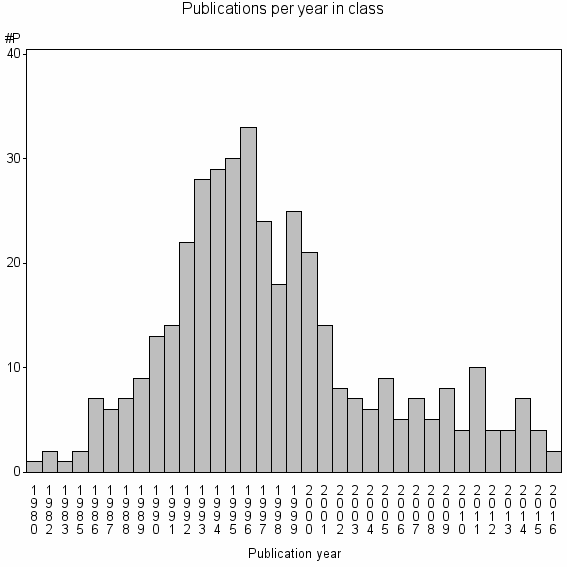
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 22039 | 396 | 33.9 | 84% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 0 | 4 | BIOCHEMISTRY & MOLECULAR BIOLOGY//CELL BIOLOGY//ONCOLOGY | 4064930 |
| 590 | 3 | TISSUE ANTIGENS//IMMUNOGENETICS//HUMAN IMMUNOLOGY | 15170 |
| 588 | 2 | TISSUE ANTIGENS//IMMUNOGENETICS//HUMAN IMMUNOLOGY | 14187 |
| 22039 | 1 | MOL IMMUNOL RUMENTAT//GENOMIC MATCHING TECHNIQUE//HAPLOSPECIFIC MARKERS | 396 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | MOL IMMUNOL RUMENTAT | address | 1019743 | 6% | 58% | 23 |
| 2 | GENOMIC MATCHING TECHNIQUE | authKW | 385543 | 1% | 100% | 5 |
| 3 | HAPLOSPECIFIC MARKERS | authKW | 231326 | 1% | 100% | 3 |
| 4 | LST1 | authKW | 214186 | 1% | 56% | 5 |
| 5 | MHC CLASS III REGION | authKW | 205620 | 1% | 67% | 4 |
| 6 | BIOINFORMAT BIOL COMP | address | 205604 | 3% | 22% | 12 |
| 7 | CR1 L | authKW | 154217 | 1% | 100% | 2 |
| 8 | G7C | authKW | 154217 | 1% | 100% | 2 |
| 9 | POALIN | authKW | 154217 | 1% | 100% | 2 |
| 10 | POLYMORPHIC FROZEN BLOCKS | authKW | 154217 | 1% | 100% | 2 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Genetics & Heredity | 7131 | 55% | 0% | 219 |
| 2 | Immunology | 4431 | 51% | 0% | 200 |
| 3 | Biotechnology & Applied Microbiology | 419 | 15% | 0% | 59 |
| 4 | Pathology | 278 | 8% | 0% | 32 |
| 5 | Cell Biology | 215 | 14% | 0% | 54 |
| 6 | Biochemistry & Molecular Biology | 183 | 21% | 0% | 85 |
| 7 | Medicine, Research & Experimental | 69 | 6% | 0% | 24 |
| 8 | Evolutionary Biology | 63 | 3% | 0% | 12 |
| 9 | Transplantation | 17 | 2% | 0% | 7 |
| 10 | Medicine, Legal | 15 | 1% | 0% | 3 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | MOL IMMUNOL RUMENTAT | 1019743 | 6% | 58% | 23 |
| 2 | BIOINFORMAT BIOL COMP | 205604 | 3% | 22% | 12 |
| 3 | GENET INFORMAT | 92565 | 6% | 5% | 22 |
| 4 | BIOCHEM ANDMOLECULAR BIOL | 77109 | 0% | 100% | 1 |
| 5 | BIOCHEM IMMUNOCHEM UNIT MRC | 77109 | 0% | 100% | 1 |
| 6 | CIGH CNRS UPR 8291 | 77109 | 0% | 100% | 1 |
| 7 | CNRS RECOMBINAISONS GENET | 77109 | 0% | 100% | 1 |
| 8 | CNRS SD 401 | 77109 | 0% | 100% | 1 |
