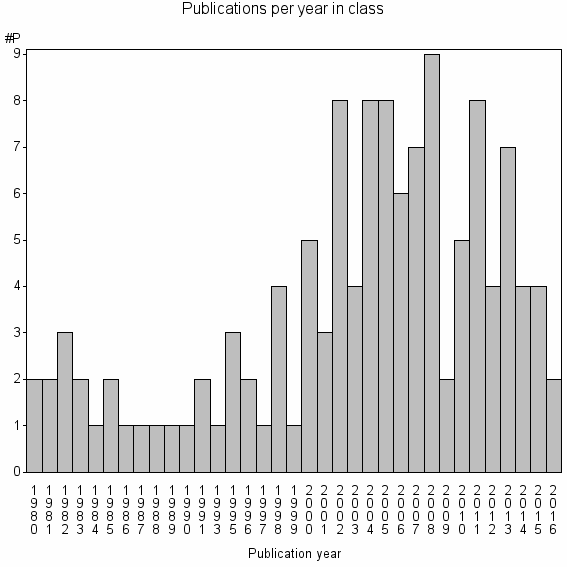
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 33386 | 125 | 49.3 | 75% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 116 | 3 | MUTATION RESEARCH//TOXICOLOGY//CARCINOGENESIS | 70215 |
| 2528 | 2 | CARCINOGENESIS//CELL TRANSFORMATION ASSAY//BHAS 42 CELLS | 3810 |
| 33386 | 1 | COMPETITIVE TEMPLATE//GIANT CANCER CELLS//RAJU CELLS | 125 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | COMPETITIVE TEMPLATE | authKW | 488569 | 2% | 100% | 2 |
| 2 | GIANT CANCER CELLS | authKW | 488569 | 2% | 100% | 2 |
| 3 | RAJU CELLS | authKW | 488569 | 2% | 100% | 2 |
| 4 | NEOSIS | authKW | 488565 | 3% | 50% | 4 |
| 5 | DEPOLYPLOIDIZATION | authKW | 325711 | 2% | 67% | 2 |
| 6 | AMITOSIS NUCLEAR FRAGMENTATION | authKW | 244285 | 1% | 100% | 1 |
| 7 | ANOMALOUS MITOSES | authKW | 244285 | 1% | 100% | 1 |
| 8 | C MYC INTERACTING GENES | authKW | 244285 | 1% | 100% | 1 |
| 9 | CARBONYL METABOLISING ENZYMES | authKW | 244285 | 1% | 100% | 1 |
| 10 | CULTURED ORGANOTYPIC EPITHELIA | authKW | 244285 | 1% | 100% | 1 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Cell Biology | 924 | 44% | 0% | 55 |
| 2 | Oncology | 529 | 34% | 0% | 43 |
| 3 | CYTOLOGY & HISTOLOGY | 126 | 2% | 0% | 2 |
| 4 | Pathology | 86 | 8% | 0% | 10 |
| 5 | Biology | 85 | 8% | 0% | 10 |
| 6 | Genetics & Heredity | 72 | 11% | 0% | 14 |
| 7 | Radiology, Nuclear Medicine & Medical Imaging | 39 | 8% | 0% | 10 |
| 8 | Biophysics | 36 | 7% | 0% | 9 |
| 9 | Medicine, Research & Experimental | 25 | 6% | 0% | 8 |
| 10 | Microscopy | 24 | 2% | 0% | 2 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | DERI ZUHREVI HASTALIKLAR ANABIHM DALI | 244285 | 1% | 100% | 1 |
| 2 | KENNETH NORRIS COMPREHENS CANC 3418 | 244285 | 1% | 100% | 1 |
| 3 | KENNETH NORRIS COMPREHENS CANC 3419 A | 244285 | 1% | 100% | 1 |
| 4 | KENNETH NORRIS COMPREHENS CANC 6418 | 244285 | 1% | 100% | 1 |
| 5 | MED ODONTOL MED PREVENT SALUD PUBL | 244285 | 1% | 100% | 1 |
| 6 | OTORHILARYNGOL | 244285 | 1% | 100% | 1 |
| 7 | UMR 848 | 244285 | 1% | 100% | 1 |
| 8 | VIRUS RICKETTSIAL DIS | 244285 | 1% | 100% | 1 |
| 9 | WESSEX ONCOL UNITCANC UK | 244285 | 1% | 100% | 1 |
| 10 | BIOCHEM TOXICOL EXPT CANC | 122141 | 1% | 50% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | CELL BIOLOGY INTERNATIONAL | 23576 | 14% | 1% | 18 |
