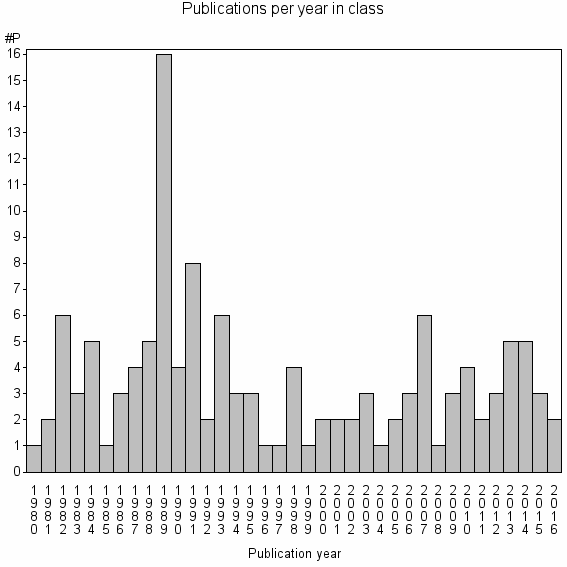
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 33141 | 128 | 20.7 | 49% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 3 | 4 | PLANT SCIENCES//AGRONOMY//FOOD SCIENCE & TECHNOLOGY | 1882812 |
| 690 | 3 | ALLELOPATHY//ALLELOPATHY JOURNAL//ALLELOCHEMICALS | 7846 |
| 2366 | 2 | ALLELOPATHY//ALLELOPATHY JOURNAL//ALLELOCHEMICALS | 4330 |
| 33141 | 1 | NYLON OLIGOMER//6 AMINOHEXANOATE DIMER HYDROLASE//CAP PLASMIDS | 128 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | NYLON OLIGOMER | authKW | 1932327 | 7% | 90% | 9 |
| 2 | 6 AMINOHEXANOATE DIMER HYDROLASE | authKW | 1431355 | 5% | 100% | 6 |
| 3 | CAP PLASMIDS | authKW | 477118 | 2% | 100% | 2 |
| 4 | INTELLIGENCE MECH TECHNOL | address | 477118 | 2% | 100% | 2 |
| 5 | 6 AMINOHEXANOATE CYCLIC DIMER HYDROLASE | authKW | 318078 | 2% | 67% | 2 |
| 6 | 6 ACA | authKW | 238559 | 1% | 100% | 1 |
| 7 | 6 AMINOHEXANOATE CYCLIC DIMER | authKW | 238559 | 1% | 100% | 1 |
| 8 | 6 AMINOHEXANOATE LINEAR DIMER HYDROLASE | authKW | 238559 | 1% | 100% | 1 |
| 9 | 6 AMINOHEXANOATE OLIGOMER HYDROLASE | authKW | 238559 | 1% | 100% | 1 |
| 10 | ACIDOVORAX FACILIS B | authKW | 238559 | 1% | 100% | 1 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biotechnology & Applied Microbiology | 826 | 34% | 0% | 44 |
| 2 | Microbiology | 376 | 23% | 0% | 30 |
| 3 | Toxicology | 177 | 12% | 0% | 15 |
| 4 | Food Science & Technology | 113 | 12% | 0% | 15 |
| 5 | Biochemistry & Molecular Biology | 41 | 19% | 0% | 24 |
| 6 | Genetics & Heredity | 31 | 8% | 0% | 10 |
| 7 | Engineering, Environmental | 23 | 4% | 0% | 5 |
| 8 | Biophysics | 12 | 5% | 0% | 6 |
| 9 | Crystallography | 11 | 3% | 0% | 4 |
| 10 | Biochemical Research Methods | 10 | 4% | 0% | 5 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | INTELLIGENCE MECH TECHNOL | 477118 | 2% | 100% | 2 |
| 2 | BASIC EDUC INTEGRATED LEARNING | 119275 | 2% | 17% | 3 |
| 3 | OSTI | 34078 | 1% | 14% | 1 |
| 4 | MAT SCI CHEM | 26751 | 9% | 1% | 12 |
| 5 | PLASMID BIOL | 23854 | 1% | 10% | 1 |
| 6 | PUSHCHINO | 19878 | 1% | 8% | 1 |
| 7 | MAT ENGN SCI | 6893 | 4% | 1% | 5 |
| 8 | INGN CIENCIAS ECON SOCIALES | 6115 | 1% | 3% | 1 |
| 9 | MOL BIOTECHNOL GRP | 5074 | 1% | 2% | 1 |
| 10 | RIKEN HARIMA | 4867 | 1% | 2% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | MIRCEN-JOURNAL OF APPLIED MICROBIOLOGY AND BIOTECHNOLOGY | 4499 | 1% | 2% | 1 |
| 2 | INTERNATIONAL BIODETERIORATION BULLETIN | 3668 | 1% | 2% | 1 |
| 3 | MUTATION RESEARCH | 2935 | 8% | 0% | 10 |
| 4 | JOURNAL OF FERMENTATION TECHNOLOGY | 2635 | 2% | 0% | 3 |
| 5 | ENZYME ENGINEERING | 2509 | 1% | 1% | 1 |
| 6 | MICROBIOLOGY | 2111 | 5% | 0% | 6 |
| 7 | CRC CRITICAL REVIEWS IN TOXICOLOGY | 2020 | 1% | 1% | 1 |
| 8 | JOURNAL OF FERMENTATION AND BIOENGINEERING | 1724 | 3% | 0% | 4 |
