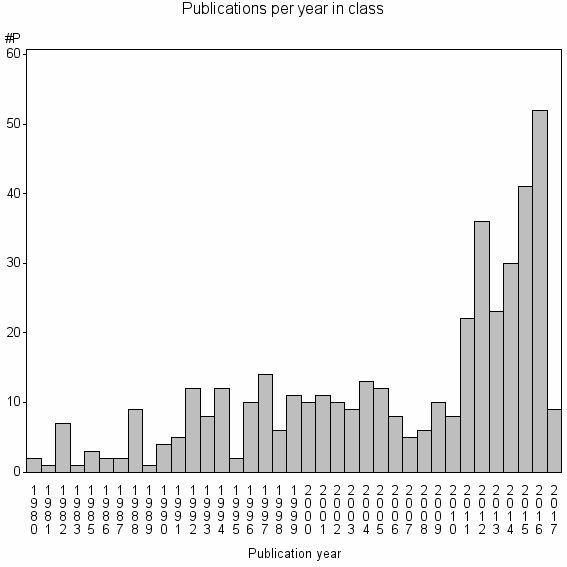
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 21258 | 427 | 40.1 | 76% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 3 | 4 | PLANT SCIENCES//AGRONOMY//FOOD SCIENCE & TECHNOLOGY | 1882812 |
| 128 | 3 | BIOTECHNOLOGY & APPLIED MICROBIOLOGY//CELLULASE//XYLANASE | 67277 |
| 331 | 2 | CELLULASE//XYLANASE//ENZYMATIC HYDROLYSIS | 18094 |
| 21258 | 1 | CELLOBIOSE DEHYDROGENASE//LYTIC POLYSACCHARIDE MONOOXYGENASE//AA9 | 427 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | CELLOBIOSE DEHYDROGENASE | authKW | 6529766 | 29% | 75% | 122 |
| 2 | LYTIC POLYSACCHARIDE MONOOXYGENASE | authKW | 1681643 | 6% | 87% | 27 |
| 3 | AA9 | authKW | 1001147 | 3% | 100% | 14 |
| 4 | GH61 | authKW | 700794 | 3% | 70% | 14 |
| 5 | CBM33 | authKW | 482691 | 2% | 75% | 9 |
| 6 | LPMO | authKW | 476730 | 2% | 67% | 10 |
| 7 | CELLOBIOSE OXIDASE | authKW | 457664 | 2% | 80% | 8 |
| 8 | ANALYT CHEM BIOCHEM STRUCT BIOL | address | 446923 | 4% | 42% | 15 |
| 9 | LYTIC POLYSACCHARIDE MONOOXYGENASE LPMO | authKW | 357552 | 1% | 100% | 5 |
| 10 | AA10 | authKW | 350397 | 2% | 70% | 7 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Biotechnology & Applied Microbiology | 3478 | 38% | 0% | 164 |
| 2 | Biochemistry & Molecular Biology | 924 | 40% | 0% | 171 |
| 3 | Biophysics | 571 | 14% | 0% | 60 |
| 4 | Microbiology | 335 | 13% | 0% | 55 |
| 5 | Biochemical Research Methods | 207 | 8% | 0% | 35 |
| 6 | Materials Science, Paper & Wood | 152 | 2% | 0% | 10 |
| 7 | Electrochemistry | 137 | 5% | 0% | 23 |
| 8 | Chemistry, Analytical | 114 | 8% | 0% | 36 |
| 9 | Energy & Fuels | 85 | 5% | 0% | 23 |
| 10 | Mycology | 58 | 2% | 0% | 7 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | ANALYT CHEM BIOCHEM STRUCT BIOL | 446923 | 4% | 42% | 15 |
| 2 | BIO ERS BIOTECHNOL FONG UMR1163 | 297959 | 1% | 83% | 5 |
| 3 | BIOCHEM UNIV WALK | 143021 | 0% | 100% | 2 |
| 4 | BIOFUELS TECHNOL | 143021 | 0% | 100% | 2 |
| 5 | PLATEFORME BIBS | 143021 | 0% | 100% | 2 |
| 6 | UMR BIO ERS BIOTECHNOL FONG 1163 | 143021 | 0% | 100% | 2 |
| 7 | CHEM BIOTECHNOL FOOD SCI | 99895 | 9% | 4% | 39 |
| 8 | CELL MOL BIOL STRUCT BIOL | 95346 | 0% | 67% | 2 |
