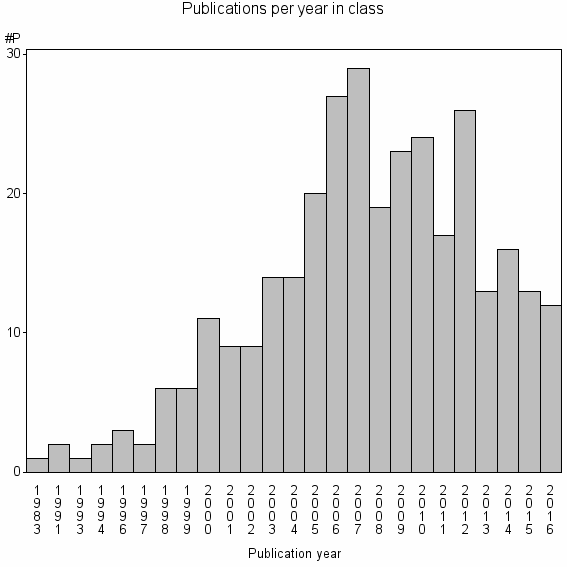
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 24226 | 319 | 35.2 | 84% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 4 | 4 | ENVIRONMENTAL SCIENCES//GEOSCIENCES, MULTIDISCIPLINARY//METEOROLOGY & ATMOSPHERIC SCIENCES | 1766162 |
| 50 | 3 | ANAEROBIC DIGESTION//MICROBIAL FUEL CELL//WATER SCIENCE AND TECHNOLOGY | 91931 |
| 1312 | 2 | MICROBIOLOGY//PLANCTOMYCETES//PROPIDIUM MONOAZIDE | 8512 |
| 24226 | 1 | T RFLP//TERMINAL RESTRICTION FRAGMENT LENGTH POLYMORPHISM//LENGTH HETEROGENEITY PCR LH PCR | 319 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | T RFLP | authKW | 519716 | 18% | 9% | 59 |
| 2 | TERMINAL RESTRICTION FRAGMENT LENGTH POLYMORPHISM | authKW | 197508 | 6% | 11% | 18 |
| 3 | LENGTH HETEROGENEITY PCR LH PCR | authKW | 191443 | 1% | 100% | 2 |
| 4 | TERMINAL RESTRICTION FRAGMENT LENGTH POLYMORPHISM T RFLP | authKW | 156499 | 3% | 15% | 11 |
| 5 | 16S RECOMBINANT RNA GENE | authKW | 95722 | 0% | 100% | 1 |
| 6 | 3 CHLOROANALINE | authKW | 95722 | 0% | 100% | 1 |
| 7 | 63F | authKW | 95722 | 0% | 100% | 1 |
| 8 | 8 27F | authKW | 95722 | 0% | 100% | 1 |
| 9 | ALGAL ANALYSIS | authKW | 95722 | 0% | 100% | 1 |
| 10 | ALIMENTAT UR 106 | address | 95722 | 0% | 100% | 1 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Microbiology | 5455 | 54% | 0% | 173 |
| 2 | Biotechnology & Applied Microbiology | 2073 | 34% | 0% | 110 |
| 3 | Biochemical Research Methods | 649 | 16% | 0% | 50 |
| 4 | Engineering, Environmental | 270 | 8% | 0% | 25 |
| 5 | Environmental Sciences | 154 | 12% | 0% | 39 |
| 6 | Soil Science | 149 | 4% | 0% | 14 |
| 7 | Water Resources | 127 | 6% | 0% | 19 |
| 8 | Ecology | 96 | 7% | 0% | 23 |
| 9 | Marine & Freshwater Biology | 90 | 6% | 0% | 18 |
| 10 | Food Science & Technology | 57 | 6% | 0% | 19 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | ALIMENTAT UR 106 | 95722 | 0% | 100% | 1 |
| 2 | ATLANGENC SILLIKER | 95722 | 0% | 100% | 1 |
| 3 | CE 1 644 | 95722 | 0% | 100% | 1 |
| 4 | ENAC ISTE ECOSYST | 95722 | 0% | 100% | 1 |
| 5 | ENAC ISTE ENVIRONM BIOTECHNOL | 95722 | 0% | 100% | 1 |
| 6 | ENVIRONM MICROBIOL STN 6 | 95722 | 0% | 100% | 1 |
| 7 | GERMNA BIOTECHNOL | 95722 | 0% | 100% | 1 |