| 9 | CYO CONNOR ERADE VILLAGE | 77109 | 0% | 100% | 1 |
| 10 | GENNET HUMANA | 77109 | 0% | 100% | 1 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | IMMUNOGENETICS | 97563 | 19% | 2% | 75 |
| 2 | EUROPEAN JOURNAL OF IMMUNOGENETICS | 19734 | 3% | 2% | 12 |
| 3 | TISSUE ANTIGENS | 16605 | 8% | 1% | 31 |
| 4 | GENOMICS | 13459 | 10% | 0% | 38 |
| 5 | HUMAN IMMUNOLOGY | 10395 | 6% | 1% | 25 |
| 6 | DNA SEQUENCE | 6764 | 2% | 1% | 9 |
| 7 | JOURNAL OF MOLECULAR EVOLUTION | 2898 | 3% | 0% | 12 |
| 8 | M S-MEDECINE SCIENCES | 2787 | 3% | 0% | 11 |
| 9 | MAMMALIAN GENOME | 2644 | 3% | 0% | 11 |
| 10 | EXPERIMENTAL AND CLINICAL IMMUNOGENETICS | 1519 | 1% | 1% | 3 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | GENOMIC MATCHING TECHNIQUE | 385543 | 1% | 100% | 5 | Search GENOMIC+MATCHING+TECHNIQUE | Search GENOMIC+MATCHING+TECHNIQUE |
| 2 | HAPLOSPECIFIC MARKERS | 231326 | 1% | 100% | 3 | Search HAPLOSPECIFIC+MARKERS | Search HAPLOSPECIFIC+MARKERS |
| 3 | LST1 | 214186 | 1% | 56% | 5 | Search LST1 | Search LST1 |
| 4 | MHC CLASS III REGION | 205620 | 1% | 67% | 4 | Search MHC+CLASS+III+REGION | Search MHC+CLASS+III+REGION |
| 5 | CR1 L | 154217 | 1% | 100% | 2 | Search CR1+L | Search CR1+L |
| 6 | G7C | 154217 | 1% | 100% | 2 | Search G7C | Search G7C |
| 7 | POALIN | 154217 | 1% | 100% | 2 | Search POALIN | Search POALIN |
| 8 | POLYMORPHIC FROZEN BLOCKS | 154217 | 1% | 100% | 2 | Search POLYMORPHIC+FROZEN+BLOCKS | Search POLYMORPHIC+FROZEN+BLOCKS |
| 9 | PROGENITOR HAPLOTYPES | 154217 | 1% | 100% | 2 | Search PROGENITOR+HAPLOTYPES | Search PROGENITOR+HAPLOTYPES |
| 10 | HLA A ALLELES | 154213 | 1% | 50% | 4 | Search HLA+A+ALLELES | Search HLA+A+ALLELES |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | SNOEK, M , TEUSCHER, C , VAN VUGT, H , (1998) MOLECULAR ANALYSIS OF THE MAJOR MHC RECOMBINATIONAL HOT SPOT LOCATED WITHIN THE G7C GENE OF THE MURINE CLASS III REGION THAT IS INVOLVED IN DISEASE SUSCEPTIBILITY.JOURNAL OF IMMUNOLOGY. VOL. 160. ISSUE 1. P. 266-272 | 32 | 76% | 20 |
| 2 | MCLURE, CA , HINCHLIFFE, P , LESTER, S , WILLIAMSON, JF , MILLMAN, JA , KEATING, PJ , STEWART, BJ , DAWKINS, RL , (2013) GENOMIC EVOLUTION AND POLYMORPHISM: SEGMENTAL DUPLICATIONS AND HAPLOTYPES AT 108 REGIONS ON 21 CHROMOSOMES.GENOMICS. VOL. 102. ISSUE 1. P. 15-26 | 24 | 56% | 4 |
