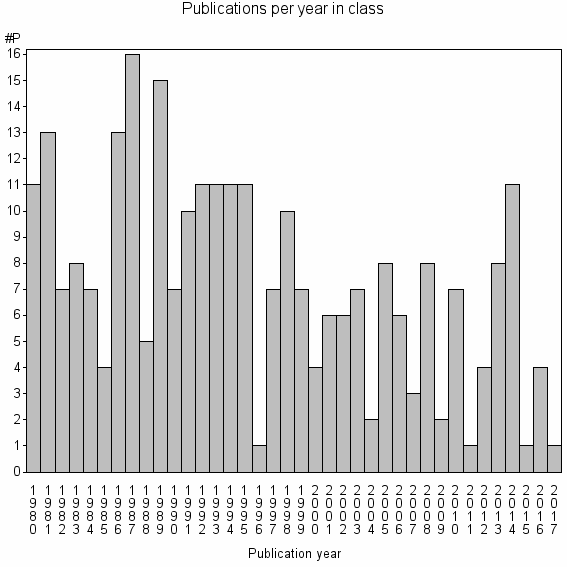
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 25686 | 274 | 20.2 | 40% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | CHROMOSOME CLASSIFICATION | authKW | 1805367 | 7% | 90% | 18 |
| 2 | AUTOMATED KARYOTYPING | authKW | 334328 | 1% | 100% | 3 |
| 3 | AUTOMATED METAPHASE FINDING | authKW | 334328 | 1% | 100% | 3 |
| 4 | METAPHASE FINDING | authKW | 334328 | 1% | 100% | 3 |
| 5 | TUKEYS POST HOC | authKW | 334328 | 1% | 100% | 3 |
| 6 | ABERRATION SCORING | authKW | 250744 | 1% | 75% | 3 |
| 7 | KARYOTYPING | authKW | 223756 | 10% | 7% | 27 |
| 8 | ANALYSIS OF CHROMOSOMES | authKW | 222885 | 1% | 100% | 2 |
| 9 | BANDED CHROMOSOME | authKW | 222885 | 1% | 100% | 2 |
| 10 | CENTROMERE FINDING | authKW | 222885 | 1% | 100% | 2 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Computer Science, Artificial Intelligence | 3301 | 29% | 0% | 79 |
| 2 | Medical Informatics | 516 | 5% | 0% | 15 |
| 3 | Mathematical & Computational Biology | 362 | 7% | 0% | 19 |
| 4 | CYTOLOGY & HISTOLOGY | 361 | 2% | 0% | 5 |
| 5 | Engineering, Biomedical | 355 | 9% | 0% | 26 |
| 6 | Computer Science, Interdisciplinary Applications | 328 | 10% | 0% | 27 |
| 7 | Genetics & Heredity | 305 | 15% | 0% | 41 |
| 8 | Engineering, Electrical & Electronic | 221 | 19% | 0% | 51 |
| 9 | Computer Science, Theory & Methods | 220 | 8% | 0% | 23 |
| 10 | Biology | 189 | 8% | 0% | 22 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | CONTROL INTELLIGNET PROC EXCELLENCE | 111443 | 0% | 100% | 1 |
| 2 | DMX MET MECAN | 111443 | 0% | 100% | 1 |
| 3 | GRP ANALYSE IMAGES BIOMED | 111443 | 0% | 100% | 1 |
| 4 | IMAGES VIS GRP | 111443 | 0% | 100% | 1 |
| 5 | MRC CLIN POPULAT CYTOGENET | 111443 | 0% | 100% | 1 |
| 6 | RADIAT HAZARDS INAGE KU | 111443 | 0% | 100% | 1 |
| 7 | DSIC ITI | 37146 | 0% | 33% | 1 |
| 8 | PHYS INAGE KU | 37146 | 0% | 33% | 1 |
| 9 | SYST INTELLIGENTS PERCEPT | 37146 | 0% | 33% | 1 |
| 10 | IMAGE VIS | 18572 | 0% | 17% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | PATTERN RECOGNITION LETTERS | 11118 | 9% | 0% | 24 |
| 2 | CYTOMETRY | 9243 | 5% | 1% | 15 |
| 3 | PATTERN RECOGNITION | 7166 | 8% | 0% | 21 |
| 4 | ANALYTICAL AND QUANTITATIVE CYTOLOGY AND HISTOLOGY | 3942 | 3% | 0% | 8 |
| 5 | PATTERN ANALYSIS AND APPLICATIONS | 3876 | 2% | 1% | 5 |
| 6 | CHEMICAL MUTAGENS-PRINCIPLES AND METHODS FOR THEIR DETECTION | 3481 | 0% | 3% | 1 |
| 7 | JOURNAL OF RADIATION RESEARCH | 3450 | 3% | 0% | 8 |
| 8 | COMPUTERS IN BIOLOGY AND MEDICINE | 2436 | 3% | 0% | 8 |
| 9 | CLINICAL GENETICS | 2178 | 4% | 0% | 10 |
| 10 | IMAGE AND VISION COMPUTING | 2113 | 3% | 0% | 7 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | CHROMOSOME CLASSIFICATION | 1805367 | 7% | 90% | 18 | Search CHROMOSOME+CLASSIFICATION | Search CHROMOSOME+CLASSIFICATION |
| 2 | AUTOMATED KARYOTYPING | 334328 | 1% | 100% | 3 | Search AUTOMATED+KARYOTYPING | Search AUTOMATED+KARYOTYPING |
| 3 | AUTOMATED METAPHASE FINDING | 334328 | 1% | 100% | 3 | Search AUTOMATED+METAPHASE+FINDING | Search AUTOMATED+METAPHASE+FINDING |