| 8 | INVEST INGN GENET BIOL MOL INGEBI CONICET | 95722 | 0% | 100% | 1 |
| 9 | MELBOURNE ENVIRONM | 95722 | 0% | 100% | 1 |
| 10 | MELBOURNE LAND | 95722 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | JOURNAL OF MICROBIOLOGICAL METHODS | 19055 | 10% | 1% | 31 |
| 2 | MICROBIAL ECOLOGY | 5680 | 4% | 0% | 14 |
| 3 | APPLIED AND ENVIRONMENTAL MICROBIOLOGY | 5651 | 13% | 0% | 42 |
| 4 | FEMS MICROBIOLOGY ECOLOGY | 5176 | 5% | 0% | 15 |
| 5 | ENVIRONMENTAL MICROBIOLOGY | 2498 | 3% | 0% | 10 |
| 6 | MICROBES AND ENVIRONMENTS | 1555 | 1% | 1% | 3 |
| 7 | JOURNAL OF BIOSCIENCE AND BIOENGINEERING | 835 | 2% | 0% | 6 |
| 8 | WATER RESEARCH | 745 | 3% | 0% | 11 |
| 9 | PCR-METHODS AND APPLICATIONS | 582 | 0% | 1% | 1 |
| 10 | BIOTECHNIQUES | 559 | 2% | 0% | 6 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | T RFLP | 519716 | 18% | 9% | 59 | Search T+RFLP | Search T+RFLP |
| 2 | TERMINAL RESTRICTION FRAGMENT LENGTH POLYMORPHISM | 197508 | 6% | 11% | 18 | Search TERMINAL+RESTRICTION+FRAGMENT+LENGTH+POLYMORPHISM | Search TERMINAL+RESTRICTION+FRAGMENT+LENGTH+POLYMORPHISM |
| 3 | LENGTH HETEROGENEITY PCR LH PCR | 191443 | 1% | 100% | 2 | Search LENGTH+HETEROGENEITY+PCR+LH+PCR | Search LENGTH+HETEROGENEITY+PCR+LH+PCR |
| 4 | TERMINAL RESTRICTION FRAGMENT LENGTH POLYMORPHISM T RFLP | 156499 | 3% | 15% | 11 | Search TERMINAL+RESTRICTION+FRAGMENT+LENGTH+POLYMORPHISM+T+RFLP | Search TERMINAL+RESTRICTION+FRAGMENT+LENGTH+POLYMORPHISM+T+RFLP |
| 5 | 16S RECOMBINANT RNA GENE | 95722 | 0% | 100% | 1 | Search 16S+RECOMBINANT+RNA+GENE | Search 16S+RECOMBINANT+RNA+GENE |
| 6 | 3 CHLOROANALINE | 95722 | 0% | 100% | 1 | Search 3+CHLOROANALINE | Search 3+CHLOROANALINE |
| 7 | 63F | 95722 | 0% | 100% | 1 | Search 63F | Search 63F |
| 8 | 8 27F | 95722 | 0% | 100% | 1 | Search 8+27F | Search 8+27F |
| 9 | ALGAL ANALYSIS | 95722 | 0% | 100% | 1 | Search ALGAL+ANALYSIS | Search ALGAL+ANALYSIS |
| 10 | AMF PLASMID DNA | 95722 | 0% | 100% | 1 | Search AMF+PLASMID+DNA | Search AMF+PLASMID+DNA |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | FREDRIKSSON, NJ , HERMANSSON, M , WILEN, BM , (2014) TOOLS FOR T-RFLP DATA ANALYSIS USING EXCEL.BMC BIOINFORMATICS. VOL. 15. ISSUE . P. - | 19 | 95% | 0 |
