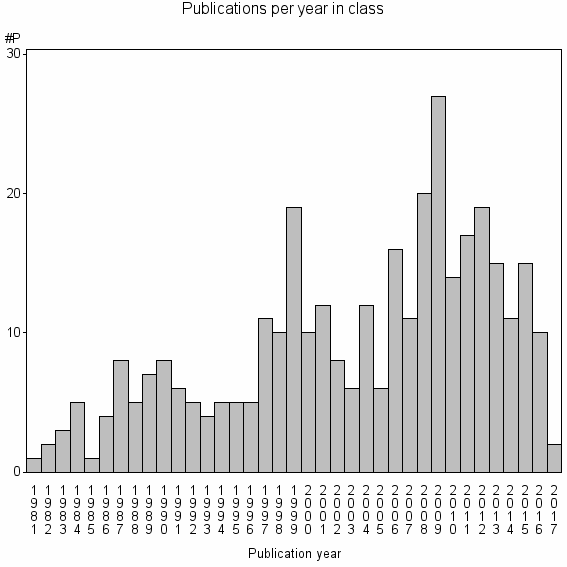
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 23453 | 345 | 35.8 | 79% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 142 | 3 | DNA REPAIR//INTERNATIONAL JOURNAL OF RADIATION BIOLOGY//RADIATION RESEARCH | 64602 |
| 1228 | 2 | BASE EXCISION REPAIR//DNA GLYCOSYLASE//OXIDATIVE DNA DAMAGE | 9086 |
| 23453 | 1 | ABASIC SITE//REGULATORY BIOORGAN CHEM//CHIM BIOORGANLEDSS | 345 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | ABASIC SITE | authKW | 730255 | 11% | 22% | 38 |
| 2 | REGULATORY BIOORGAN CHEM | address | 327291 | 4% | 26% | 14 |
| 3 | CHIM BIOORGANLEDSS | address | 177015 | 1% | 100% | 2 |
| 4 | FLUORESCENCE SELECTIVITY | authKW | 177015 | 1% | 100% | 2 |
| 5 | GG MISMATCH | authKW | 177015 | 1% | 100% | 2 |
| 6 | INDUCED HAIRPIN FORMATION | authKW | 177015 | 1% | 100% | 2 |
| 7 | UREA PHENANTHRIDINIUM | authKW | 177015 | 1% | 100% | 2 |
| 8 | CHEM BIOL CRYSTALLOG | address | 144822 | 2% | 27% | 6 |
| 9 | STUDY INTERACT BIOMACROMOL | address | 138286 | 1% | 31% | 5 |
| 10 | NUCLEOBASE RECOGNITION | authKW | 132758 | 1% | 50% | 3 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Chemistry, Organic | 626 | 20% | 0% | 70 |
| 2 | Chemistry, Medicinal | 556 | 12% | 0% | 43 |
| 3 | Biochemistry & Molecular Biology | 494 | 34% | 0% | 116 |
| 4 | Chemistry, Multidisciplinary | 372 | 23% | 0% | 80 |
| 5 | Chemistry, Analytical | 155 | 10% | 0% | 36 |
| 6 | Biophysics | 83 | 7% | 0% | 23 |
| 7 | Electrochemistry | 31 | 3% | 0% | 11 |
| 8 | Biochemical Research Methods | 24 | 4% | 0% | 13 |
| 9 | Toxicology | 22 | 3% | 0% | 11 |
| 10 | Crystallography | 18 | 3% | 0% | 9 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | REGULATORY BIOORGAN CHEM | 327291 | 4% | 26% | 14 |
| 2 | CHIM BIOORGANLEDSS | 177015 | 1% | 100% | 2 |
| 3 | CHEM BIOL CRYSTALLOG | 144822 | 2% | 27% | 6 |
| 4 | STUDY INTERACT BIOMACROMOL | 138286 | 1% | 31% | 5 |
| 5 | UMR 5616 | 100227 | 5% | 7% | 16 |
| 6 | CHEM MODELLING IMAGING BIOL | 88508 | 0% | 100% | 1 |
| 7 | CHIM BIOORGAN CNRS UA488 | 88508 | 0% | 100% | 1 |
| 8 | CNRSINSERMU1196UMR9187 | 88508 | 0% | 100% | 1 |