| 4 | METAPHASE FINDING | 334328 | 1% | 100% | 3 | Search METAPHASE+FINDING | Search METAPHASE+FINDING |
| 5 | TUKEYS POST HOC | 334328 | 1% | 100% | 3 | Search TUKEYS+POST+HOC | Search TUKEYS+POST+HOC |
| 6 | ABERRATION SCORING | 250744 | 1% | 75% | 3 | Search ABERRATION+SCORING | Search ABERRATION+SCORING |
| 7 | KARYOTYPING | 223756 | 10% | 7% | 27 | Search KARYOTYPING | Search KARYOTYPING |
| 8 | ANALYSIS OF CHROMOSOMES | 222885 | 1% | 100% | 2 | Search ANALYSIS+OF+CHROMOSOMES | Search ANALYSIS+OF+CHROMOSOMES |
| 9 | BANDED CHROMOSOME | 222885 | 1% | 100% | 2 | Search BANDED+CHROMOSOME | Search BANDED+CHROMOSOME |
| 10 | CENTROMERE FINDING | 222885 | 1% | 100% | 2 | Search CENTROMERE+FINDING | Search CENTROMERE+FINDING |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | ZHENG, B , LI, SB , MULVIHILL, JJ , LIU, H , WANG, XW , (2008) DEVELOPMENT AND ASSESSMENT OF AN INTEGRATED COMPUTER-AIDED DETECTION SCHEME FOR DIGITAL MICROSCOPIC IMAGES OF METAPHASE CHROMOSOMES.JOURNAL OF ELECTRONIC IMAGING. VOL. 17. ISSUE 4. P. - | 23 | 85% | 1 |
| 2 | WANG, XW , ZHENG, B , LI, SB , MULVIHILL, JJ , WOOD, MC , LIU, H , (2009) AUTOMATED CLASSIFICATION OF METAPHASE CHROMOSOMES: OPTIMIZATION OF AN ADAPTIVE COMPUTERIZED SCHEME.JOURNAL OF BIOMEDICAL INFORMATICS. VOL. 42. ISSUE 1. P. 22 -31 | 21 | 84% | 12 |
| 3 | ARORA, T , DHIR, R , (2016) A REVIEW OF METAPHASE CHROMOSOME IMAGE SELECTION TECHNIQUES FOR AUTOMATIC KARYOTYPE GENERATION.MEDICAL & BIOLOGICAL ENGINEERING & COMPUTING. VOL. 54. ISSUE 8. P. 1147 -1157 | 17 | 89% | 0 |
| 4 | POLETTI, E , GRISAN, E , RUGGERI, A , (2012) A MODULAR FRAMEWORK FOR THE AUTOMATIC CLASSIFICATION OF CHROMOSOMES IN Q-BAND IMAGES.COMPUTER METHODS AND PROGRAMS IN BIOMEDICINE. VOL. 105. ISSUE 2. P. 120 -130 | 16 | 94% | 4 |
| 5 | FENG, XW , CONG, PS , ZHU, ZL , DU, XY , (2012) AUTOMATED PAIRING OF HUMAN CHROMOSOMES APPLYING GRADIENT PROFILE AND SIMILARITY MATCHING ALGORITHM.CHEMOMETRICS AND INTELLIGENT LABORATORY SYSTEMS. VOL. 111. ISSUE 1. P. 46 -52 | 19 | 73% | 5 |
| 6 | GRISAN, E , POLETTI, E , RUGGERI, A , (2009) AUTOMATIC SEGMENTATION AND DISENTANGLING OF CHROMOSOMES IN Q-BAND PROMETAPHASE IMAGES.IEEE TRANSACTIONS ON INFORMATION TECHNOLOGY IN BIOMEDICINE. VOL. 13. ISSUE 4. P. 575 -581 | 14 | 100% | 17 |
| 7 | KAO, JH , CHUANG, JH , WANG, T , (2008) CHROMOSOME CLASSIFICATION BASED ON THE BAND PROFILE SIMILARITY ALONG APPROXIMATE MEDIAL AXIS.PATTERN RECOGNITION. VOL. 41. ISSUE 1. P. 77-89 | 16 | 89% | 16 |
| 8 | SIVARAMAKRISHNAN, R , ARUN, C , (2014) LUKASIEWICZ LOGIC BASED FUZZY SIMILARITY CLASSIFIER FOR DENVER GROUP CHROMOSOMAL CLASSIFICATION.BIOSCIENCE JOURNAL. VOL. 30. ISSUE 3. P. 843 -U533 | 17 | 74% | 1 |
| 9 | WANG, X , ZHENG, B , LI, S , MULVIHILL, JJ , LIU, H , (2008) A RULE-BASED COMPUTER SCHEME FOR CENTROMERE IDENTIFICATION AND POLARITY ASSIGNMENT OF METAPHASE CHROMOSOMES.COMPUTER METHODS AND PROGRAMS IN BIOMEDICINE. VOL. 89. ISSUE 1. P. 33 -42 | 14 | 88% | 11 |
| 10 | CAROTHERS, A , PIPER, J , (1994) COMPUTER-AIDED CLASSIFICATION OF HUMAN-CHROMOSOMES - A REVIEW.STATISTICS AND COMPUTING. VOL. 4. ISSUE 3. P. 161-171 | 20 | 87% | 55 |
Classes with closest relation at Level 1 |