| 9 | PROCESS CONTROL AND QUALITY | 1632 | 1% | 1% | 1 |
| 10 | JOURNAL OF BIOTECHNOLOGY | 1295 | 5% | 0% | 6 |
Author Key Words |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | NAGAI, K , YASUHIRA, K , TANAKA, Y , KATO, D , TAKEO, M , HIGUCHI, Y , NEGORO, S , SHIBATA, N , (2013) CRYSTALLIZATION AND X-RAY DIFFRACTION ANALYSIS OF NYLON HYDROLASE (NYLC) FROM ARTHROBACTER SP KI72.ACTA CRYSTALLOGRAPHICA SECTION F-STRUCTURAL BIOLOGY COMMUNICATIONS. VOL. 69. ISSUE . P. 1151-1154 | 14 | 82% | 0 |
| 2 | ESIKOVA, TZ , TARAN, SA , (2016) A NOVEL STRAIN GULOSIBACTER SP BS4 DEGRADING EPSILON-CAPROLACTAM AND NYLON-6 OLIGOMERS.MICROBIOLOGY. VOL. 85. ISSUE 5. P. 642 -645 | 10 | 91% | 0 |
| 3 | YASUHIRA, K , SHIBATA, N , TANAKA, Y , KUMAGAI, N , TANAKA, Y , NAGAI, K , KATO, D , TAKEO, M , NEGORO, S , HIGUCHI, Y , (2011) CRYSTALLIZATION AND X-RAY DIFFRACTION ANALYSIS OF NYLON-OLIGOMER HYDROLASE (NYLC) FROM AGROMYCES SP KY5R.ACTA CRYSTALLOGRAPHICA SECTION F-STRUCTURAL BIOLOGY AND CRYSTALLIZATION COMMUNICATIONS. VOL. 67. ISSUE . P. 892 -895 | 13 | 76% | 1 |
| 4 | NEGORO, S , KAWASHIMA, Y , SHIBATA, N , KOBAYASHI, T , BABA, T , LEE, YH , KAMIYA, K , SHIGETA, Y , NAGAI, K , TAKEHARA, I , ET AL (2016) MUTATIONS AFFECTING THE INTERNAL EQUILIBRIUM OF THE REACTION CATALYZED BY 6-AMINOHEXANOATE-DIMER HYDROLASE.FEBS LETTERS. VOL. 590. ISSUE 18. P. 3133 -3143 | 20 | 39% | 0 |
| 5 | NEGORO, S , (2000) BIODEGRADATION OF NYLON OLIGOMERS.APPLIED MICROBIOLOGY AND BIOTECHNOLOGY. VOL. 54. ISSUE 4. P. 461-466 | 21 | 50% | 48 |
| 6 | ESIKOVA, TZ , AKATOVA, EV , TARAN, SA , (2014) BACTERIA THAT DEGRADE LOW-MOLECULAR LINEAR EPSILON-CAPROLACTAM OLIGOMERS.APPLIED BIOCHEMISTRY AND MICROBIOLOGY. VOL. 50. ISSUE 5. P. 463 -470 | 8 | 89% | 0 |
| 7 | KAWASHIMA, Y , OHKI, T , SHIBATA, N , HIGUCHI, Y , WAKITANI, Y , MATSUURA, Y , NAKATA, Y , TAKEO, M , KATO, D , NEGORO, S , (2009) MOLECULAR DESIGN OF A NYLON-6 BYPRODUCT-DEGRADING ENZYME FROM A CARBOXYLESTERASE WITH A BETA-LACTAMASE FOLD.FEBS JOURNAL. VOL. 276. ISSUE 9. P. 2547-2556 | 12 | 57% | 7 |
| 8 | NEGORO, S , (1992) GENETIC ORGANIZATION OF THE ENZYMES RESPONSIBLE FOR DEGRADATION OF NYLON OLIGOMERS.HAKKOKOGAKU KAISHI-JOURNAL OF THE SOCIETY OF FERMENTATION TECHNOLOGY. VOL. 70. ISSUE 2. P. 115-131 | 13 | 93% | 0 |
| 9 | ESIKOVA, T , PONAMOREVA, O , BASKUNOV, B , TARAN, S , BORONIN, A , (2012) TRANSFORMATION OF LOW-MOLECULAR LINEAR CAPROLACTAM OLIGOMERS BY CAPROLACTAM-DEGRADING BACTERIA.JOURNAL OF CHEMICAL TECHNOLOGY AND BIOTECHNOLOGY. VOL. 87. ISSUE 9. P. 1284 -1290 | 8 | 73% | 4 |
| 10 | BABA, T , KAMIYA, K , MATSUI, T , SHIBATA, N , HIGUCHI, Y , KOBAYASHI, T , NEGORO, S , SHIGETA, Y , (2011) MOLECULAR DYNAMICS STUDIES ON THE MUTATIONAL STRUCTURES OF A NYLON-6 BYPRODUCT-DEGRADING ENZYME.CHEMICAL PHYSICS LETTERS. VOL. 507. ISSUE 1-3. P. 157 -161 | 11 | 55% | 0 |
Classes with closest relation at Level 1 |