| 2 | FREDRIKSSON, NJ , HERMANSSON, M , WILEN, BM , (2014) IMPACT OF T-RFLP DATA ANALYSIS CHOICES ON ASSESSMENTS OF MICROBIAL COMMUNITY STRUCTURE AND DYNAMICS.BMC BIOINFORMATICS. VOL. 15. ISSUE . P. - | 22 | 76% | 4 |
| 3 | SCHUTTE, UME , ABDO, Z , BENT, SJ , SHYU, C , WILLIAMS, CJ , PIERSON, JD , FORNEY, LJ , (2008) ADVANCES IN THE USE OF TERMINAL RESTRICTION FRAGMENT LENGTH POLYMORPHISM (T-RFLP) ANALYSIS OF 16S RRNA GENES TO CHARACTERIZE MICROBIAL COMMUNITIES.APPLIED MICROBIOLOGY AND BIOTECHNOLOGY. VOL. 80. ISSUE 3. P. 365 -380 | 39 | 35% | 179 |
| 4 | CULMAN, SW , BUKOWSKI, R , GAUCH, HG , CADILLO-QUIROZ, H , BUCKLEY, DH , (2009) T-REX: SOFTWARE FOR THE PROCESSING AND ANALYSIS OF T-RFLP DATA.BMC BIOINFORMATICS. VOL. 10. ISSUE . P. - | 13 | 81% | 207 |
| 5 | PRAKASH, O , PANDEY, PK , KULKARNI, GJ , MAHALE, KN , SHOUCHE, YS , (2014) TECHNICALITIES AND GLITCHES OF TERMINAL RESTRICTION FRAGMENT LENGTH POLYMORPHISM (T-RFLP).INDIAN JOURNAL OF MICROBIOLOGY. VOL. 54. ISSUE 3. P. 255-261 | 25 | 52% | 5 |
| 6 | THIES, JE , (2007) SOIL MICROBIAL COMMUNITY ANALYSIS USING TERMINAL RESTRICTION FRAGMENT LENGTH POLYMORPHISMS.SOIL SCIENCE SOCIETY OF AMERICA JOURNAL. VOL. 71. ISSUE 2. P. 579-591 | 50 | 26% | 63 |
| 7 | ROSSI, P , GILLET, F , ROHRBACH, E , DIABY, N , HOLLIGER, C , (2009) STATISTICAL ASSESSMENT OF VARIABILITY OF TERMINAL RESTRICTION FRAGMENT LENGTH POLYMORPHISM ANALYSIS APPLIED TO COMPLEX MICROBIAL COMMUNITIES.APPLIED AND ENVIRONMENTAL MICROBIOLOGY. VOL. 75. ISSUE 22. P. 7268 -7270 | 17 | 71% | 7 |
| 8 | ZHANG, R , THIYAGARAJAN, V , QIAN, PY , (2008) EVALUATION OF TERMINAL-RESTRICTION FRAGMENT LENGTH POLYMORPHISM ANALYSIS IN CONTRASTING MARINE ENVIRONMENTS.FEMS MICROBIOLOGY ECOLOGY. VOL. 65. ISSUE 1. P. 169 -178 | 19 | 59% | 39 |
| 9 | KOPECKY, J , NOVOTNA, G , SAGOVA-MARECKOVA, M , (2009) MODIFICATION OF THE TERMINAL RESTRICTION FRAGMENT LENGTH POLYMORPHISM ANALYSIS FOR ASSESSMENT OF A SPECIFIC TAXONOMIC GROUP WITHIN A SOIL MICROBIAL COMMUNITY.PLANT SOIL AND ENVIRONMENT. VOL. 55. ISSUE 9. P. 397-403 | 16 | 64% | 1 |
| 10 | AIKEN, JT , (2011) TERMINAL RESTRICTION FRAGMENT LENGTH POLYMORPHISM FOR SOIL MICROBIAL COMMUNITY FINGERPRINTING.SOIL SCIENCE SOCIETY OF AMERICA JOURNAL. VOL. 75. ISSUE 1. P. 102-111 | 15 | 58% | 5 |
Classes with closest relation at Level 1 |