| 2 | IN VITRO-JOURNAL OF THE TISSUE CULTURE ASSOCIATION | 6561 | 3% | 1% | 4 |
| 3 | CANCER CELL INTERNATIONAL | 2864 | 2% | 0% | 3 |
| 4 | ATLA-ALTERNATIVES TO LABORATORY ANIMALS | 2851 | 3% | 0% | 4 |
| 5 | CANCER COMMUNICATIONS | 2393 | 1% | 1% | 1 |
| 6 | CANCER GENETICS AND CYTOGENETICS | 1957 | 6% | 0% | 7 |
| 7 | MOLECULAR DIAGNOSIS | 1461 | 1% | 1% | 1 |
| 8 | RADIATION RESEARCH | 1333 | 5% | 0% | 6 |
| 9 | MOLECULAR CANCER | 1240 | 2% | 0% | 3 |
| 10 | IN VITRO CELLULAR & DEVELOPMENTAL BIOLOGY | 1005 | 2% | 0% | 2 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | COMPETITIVE TEMPLATE | 488569 | 2% | 100% | 2 | Search COMPETITIVE+TEMPLATE | Search COMPETITIVE+TEMPLATE |
| 2 | GIANT CANCER CELLS | 488569 | 2% | 100% | 2 | Search GIANT+CANCER+CELLS | Search GIANT+CANCER+CELLS |
| 3 | RAJU CELLS | 488569 | 2% | 100% | 2 | Search RAJU+CELLS | Search RAJU+CELLS |
| 4 | NEOSIS | 488565 | 3% | 50% | 4 | Search NEOSIS | Search NEOSIS |
| 5 | DEPOLYPLOIDIZATION | 325711 | 2% | 67% | 2 | Search DEPOLYPLOIDIZATION | Search DEPOLYPLOIDIZATION |
| 6 | AMITOSIS NUCLEAR FRAGMENTATION | 244285 | 1% | 100% | 1 | Search AMITOSIS+NUCLEAR+FRAGMENTATION | Search AMITOSIS+NUCLEAR+FRAGMENTATION |
| 7 | ANOMALOUS MITOSES | 244285 | 1% | 100% | 1 | Search ANOMALOUS+MITOSES | Search ANOMALOUS+MITOSES |
| 8 | C MYC INTERACTING GENES | 244285 | 1% | 100% | 1 | Search C+MYC+INTERACTING+GENES | Search C+MYC+INTERACTING+GENES |
| 9 | CARBONYL METABOLISING ENZYMES | 244285 | 1% | 100% | 1 | Search CARBONYL+METABOLISING+ENZYMES | Search CARBONYL+METABOLISING+ENZYMES |
| 10 | CULTURED ORGANOTYPIC EPITHELIA | 244285 | 1% | 100% | 1 | Search CULTURED+ORGANOTYPIC+EPITHELIA | Search CULTURED+ORGANOTYPIC+EPITHELIA |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | ERENPREISA, J , KALEJS, M , IANZINI, F , KOSMACEK, EA , MACKEY, MA , EMZINSH, D , CRAGG, MS , IVANOV, A , ILLIDGE, TM , (2005) SEGREGATION OF GENOMES IN POLYPLOID TUMOUR CELLS FOLLOWING MITOTIC CATASTROPHE.CELL BIOLOGY INTERNATIONAL. VOL. 29. ISSUE 12. P. 1005 -1011 | 15 | 79% | 48 |
| 2 | ERENPREISA, J , SALMINA, K , HUNA, A , KOSMACEK, EA , CRAGG, MS , IANZINI, F , ANISIMOV, AP , (2011) POLYPLOID TUMOUR CELLS ELICIT PARADIPLOID PROGENY THROUGH DEPOLYPLOIDIZING DIVISIONS AND REGULATED AUTOPHAGIC DEGRADATION.CELL BIOLOGY INTERNATIONAL. VOL. 35. ISSUE 7. P. 687 -695 | 20 | 38% | 15 |