| 9 | INSERMU1196 | 88508 | 0% | 100% | 1 |
| 10 | JST ENGN SYNTHET CHEM BIOL CHEM | 88508 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | NUCLEOSIDES NUCLEOTIDES & NUCLEIC ACIDS | 3192 | 4% | 0% | 13 |
| 2 | CLAVIER | 1922 | 0% | 2% | 1 |
| 3 | ORGANIC & BIOMOLECULAR CHEMISTRY | 1922 | 5% | 0% | 16 |
| 4 | CHEMICAL COMMUNICATIONS | 890 | 6% | 0% | 21 |
| 5 | NUCLEIC ACIDS RESEARCH | 871 | 6% | 0% | 20 |
| 6 | CHEMISTRY-A EUROPEAN JOURNAL | 834 | 4% | 0% | 15 |
| 7 | BIOORGANIC & MEDICINAL CHEMISTRY | 809 | 3% | 0% | 11 |
| 8 | BIOCHEMISTRY | 776 | 6% | 0% | 22 |
| 9 | BIOORGANIC & MEDICINAL CHEMISTRY LETTERS | 770 | 4% | 0% | 15 |
| 10 | ANTI-CANCER DRUG DESIGN | 561 | 1% | 0% | 2 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | ABASIC SITE | 730255 | 11% | 22% | 38 | Search ABASIC+SITE | Search ABASIC+SITE |
| 2 | FLUORESCENCE SELECTIVITY | 177015 | 1% | 100% | 2 | Search FLUORESCENCE+SELECTIVITY | Search FLUORESCENCE+SELECTIVITY |
| 3 | GG MISMATCH | 177015 | 1% | 100% | 2 | Search GG+MISMATCH | Search GG+MISMATCH |
| 4 | INDUCED HAIRPIN FORMATION | 177015 | 1% | 100% | 2 | Search INDUCED+HAIRPIN+FORMATION | Search INDUCED+HAIRPIN+FORMATION |
| 5 | UREA PHENANTHRIDINIUM | 177015 | 1% | 100% | 2 | Search UREA+PHENANTHRIDINIUM | Search UREA+PHENANTHRIDINIUM |
| 6 | NUCLEOBASE RECOGNITION | 132758 | 1% | 50% | 3 | Search NUCLEOBASE+RECOGNITION | Search NUCLEOBASE+RECOGNITION |
| 7 | MISMATCH BINDING LIGAND | 118009 | 1% | 67% | 2 | Search MISMATCH+BINDING+LIGAND | Search MISMATCH+BINDING+LIGAND |
| 8 | NAPHTHYRIDINE DERIVATIVE | 118009 | 1% | 67% | 2 | Search NAPHTHYRIDINE+DERIVATIVE | Search NAPHTHYRIDINE+DERIVATIVE |
| 9 | ABASIC DNA | 113792 | 1% | 43% | 3 | Search ABASIC+DNA | Search ABASIC+DNA |
| 10 | 2 AMINO 7 METHYL 1 8 NAPHTHYRIDINE | 88508 | 0% | 100% | 1 | Search 2+AMINO+7+METHYL+1+8+NAPHTHYRIDINE | Search 2+AMINO+7+METHYL+1+8+NAPHTHYRIDINE |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | LI, MJ , WU, YP , ZHANG, BT , GUO, LH , (2013) DETECTION AND APPLICATION OF DNA ABASIC SITES.CHINESE JOURNAL OF ANALYTICAL CHEMISTRY. VOL. 41. ISSUE 7. P. 965-972 | 34 | 60% | 3 |
