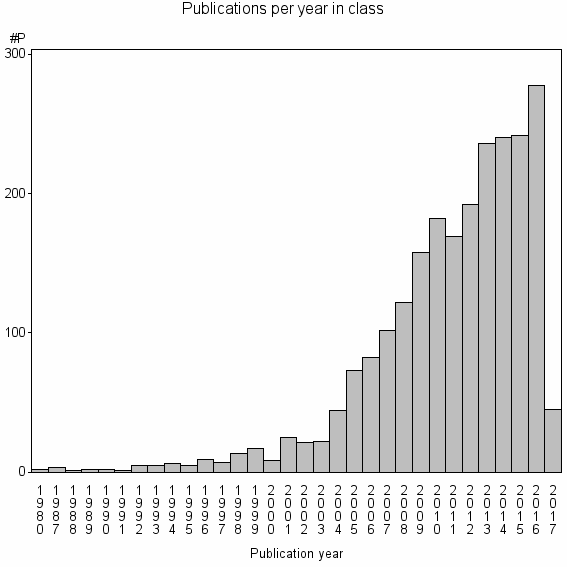
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 2218 | 2319 | 48.6 | 89% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 103 | 3 | O GLCNACYLATION//O GLCNAC TRANSFERASE//O GLCNACASE | 73436 |
| 321 | 2 | N GLYCAN//GLYCOBIOLOGY//GLYCOPROTEOMICS | 18230 |
| 2218 | 1 | GLYCOPROTEOMICS//GLYCOMICS//LECTIN MICROARRAY | 2319 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | GLYCOPROTEOMICS | authKW | 1691573 | 8% | 68% | 189 |
| 2 | GLYCOMICS | authKW | 1044385 | 7% | 48% | 164 |
| 3 | LECTIN MICROARRAY | authKW | 358681 | 2% | 72% | 38 |
| 4 | N GLYCAN | authKW | 296203 | 5% | 18% | 123 |
| 5 | GLYCAN | authKW | 258051 | 5% | 17% | 113 |
| 6 | GLYCAN PROFILING | authKW | 257943 | 1% | 85% | 23 |
| 7 | GLYCOPEPTIDES | authKW | 239658 | 6% | 12% | 148 |
| 8 | DUBLIN OXFORD GLYCOBIOL | address | 230013 | 1% | 56% | 31 |
| 9 | GLYC GLYCOPROTE | address | 193888 | 1% | 82% | 18 |
| 10 | FUCOSYLATION | authKW | 191391 | 2% | 26% | 55 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biochemical Research Methods | 38886 | 43% | 0% | 998 |
| 2 | Chemistry, Analytical | 13176 | 33% | 0% | 759 |
| 3 | Biochemistry & Molecular Biology | 2350 | 29% | 0% | 676 |
| 4 | Spectroscopy | 794 | 6% | 0% | 132 |
| 5 | Mathematical & Computational Biology | 145 | 2% | 0% | 41 |
| 6 | Biophysics | 133 | 4% | 0% | 90 |
| 7 | Medical Laboratory Technology | 106 | 1% | 0% | 32 |
| 8 | Biotechnology & Applied Microbiology | 89 | 4% | 0% | 97 |
| 9 | Oncology | 27 | 4% | 0% | 95 |
| 10 | Chemistry, Multidisciplinary | 18 | 5% | 0% | 121 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | DUBLIN OXFORD GLYCOBIOL | 230013 | 1% | 56% | 31 |
| 2 | GLYC GLYCOPROTE | 193888 | 1% | 82% | 18 |
| 3 | BIOMOL FRONTIERS | 172973 | 1% | 45% | 29 |
| 4 | NIBRT | 155399 | 1% | 54% | 22 |
| 5 | MOL BIOCHEM CLIN INVEST | 152483 | 1% | 35% | 33 |
| 6 | NIBRT GLYCOSCI GRP | 118479 | 1% | 60% | 15 |
| 7 | MED GLYCOSCI | 115047 | 2% | 22% | 40 |
| 8 | FIELD DRUG DISCOVERY | 82417 | 1% | 52% | 12 |
| 9 | NIBRT DUBLIN OXFORD GLYCOBIOL | 77437 | 0% | 59% | 10 |
| 10 | GLYCOBIOTECHNOL | 77346 | 1% | 24% | 24 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | JOURNAL OF PROTEOME RESEARCH | 112221 | 10% | 4% | 228 |
| 2 | MOLECULAR & CELLULAR PROTEOMICS | 41772 | 4% | 3% | 99 |
| 3 | GLYCOBIOLOGY | 34205 | 4% | 3% | 86 |
| 4 | PROTEOMICS | 20225 | 4% | 2% | 94 |
| 5 | CLINICAL PROTEOMICS | 19524 | 1% | 11% | 14 |
| 6 | ANALYTICAL CHEMISTRY | 19424 | 10% | 1% | 231 |
| 7 | GLYCOCONJUGATE JOURNAL | 15367 | 2% | 2% | 48 |
| 8 | PROTEOMICS CLINICAL APPLICATIONS | 8196 | 1% | 3% | 24 |
| 9 | JOURNAL OF PROTEOMICS | 7154 | 2% | 1% | 38 |
| 10 | ELECTROPHORESIS | 4980 | 3% | 1% | 67 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | GLYCOPROTEOMICS | 1691573 | 8% | 68% | 189 | Search GLYCOPROTEOMICS | Search GLYCOPROTEOMICS |
| 2 | GLYCOMICS | 1044385 | 7% | 48% | 164 | Search GLYCOMICS | Search GLYCOMICS |
