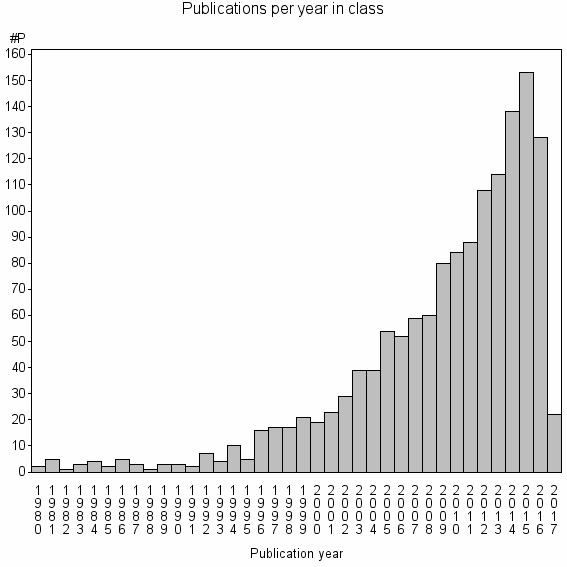
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 6978 | 1420 | 31.7 | 62% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 10 | 4 | OPTICS//PHYSICS, PARTICLES & FIELDS//PHYSICS, MULTIDISCIPLINARY | 1131262 |
| 359 | 3 | JOURNAL OF BIOMEDICAL OPTICS//BIOMEDICAL OPTICS EXPRESS//OPTICS | 34746 |
| 1278 | 2 | NANOBIOPHOTON//SUPER RESOLUTION MICROSCOPY//HIGH CONTENT SCREENING | 8720 |
| 6978 | 1 | CELL SEGMENTATION//HISTOPATHOLOGICAL IMAGE ANALYSIS//NUCLEI SEGMENTATION | 1420 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | CELL SEGMENTATION | authKW | 563962 | 3% | 66% | 40 |
| 2 | HISTOPATHOLOGICAL IMAGE ANALYSIS | authKW | 494548 | 2% | 100% | 23 |
| 3 | NUCLEI SEGMENTATION | authKW | 250198 | 1% | 73% | 16 |
| 4 | TISSUE COUNTER ANALYSIS | authKW | 165396 | 1% | 77% | 10 |
| 5 | NUCLEAR SEGMENTATION | authKW | 145135 | 1% | 75% | 9 |
| 6 | ANALYT MORPHOL DERMATOL | address | 144533 | 1% | 61% | 11 |
| 7 | CLUMP SPLITTING | authKW | 137610 | 1% | 80% | 8 |
| 8 | MITOSIS DETECTION | authKW | 110581 | 0% | 86% | 6 |
| 9 | DIGITAL PATHOLOGY | authKW | 110216 | 2% | 18% | 29 |
| 10 | BREAST CANCER GRADING | authKW | 107511 | 0% | 100% | 5 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Engineering, Biomedical | 9969 | 21% | 0% | 302 |
| 2 | Microscopy | 8476 | 8% | 0% | 120 |
| 3 | Computer Science, Artificial Intelligence | 7599 | 19% | 0% | 276 |
| 4 | Mathematical & Computational Biology | 4695 | 11% | 0% | 153 |
| 5 | Imaging Science & Photographic Technology | 4101 | 8% | 0% | 110 |
| 6 | Computer Science, Interdisciplinary Applications | 4029 | 15% | 0% | 210 |
| 7 | Medical Informatics | 3368 | 6% | 0% | 87 |
| 8 | Biochemical Research Methods | 2873 | 16% | 0% | 222 |
| 9 | Radiology, Nuclear Medicine & Medical Imaging | 1384 | 13% | 0% | 186 |
| 10 | Cell Biology | 1172 | 16% | 0% | 231 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | ANALYT MORPHOL DERMATOL | 144533 | 1% | 61% | 11 |
| 2 | JIANGSU BIG DATA ANAL TECH | 107511 | 0% | 100% | 5 |
| 3 | HCNR BIOINFORMAT | 91735 | 1% | 53% | 8 |
| 4 | NEUROIMMUNOL MOL PATTERN RECOGNIT GRP | 64506 | 0% | 100% | 3 |
| 5 | HARVARD NEURODEGENERAT REPAIR | 60490 | 1% | 26% | 11 |
| 6 | BIOMED COMP VIS GRP | 48853 | 1% | 23% | 10 |