| 2 | BENNER, K , BERGEN, A , IHMELS, H , PITHAN, PM , (2014) SELECTIVE STABILIZATION OF ABASIC SITE-CONTAINING DNA BY INSERTION OF STERICALLY DEMANDING BIARYL LIGANDS.CHEMISTRY-A EUROPEAN JOURNAL. VOL. 20. ISSUE 32. P. 9883 -9887 | 32 | 57% | 5 |
| 3 | LHOMME, J , CONSTANT, JF , DEMEUNYNCK, M , (1999) ABASIC DNA STRUCTURE, REACTIVITY, AND RECOGNITION.BIOPOLYMERS. VOL. 52. ISSUE 2. P. 65 -83 | 56 | 41% | 132 |
| 4 | SATO, Y , ZHANG, YS , SEINO, T , SUGIMOTO, T , NISHIZAWA, S , TERAMAE, N , (2012) HIGHLY SELECTIVE BINDING OF NAPHTHYRIDINE WITH A TRIFLUOROMETHYL GROUP TO CYTOSINE OPPOSITE AN ABASIC SITE IN DNA DUPLEXES.ORGANIC & BIOMOLECULAR CHEMISTRY. VOL. 10. ISSUE 20. P. 4003-4006 | 19 | 70% | 8 |
| 5 | AYADI, L , FORGET, D , MARTELLI, A , CONSTANT, JF , DEMEUNYNCK, M , COULOMBEAU, C , (2000) MOLECULAR MODELING STUDY OF DNA ABASIC SITES.THEORETICAL CHEMISTRY ACCOUNTS. VOL. 104. ISSUE 3-4. P. 284 -289 | 26 | 74% | 3 |
| 6 | MALINA, J , SCOTT, P , BRABEC, V , (2015) SHAPE-SELECTIVE RECOGNITION OF DNA ABASIC SITES BY METALLOHELICES: INHIBITION OF HUMAN AP ENDONUCLEASE 1.NUCLEIC ACIDS RESEARCH. VOL. 43. ISSUE 11. P. 5297 -5306 | 21 | 53% | 3 |
| 7 | NISHIZAWA, S , SATO, Y , TERAMAE, N , (2014) RECENT PROGRESS IN ABASIC SITE-BINDING SMALL MOLECULES FOR DETECTING SINGLE-BASE MUTATIONS IN DNA.ANALYTICAL SCIENCES. VOL. 30. ISSUE 1. P. 137 -142 | 28 | 40% | 4 |
| 8 | BENNER, K , IHMELS, H , KOLSCH, S , PITHAN, PM , (2014) TARGETING ABASIC SITE-CONTAINING DNA WITH ANNELATED QUINOLIZINIUM DERIVATIVES: THE INFLUENCE OF SIZE, SHAPE AND SUBSTITUENTS.ORGANIC & BIOMOLECULAR CHEMISTRY. VOL. 12. ISSUE 11. P. 1725 -1734 | 25 | 41% | 9 |
| 9 | SATO, Y , KAGEYAMA, T , TERAMAE, N , NISHIZAWA, S , (2013) COMPETITIVE BINDING OF ABASIC SITE-BINDING LIGANDS AND MASKING LIGANDS TO DNA DUPLEXES FOR THE ANALYSIS OF SINGLE-BASE MUTATION.ANALYTICAL SCIENCES. VOL. 29. ISSUE 1. P. 15-19 | 18 | 58% | 4 |
| 10 | GRECO, NJ , TOR, YZ , (2007) SYNTHESIS AND SITE-SPECIFIC INCORPORATION OF A SIMPLE FLUORESCENT PYRIMIDINE.NATURE PROTOCOLS. VOL. 2. ISSUE 2. P. 305-316 | 32 | 37% | 18 |
Classes with closest relation at Level 1 |