| 3 | KULSKI, JK , GAUDIERI, S , DAWKINS, RL , (2000) USING ALU J ELEMENTS AS MOLECULAR CLOCKS TO TRACE THE EVOLUTIONARY RELATIONSHIPS BETWEEN DUPLICATED HLA CLASS I GENOMIC SEGMENTS.JOURNAL OF MOLECULAR EVOLUTION. VOL. 50. ISSUE 6. P. 510 -519 | 26 | 74% | 11 |
| 4 | SHI, L , KULSKI, JK , ZHANG, H , DONG, ZM , CAO, DF , ZHOU, JX , YU, JK , YAO, YF , SHI, L , (2014) ASSOCIATION AND DIFFERENTIATION OF MHC CLASS I AND II POLYMORPHIC ALU INSERTIONS AND HLA-A, -B, -C AND-DRB1 ALLELES IN THE CHINESE HAN POPULATION.MOLECULAR GENETICS AND GENOMICS. VOL. 289. ISSUE 1. P. 93-101 | 16 | 57% | 1 |
| 5 | PICHON, L , GIFFON, T , CHAUVEL, B , LEGALL, JY , DAVID, V , (1996) THE CLASS I REGION OF THE MHC GENES IS ONE OF THE MOST COMPLEX IN THE WHOLE HUMAN GENOME.M S-MEDECINE SCIENCES. VOL. 12. ISSUE 11. P. 1209-1218 | 23 | 66% | 2 |
| 6 | SNOEK, M , VANDINTEN, L , VANVUGT, H , (1996) A NOVEL GENE, G7E, RESEMBLING A VIRAL ENVELOPE GENE, IS LOCATED AT THE RECOMBINATIONAL HOT SPOT IN THE CLASS III REGION OF THE MOUSE MHC.GENOMICS. VOL. 38. ISSUE 1. P. 5-12 | 22 | 67% | 10 |
| 7 | KULSKI, JK , ANZAI, T , INOKO, H , (2005) ERVK9, TRANSPOSONS AND THE EVOLUTION OF MHC CLASS I DUPLICONS WITHIN THE ALPHA-BLOCK OF THE HUMAN AND CHIMPANZEE.CYTOGENETIC AND GENOME RESEARCH. VOL. 110. ISSUE 1-4. P. 181-192 | 24 | 45% | 11 |
| 8 | CULLEN, M , NOBLE, J , ERLICH, H , THORPE, K , BECK, S , KLITZ, W , TROWSDALE, J , CARRINGTON, M , (1997) CHARACTERIZATION OF RECOMBINATION IN THE HLA CLASS II REGION.AMERICAN JOURNAL OF HUMAN GENETICS. VOL. 60. ISSUE 2. P. 397-407 | 29 | 43% | 84 |
| 9 | SHIINA, T , TAMIYA, G , OKA, A , TAKISHIMA, N , INOKO, H , (1999) GENOME SEQUENCING ANALYSIS OF THE 1.8 MB ENTIRE HUMAN MHC CLASS I REGION.IMMUNOLOGICAL REVIEWS. VOL. 167. ISSUE . P. 193 -199 | 24 | 50% | 37 |
| 10 | STEELE, EJ , (2014) REFLECTIONS ON ANCESTRAL HAPLOTYPES MEDICAL GENOMICS, EVOLUTION, AND HUMAN INDIVIDUALITY.PERSPECTIVES IN BIOLOGY AND MEDICINE. VOL. 57. ISSUE 2. P. 179 -197 | 22 | 37% | 0 |
Classes with closest relation at Level 1 |