| 7 | CELL BIOL GENOMES GRP | 48378 | 0% | 75% | 3 |
| 8 | OERSTED DTU AUTOMAT | 48378 | 0% | 75% | 3 |
| 9 | BIOSPECIMENS TUMOR BANK | 43004 | 0% | 100% | 2 |
| 10 | CONCEPT OPTIMIZAT MODELLING SYST | 43004 | 0% | 100% | 2 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | CYTOMETRY PART A | 57856 | 4% | 4% | 63 |
| 2 | MEDICAL IMAGE ANALYSIS | 30395 | 3% | 4% | 40 |
| 3 | ANALYTICAL AND QUANTITATIVE CYTOLOGY AND HISTOLOGY | 26267 | 3% | 3% | 47 |
| 4 | JOURNAL OF MICROSCOPY | 22807 | 4% | 2% | 52 |
| 5 | IEEE TRANSACTIONS ON MEDICAL IMAGING | 21013 | 4% | 2% | 61 |
| 6 | COMPUTERIZED MEDICAL IMAGING AND GRAPHICS | 11995 | 2% | 2% | 31 |
| 7 | ANALYTICAL CELLULAR PATHOLOGY | 9498 | 1% | 3% | 17 |
| 8 | IEEE TRANSACTIONS ON BIOMEDICAL ENGINEERING | 6308 | 3% | 1% | 48 |
| 9 | COMPUTERS IN BIOLOGY AND MEDICINE | 4945 | 2% | 1% | 26 |
| 10 | PATTERN RECOGNITION | 4720 | 3% | 1% | 39 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | CELL SEGMENTATION | 563962 | 3% | 66% | 40 | Search CELL+SEGMENTATION | Search CELL+SEGMENTATION |
| 2 | HISTOPATHOLOGICAL IMAGE ANALYSIS | 494548 | 2% | 100% | 23 | Search HISTOPATHOLOGICAL+IMAGE+ANALYSIS | Search HISTOPATHOLOGICAL+IMAGE+ANALYSIS |
| 3 | NUCLEI SEGMENTATION | 250198 | 1% | 73% | 16 | Search NUCLEI+SEGMENTATION | Search NUCLEI+SEGMENTATION |
| 4 | TISSUE COUNTER ANALYSIS | 165396 | 1% | 77% | 10 | Search TISSUE+COUNTER+ANALYSIS | Search TISSUE+COUNTER+ANALYSIS |
| 5 | NUCLEAR SEGMENTATION | 145135 | 1% | 75% | 9 | Search NUCLEAR+SEGMENTATION | Search NUCLEAR+SEGMENTATION |
| 6 | CLUMP SPLITTING | 137610 | 1% | 80% | 8 | Search CLUMP+SPLITTING | Search CLUMP+SPLITTING |
| 7 | MITOSIS DETECTION | 110581 | 0% | 86% | 6 | Search MITOSIS+DETECTION | Search MITOSIS+DETECTION |
| 8 | DIGITAL PATHOLOGY | 110216 | 2% | 18% | 29 | Search DIGITAL+PATHOLOGY | Search DIGITAL+PATHOLOGY |
| 9 | BREAST CANCER GRADING | 107511 | 0% | 100% | 5 | Search BREAST+CANCER+GRADING | Search BREAST+CANCER+GRADING |
| 10 | PAP SMEAR IMAGES | 107511 | 0% | 100% | 5 | Search PAP+SMEAR+IMAGES | Search PAP+SMEAR+IMAGES |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | CHEN, C , WANG, W , OZOLEK, JA , ROHDE, GK , (2013) A FLEXIBLE AND ROBUST APPROACH FOR SEGMENTING CELL NUCLEI FROM 2D MICROSCOPY IMAGES USING SUPERVISED LEARNING AND TEMPLATE MATCHING.CYTOMETRY PART A. VOL. 83A. ISSUE 5. P. 495 -507 | 42 | 86% | 17 |
