRY | 5137 | 3% | 1% | 48 |
| 9 | NUCLEAR INSTRUMENTS & METHODS IN PHYSICS RESEARCH SECTION B-BEAM INTERACTIONS WITH MATERIALS AND ATOMS | 4397 | 6% | 0% | 81 |
| 10 | DNA REPAIR | 1780 | 1% | 1% | 13 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | CLUSTERED DNA DAMAGE | 451984 | 2% | 77% | 27 | Search CLUSTERED+DNA+DAMAGE | Search CLUSTERED+DNA+DAMAGE |
| 2 | NANODOSIMETRY | 431014 | 2% | 83% | 24 | Search NANODOSIMETRY | Search NANODOSIMETRY |
| 3 | GEANT4 DNA | 407478 | 2% | 72% | 26 | Search GEANT4+DNA | Search GEANT4+DNA |
| 4 | TRACK STRUCTURE | 225965 | 2% | 39% | 27 | Search TRACK+STRUCTURE | Search TRACK+STRUCTURE |
| 5 | ULTRASOFT X RAYS | 223205 | 1% | 86% | 12 | Search ULTRASOFT+X+RAYS | Search ULTRASOFT+X+RAYS |
| 6 | CLUSTERED DAMAGE | 156236 | 1% | 60% | 12 | Search CLUSTERED+DAMAGE | Search CLUSTERED+DAMAGE |
| 7 | MULTIPLY DAMAGED SITES | 111602 | 0% | 86% | 6 | Search MULTIPLY+DAMAGED+SITES | Search MULTIPLY+DAMAGED+SITES |
| 8 | MICRODOSIMETRY | 91496 | 2% | 12% | 34 | Search MICRODOSIMETRY | Search MICRODOSIMETRY |
| 9 | IONIZING RADIATION DNA DAMAGE | 86803 | 0% | 100% | 4 | Search IONIZING+RADIATION+DNA+DAMAGE | Search IONIZING+RADIATION+DNA+DAMAGE |
| 10 | DIELECTRIC THEORY | 86791 | 1% | 40% | 10 | Search DIELECTRIC+THEORY | Search DIELECTRIC+THEORY |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | GEORGAKILAS, AG , O'NEILL, P , STEWART, RD , (2013) INDUCTION AND REPAIR OF CLUSTERED DNA LESIONS: WHAT DO WE KNOW SO FAR?.RADIATION RESEARCH. VOL. 180. ISSUE 1. P. 100-109 | 74 | 73% | 48 |
| 2 | NIKITAKI, Z , NIKOLOV, V , MAVRAGANI, IV , PLANTE, I , EMFIETZOGLOU, D , ILIAKIS, G , GEORGAKILAS, AG , (2016) NON-DSB CLUSTERED DNA LESIONS. DOES THEORY COLOCALIZE WITH THE EXPERIMENT?.RADIATION PHYSICS AND CHEMISTRY. VOL. 128. ISSUE . P. 26 -35 | 59 | 66% | 1 |
| 3 | SEMSARHA, F , RAISALI, G , GOLIAEI, B , KHALAFI, H , (2016) MICRODOSIMETRY OF DNA CONFORMATIONS: RELATION BETWEEN DIRECT EFFECT OF CO-60 GAMMA RAYS AND TOPOLOGY OF DNA GEOMETRICAL MODELS IN THE CALCULATION OF A-, B- AND Z-DNA RADIATION-INDUCED DAMAGE YIELDS.RADIATION AND ENVIRONMENTAL BIOPHYSICS. VOL. 55. ISSUE 2. P. 243 -254 | 50 | 81% | 0 |
| 4 | BUENO, M , SCHULTE, R , MEYLAN, S , VILLAGRASA, C , (2015) INFLUENCE OF THE GEOMETRICAL DETAIL IN THE DESCRIPTION OF DNA AND THE SCORING METHOD OF IONIZATION CLUSTERING ON NANODOSIMETRIC PARAMETERS OF TRACK STRUCTURE: A MONTE CARLO STUDY USING GEANT4-DNA.PHYSICS IN MEDICINE AND BIOLOGY. VOL. 60. ISSUE 21. P. 8583 -8599 | 43 | 91% | 1 |
| 5 | TAJIK, M , ROZATIAN, ASH , SEMSARHA, F , (2015) SIMULATION OF ULTRASOFT X-RAYS INDUCED DNA DAMAGE USING THE GEANT4 MONTE CARLO TOOLKIT.NUCLEAR INSTRUMENTS & METHODS IN PHYSICS RESEARCH SECTION B-BEAM INTERACTIONS WITH MATERIALS AND ATOMS. VOL. 342. ISSUE . P. 258 -265 | 49 | 74% | 3 |
| 6 | TAJIK, M , ROZATIAN, ASH , SEMSARHA, F , (2015) CALCULATION OF DIRECT EFFECTS OF CO-60 GAMMA RAYS ON THE DIFFERENT DNA STRUCTURAL LEVELS: A SIMULATION STUDY USING THE GEANT4-DNA TOOLKIT.NUCLEAR INSTRUMENTS & METHODS IN PHYSICS RESEARCH SECTION B-BEAM INTERACTIONS WITH MATERIALS AND ATOMS. VOL. 346. ISSUE . P. 53 -60 | 45 | 82% | 2 |
| 7 | ECCLES, LJ , O'NEILL, P , LOMAX, ME , (2011) DELAYED REPAIR OF RADIATION INDUCED CLUSTERED DNA DAMAGE: FRIEND OR FOE?.MUTATION RESEARCH-FUNDAMENTAL AND MOLECULAR MECHANISMS OF MUTAGENESIS. VOL. 711. ISSUE 1-2. P. 134 -141 | 49 | 69% | 65 |
| 8 | MAGNANDER, K , ELMROTH, K , (2012) BIOLOGICAL CONSEQUENCES OF FORMATION AND REPAIR OF COMPLEX DNA DAMAGE.CANCER LETTERS. VOL. 327. ISSUE 1-2. P. 90-96 | 64 | 60% | 13 |
| 9 | SHIKAZONO, N , NOGUCHI, M , FUJII, K , URUSHIBARA, A , YOKOYA, A , (2009) THE YIELD, PROCESSING, AND BIOLOGICAL CONSEQUENCES OF CLUSTERED DNA DAMAGE INDUCED BY IONIZING RADIATION.JOURNAL OF RADIATION RESEARCH. VOL. 50. ISSUE 1. P. 27-36 | 54 | 70% | 54 |
| 10 | KYRIAKOU, I , SEFL, M , NOURRY, V , INCERTI, S , (2016) THE IMPACT OF NEW GEANT4-DNA CROSS SECTION MODELS ON ELECTRON TRACK STRUCTURE SIMULATIONS IN LIQUID WATER.JOURNAL OF APPLIED PHYSICS. VOL. 119. ISSUE 19. P. - | 42 | 74% | 1 |
Classes with closest relation at Level 1 |