| 3 | ERENPREISA, J , CRAGG, MS , (2010) MOS, ANEUPLOIDY AND THE PLOIDY CYCLE OF CANCER CELLS.ONCOGENE. VOL. 29. ISSUE 40. P. 5447 -5451 | 18 | 36% | 19 |
| 4 | ERENPREISA, J , CRAGG, MS , SALMINA, K , HAUSMANN, M , SCHERTHAN, H , (2009) THE ROLE OF MEIOTIC COHESIN REC8 IN CHROMOSOME SEGREGATION IN GAMMA IRRADIATION-INDUCED ENDOPOLYPLOID TUMOUR CELLS.EXPERIMENTAL CELL RESEARCH. VOL. 315. ISSUE 15. P. 2593 -2603 | 17 | 33% | 14 |
| 5 | NIU, N , ZHANG, J , ZHANG, N , MERCADO-URIBE, I , TAO, F , HAN, Z , PATHAK, S , MULTANI, AS , KUANG, J , YAO, J , ET AL (2016) LINKING GENOMIC REORGANIZATION TO TUMOR INITIATION VIA THE GIANT CELL CYCLE.ONCOGENESIS. VOL. 5. ISSUE . P. - | 14 | 32% | 0 |
| 6 | ERENPREISA, J , CRAGG, MS , (2013) THREE STEPS TO THE IMMORTALITY OF CANCER CELLS: SENESCENCE, POLYPLOIDY AND SELF-RENEWAL.CANCER CELL INTERNATIONAL. VOL. 13. ISSUE . P. - | 23 | 18% | 17 |
| 7 | WALEN, KH , (2007) BIPOLAR GENOME REDUCTIONAL DIVISION OF HUMAN NEAR-SENESCENT, POLYPLOID FIBROBLAST CELLS.CANCER GENETICS AND CYTOGENETICS. VOL. 173. ISSUE 1. P. 43-50 | 13 | 39% | 6 |
| 8 | ERENPREISA, J , IVANOV, A , WHEATLEY, SP , KOSMACEK, EA , IANZINI, F , ANISIMOV, AP , MACKEY, M , DAVIS, PJ , PLAKHINS, G , ILLIDGE, TM , (2008) ENDOPOLYPLOIDY IN IRRADIATED P53-DEFICIENT TUMOUR CELL LINES: PERSISTENCE OF CELL DIVISION ACTIVITY IN GIANT CELLS EXPRESSING AURORA-B KINASE.CELL BIOLOGY INTERNATIONAL. VOL. 32. ISSUE 9. P. 1044 -1056 | 12 | 36% | 24 |
| 9 | IANZINI, F , KOSMACEK, EA , NELSON, ES , NAPOLI, E , ERENPREISA, J , KALEJS, M , MACKEY, MA , (2009) ACTIVATION OF MEIOSIS-SPECIFIC GENES IS ASSOCIATED WITH DEPOLYPLOIDIZATION OF HUMAN TUMOR CELLS FOLLOWING RADIATION-INDUCED MITOTIC CATASTROPHE.CANCER RESEARCH. VOL. 69. ISSUE 6. P. 2296-2304 | 13 | 29% | 31 |
| 10 | STAAB, CA , VONDRACEK, M , CUSTODIO, H , JOHANSSON, K , NILSSON, JA , MORGAN, P , HOOG, JO , COTGREAVE, I , GRASFSTROM, RC , (2004) MODELLING OF NORMAL AND PREMALIGNANT ORAL TISSUE BY USING THE IMMORTALISED CELL LINE, SVPGC2A: A REVIEW OF THE VALUE OF THE MODEL.ATLA-ALTERNATIVES TO LABORATORY ANIMALS. VOL. 32. ISSUE 4. P. 401-405 | 7 | 54% | 4 |
Classes with closest relation at Level 1 |