| 3 | LECTIN MICROARRAY | 358681 | 2% | 72% | 38 | Search LECTIN+MICROARRAY | Search LECTIN+MICROARRAY |
| 4 | N GLYCAN | 296203 | 5% | 18% | 123 | Search N+GLYCAN | Search N+GLYCAN |
| 5 | GLYCAN | 258051 | 5% | 17% | 113 | Search GLYCAN | Search GLYCAN |
| 6 | GLYCAN PROFILING | 257943 | 1% | 85% | 23 | Search GLYCAN+PROFILING | Search GLYCAN+PROFILING |
| 7 | GLYCOPEPTIDES | 239658 | 6% | 12% | 148 | Search GLYCOPEPTIDES | Search GLYCOPEPTIDES |
| 8 | FUCOSYLATION | 191391 | 2% | 26% | 55 | Search FUCOSYLATION | Search FUCOSYLATION |
| 9 | GLYCOPEPTIDE ENRICHMENT | 171154 | 1% | 100% | 13 | Search GLYCOPEPTIDE+ENRICHMENT | Search GLYCOPEPTIDE+ENRICHMENT |
| 10 | GLYCAN ANALYSIS | 165450 | 1% | 47% | 27 | Search GLYCAN+ANALYSIS | Search GLYCAN+ANALYSIS |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | ALLEY, WR , MANN, BF , NOVOTNY, MV , (2013) HIGH-SENSITIVITY ANALYTICAL APPROACHES FOR THE STRUCTURAL CHARACTERIZATION OF GLYCOPROTEINS.CHEMICAL REVIEWS. VOL. 113. ISSUE 4. P. 2668 -2732 | 255 | 43% | 94 |
| 2 | THAYSEN-ANDERSEN, M , PACKER, NH , SCHULZ, BL , (2016) MATURING GLYCOPROTEOMICS TECHNOLOGIES PROVIDE UNIQUE STRUCTURAL INSIGHTS INTO THE N-GLYCOPROTEOME AND ITS REGULATION IN HEALTH AND DISEASE.MOLECULAR & CELLULAR PROTEOMICS. VOL. 15. ISSUE 6. P. 1773 -1790 | 119 | 72% | 6 |
| 3 | GAUNITZ, S , NAGY, G , POHL, NLB , NOYOTNY, MV , (2017) RECENT ADVANCES IN THE ANALYSIS OF COMPLEX GLYCOPROTEINS.ANALYTICAL CHEMISTRY. VOL. 89. ISSUE 1. P. 389 -413 | 159 | 68% | 0 |
| 4 | THAYSEN-ANDERSEN, M , PACKER, NH , (2014) ADVANCES IN LC-MS/MS-BASED GLYCOPROTEOMICS: GETTING CLOSER TO SYSTEM-WIDE SITE-SPECIFIC MAPPING OF THE N- AND O-GLYCOPROTEOME.BIOCHIMICA ET BIOPHYSICA ACTA-PROTEINS AND PROTEOMICS. VOL. 1844. ISSUE 9. P. 1437 -1452 | 110 | 71% | 48 |
| 5 | BOICHENKO, A , GOVORUKHINA, N , BISCHOFF, R , ONGAY, S , (2012) GLYCOPEPTIDE ENRICHMENT AND SEPARATION FOR PROTEIN GLYCOSYLATION ANALYSIS.JOURNAL OF SEPARATION SCIENCE. VOL. 35. ISSUE 18. P. 2341 -2372 | 140 | 66% | 45 |
| 6 | AHN, YH , KIM, JY , YOO, JS , (2015) QUANTITATIVE MASS SPECTROMETRIC ANALYSIS OF GLYCOPROTEINS COMBINED WITH ENRICHMENT METHODS.MASS SPECTROMETRY REVIEWS. VOL. 34. ISSUE 2. P. 148 -165 | 104 | 78% | 9 |
| 7 | LI, LJ , FROST, DC , (2014) RECENT ADVANCES IN MASS SPECTROMETRY-BASED GLYCOPROTEOMICS.PROTEOMICS IN BIOMEDICINE AND PHARMACOLOGY. VOL. 95. ISSUE . P. 71 -+ | 139 | 62% | 5 |
| 8 | BANAZADEH, A , VEILLON, L , WOODING, KM , ZABET-MOGHADDAM, M , MECHREF, Y , (2017) RECENT ADVANCES IN MASS SPECTROMETRIC ANALYSIS OF GLYCOPROTEINS.ELECTROPHORESIS. VOL. 38. ISSUE 1. P. 162 -189 | 100 | 78% | 0 |
| 9 | NOVOTNY, MV , ALLEY, WR , MANN, BF , (2013) ANALYTICAL GLYCOBIOLOGY AT HIGH SENSITIVITY: CURRENT APPROACHES AND DIRECTIONS.GLYCOCONJUGATE JOURNAL. VOL. 30. ISSUE 2. P. 89 -117 | 124 | 58% | 23 |
| 10 | LAZAR, IM , DENG, JR , IKENISHI, F , LAZAR, AC , (2015) EXPLORING THE GLYCOPROTEOMICS LANDSCAPE WITH ADVANCED MS TECHNOLOGIES.ELECTROPHORESIS. VOL. 36. ISSUE 1. P. 225 -237 | 79 | 82% | 9 |
Classes with closest relation at Level 1 |