| 9 | ESIL POLYTECH MARSEILLE | 95346 | 0% | 67% | 2 |
| 10 | FOOD BIOTECHNOL | 74194 | 11% | 2% | 46 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BIOTECHNOLOGY FOR BIOFUELS | 14388 | 4% | 1% | 15 |
| 2 | APPLIED MICROBIOLOGY AND BIOTECHNOLOGY | 2040 | 5% | 0% | 20 |
| 3 | ENZYME AND MICROBIAL TECHNOLOGY | 2021 | 3% | 0% | 13 |
| 4 | BIOTECHNOLOGY JOURNAL | 1807 | 1% | 1% | 5 |
| 5 | BIOELECTROCHEMISTRY | 1773 | 1% | 0% | 6 |
| 6 | APPLIED AND ENVIRONMENTAL MICROBIOLOGY | 932 | 5% | 0% | 20 |
| 7 | BIOMOLECULAR NMR ASSIGNMENTS | 916 | 1% | 0% | 3 |
| 8 | APPLIED BIOCHEMISTRY AND MICROBIOLOGY | 758 | 1% | 0% | 5 |
| 9 | CHEMIKER-ZEITUNG | 736 | 1% | 0% | 3 |
| 10 | EUROPEAN JOURNAL OF APPLIED MICROBIOLOGY AND BIOTECHNOLOGY | 576 | 0% | 0% | 2 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | CELLOBIOSE DEHYDROGENASE | 6529766 | 29% | 75% | 122 | Search CELLOBIOSE+DEHYDROGENASE | Search CELLOBIOSE+DEHYDROGENASE |
| 2 | LYTIC POLYSACCHARIDE MONOOXYGENASE | 1681643 | 6% | 87% | 27 | Search LYTIC+POLYSACCHARIDE+MONOOXYGENASE | Search LYTIC+POLYSACCHARIDE+MONOOXYGENASE |
| 3 | AA9 | 1001147 | 3% | 100% | 14 | Search AA9 | Search AA9 |
| 4 | GH61 | 700794 | 3% | 70% | 14 | Search GH61 | Search GH61 |
| 5 | CBM33 | 482691 | 2% | 75% | 9 | Search CBM33 | Search CBM33 |
| 6 | LPMO | 476730 | 2% | 67% | 10 | Search LPMO | Search LPMO |
| 7 | CELLOBIOSE OXIDASE | 457664 | 2% | 80% | 8 | Search CELLOBIOSE+OXIDASE | Search CELLOBIOSE+OXIDASE |
| 8 | LYTIC POLYSACCHARIDE MONOOXYGENASE LPMO | 357552 | 1% | 100% | 5 | Search LYTIC+POLYSACCHARIDE+MONOOXYGENASE+LPMO | Search LYTIC+POLYSACCHARIDE+MONOOXYGENASE+LPMO |
| 9 | AA10 | 350397 | 2% | 70% | 7 | Search AA10 | Search AA10 |
| 10 | CELLOBIONIC ACID | 228832 | 1% | 80% | 4 | Search CELLOBIONIC+ACID | Search CELLOBIONIC+ACID |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | ZAMOCKY, M , LUDWIG, R , PETERBAUER, C , HALLBERG, BM , DIVNE, C , NICHOLLS, P , HALTRICH, D , (2006) CELLOBIOSE DEHYDROGENASE - A FLAVOCYTOCHROME FROM WOOD-DEGRADING, PHYTOPATHOGENIC AND SAPROTROPIC FUNGI.CURRENT PROTEIN & PEPTIDE SCIENCE. VOL. 7. ISSUE 3. P. 255 -280 | 85 | 67% | 110 |