| 2 | ARSLAN, S , ERSAHIN, T , CETIN-ATALAY, R , GUNDUZ-DEMIR, C , (2013) ATTRIBUTED RELATIONAL GRAPHS FOR CELL NUCLEUS SEGMENTATION IN FLUORESCENCE MICROSCOPY IMAGES.IEEE TRANSACTIONS ON MEDICAL IMAGING. VOL. 32. ISSUE 6. P. 1121 -1131 | 28 | 88% | 8 |
| 3 | MATULA, P , MASKA, M , SOROKIN, DV , MATULA, P , ORTIZ-DE-SOLORZANO, C , KOZUBEK, M , (2015) CELL TRACKING ACCURACY MEASUREMENT BASED ON COMPARISON OF ACYCLIC ORIENTED GRAPHS.PLOS ONE. VOL. 10. ISSUE 12. P. - | 24 | 92% | 1 |
| 4 | XU, J , XIANG, L , LIU, QS , GILMORE, H , WU, JZ , TANG, JH , MADABHUSHI, A , (2016) STACKED SPARSE AUTOENCODER (SSAE) FOR NUCLEI DETECTION ON BREAST CANCER HISTOPATHOLOGY IMAGES.IEEE TRANSACTIONS ON MEDICAL IMAGING. VOL. 35. ISSUE 1. P. 119 -130 | 23 | 74% | 4 |
| 5 | LU, C , XU, HM , XU, J , GILMORE, H , MANDAL, M , MADABHUSHI, A , (2016) MULTI-PASS ADAPTIVE VOTING FOR NUCLEI DETECTION IN HISTOPATHOLOGICAL IMAGES.SCIENTIFIC REPORTS. VOL. 6. ISSUE . P. - | 26 | 84% | 0 |
| 6 | MEIJERING, E , SMAL, I , VAN CAPPELLEN, WA , DZYUBACHYK, O , (2009) TRACKING IN CELL AND DEVELOPMENTAL BIOLOGY.SEMINARS IN CELL & DEVELOPMENTAL BIOLOGY. VOL. 20. ISSUE 8. P. 894 -902 | 42 | 49% | 86 |
| 7 | KOYUNCU, CF , AKHAN, E , ERSAHIN, T , CETIN-ATALAY, R , GUNDUZ-DEMIR, C , (2016) ITERATIVE H-MINIMA-BASED MARKER-CONTROLLED WATERSHED FOR CELL NUCLEUS SEGMENTATION.CYTOMETRY PART A. VOL. 89A. ISSUE 4. P. 338 -349 | 21 | 100% | 0 |
| 8 | MASKA, M , ULMAN, V , SVOBODA, D , MATULA, P , MATULA, P , EDERRA, C , URBIOLA, A , ESPANA, T , VENKATESAN, S , BALAK, DMW , ET AL (2014) A BENCHMARK FOR COMPARISON OF CELL TRACKING ALGORITHMS.BIOINFORMATICS. VOL. 30. ISSUE 11. P. 1609-1617 | 19 | 79% | 48 |
| 9 | MEIJERING, E , DZYUBACHYK, O , SMAL, I , (2012) METHODS FOR CELL AND PARTICLE TRACKING.IMAGING AND SPECTROSCOPIC ANALYSIS OF LIVING CELLS: OPTICAL AND SPECTROSCOPIC TECHNIQUES. VOL. 504. ISSUE . P. 183 -200 | 20 | 53% | 330 |
| 10 | BARKER, J , HOOGI, A , DEPEURSINGE, A , RUBIN, DL , (2016) AUTOMATED CLASSIFICATION OF BRAIN TUMOR TYPE IN WHOLE-SLIDE DIGITAL PATHOLOGY IMAGES USING LOCAL REPRESENTATIVE TILES.MEDICAL IMAGE ANALYSIS. VOL. 30. ISSUE . P. 60 -71 | 30 | 59% | 1 |
Classes with closest relation at Level 1 |