| 2 | HEMSWORTH, GR , JOHNSTON, EM , DAVIES, GJ , WALTON, PH , (2015) LYTIC POLYSACCHARIDE MONOOXYGENASES IN BIOMASS CONVERSION.TRENDS IN BIOTECHNOLOGY. VOL. 33. ISSUE 12. P. 747 -761 | 56 | 60% | 23 |
| 3 | HENRIKSSON, G , JOHANSSON, G , PETTERSSON, G , (2000) A CRITICAL REVIEW OF CELLOBIOSE DEHYDROGENASES.JOURNAL OF BIOTECHNOLOGY. VOL. 78. ISSUE 2. P. 93 -113 | 63 | 76% | 235 |
| 4 | LOOSE, JSM , FORSBERG, Z , KRACHER, D , SCHEIBLBRANDNER, S , LUDWIG, R , EIJSINK, VGH , VAAJE-KOLSTAD, G , (2016) ACTIVATION OF BACTERIAL LYTIC POLYSACCHARIDE MONOOXYGENASES WITH CELLOBIOSE DEHYDROGENASE.PROTEIN SCIENCE. VOL. 25. ISSUE 12. P. 2175 -2186 | 46 | 85% | 0 |
| 5 | COURTADE, G , WIMMER, R , ROHR, AK , PREIMS, M , FELICE, AKG , DIMAROGONA, M , VAAJE-KOLSTAD, G , SORLIE, M , SANDGREN, M , LUDWIG, R , ET AL (2016) INTERACTIONS OF A FUNGAL LYTIC POLYSACCHARIDE MONOOXYGENASE WITH BETA-GLUCAN SUBSTRATES AND CELLOBIOSE DEHYDROGENASE.PROCEEDINGS OF THE NATIONAL ACADEMY OF SCIENCES OF THE UNITED STATES OF AMERICA. VOL. 113. ISSUE 21. P. 5922 -5927 | 34 | 71% | 6 |
| 6 | CORREA, TLR , DOS SANTOS, LV , PEREIRA, GAG , (2016) AA9 AND AA10: FROM ENIGMATIC TO ESSENTIAL ENZYMES.APPLIED MICROBIOLOGY AND BIOTECHNOLOGY. VOL. 100. ISSUE 1. P. 9 -16 | 37 | 74% | 3 |
| 7 | WALTON, PH , DAVIES, GJ , (2016) ON THE CATALYTIC MECHANISMS OF LYTIC POLYSACCHARIDE MONOOXYGENASES.CURRENT OPINION IN CHEMICAL BIOLOGY. VOL. 31. ISSUE . P. 195 -207 | 34 | 79% | 3 |
| 8 | FRANDSEN, KEH , LO LEGGIO, L , (2016) LYTIC POLYSACCHARIDE MONOOXYGENASES: A CRYSTALLOGRAPHER'S VIEW ON A NEW CLASS OF BIOMASS-DEGRADING ENZYMES.IUCRJ. VOL. 3. ISSUE . P. 448 -467 | 59 | 54% | 0 |
| 9 | GARAJOVA, S , MATHIEU, Y , BECCIA, MR , BENNATI-GRANIER, C , BIASO, F , FANUEL, M , ROPARTZ, D , GUIGLIARELLI, B , RECORD, E , ROGNIAUX, H , ET AL (2016) SINGLE-DOMAIN FLAVOENZYMES TRIGGER LYTIC POLYSACCHARIDE MONOOXYGENASES FOR OXIDATIVE DEGRADATION OF CELLULOSE.SCIENTIFIC REPORTS. VOL. 6. ISSUE . P. - | 32 | 76% | 4 |
| 10 | ZHANG, RF , FAN, ZL , KASUGA, T , (2011) EXPRESSION OF CELLOBIOSE DEHYDROGENASE FROM NEUROSPORA CRASSA IN PICHIA PASTORIS AND ITS PURIFICATION AND CHARACTERIZATION.PROTEIN EXPRESSION AND PURIFICATION. VOL. 75. ISSUE 1. P. 63-69 | 34 | 92% | 12 |
Classes with closest relation at Level 1 |