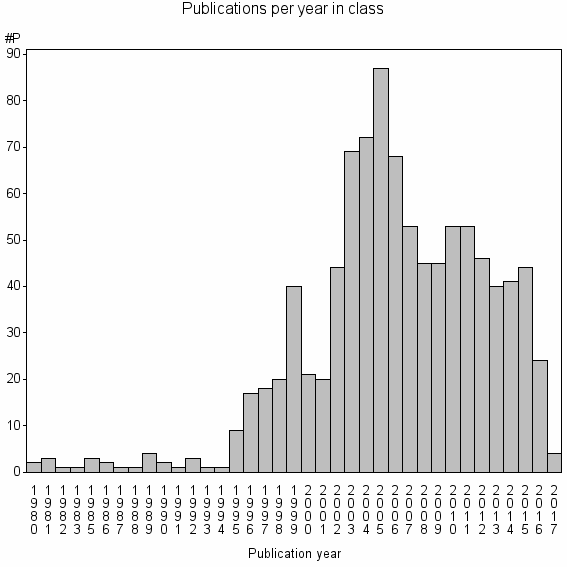
Class information for: |
Basic class information |
| Class id | #P | Avg. number of references |
Database coverage of references |
|---|---|---|---|
| 11719 | 959 | 20.9 | 49% |
Hierarchy of classes |
The table includes all classes above and classes immediately below the current class. |
| Cluster id | Level | Cluster label | #P |
|---|---|---|---|
| 9 | 4 | COMPUTER SCIENCE, THEORY & METHODS//COMPUTER SCIENCE, ARTIFICIAL INTELLIGENCE//COMPUTER SCIENCE, INFORMATION SYSTEMS | 1247339 |
| 621 | 3 | MEMBRANE COMPUTING//THEORETICAL COMPUTER SCIENCE//LOGIC | 13680 |
| 1059 | 2 | MEMBRANE COMPUTING//THEORETICAL COMPUTER SCIENCE//P SYSTEMS | 10031 |
| 11719 | 1 | DNA COMPUTING//DNA COMPUTATION//SPLICING SYSTEMS | 959 |
Terms with highest relevance score |
| rank | Term | termType | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|---|
| 1 | DNA COMPUTING | authKW | 4429240 | 22% | 66% | 210 |
| 2 | DNA COMPUTATION | authKW | 877381 | 4% | 67% | 41 |
| 3 | SPLICING SYSTEMS | authKW | 868340 | 3% | 91% | 30 |
| 4 | ADLEMAN LIPTON MODEL | authKW | 509429 | 2% | 100% | 16 |
| 5 | DNA BASED COMPUTING | authKW | 447739 | 2% | 94% | 15 |
| 6 | MOLECULAR COMPUTING | authKW | 445694 | 4% | 33% | 42 |
| 7 | STICKER MODEL | authKW | 390028 | 1% | 88% | 14 |
| 8 | EVOLUTIONARY PROCESSOR | authKW | 382072 | 1% | 100% | 12 |
| 9 | NP COMPLETE PROBLEM | authKW | 366821 | 4% | 30% | 39 |
| 10 | HAIRPIN COMPLETION | authKW | 318393 | 1% | 100% | 10 |
Web of Science journal categories |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | Computer Science, Theory & Methods | 16802 | 36% | 0% | 350 |
| 2 | Mathematical & Computational Biology | 9332 | 18% | 0% | 175 |
| 3 | Computer Science, Artificial Intelligence | 1974 | 12% | 0% | 118 |
| 4 | Biology | 907 | 9% | 0% | 89 |
| 5 | Computer Science, Information Systems | 714 | 8% | 0% | 73 |
| 6 | Mathematics, Applied | 691 | 12% | 0% | 113 |
| 7 | Computer Science, Interdisciplinary Applications | 542 | 7% | 0% | 67 |
| 8 | Computer Science, Software Engineering | 482 | 6% | 0% | 54 |
| 9 | Biochemical Research Methods | 299 | 7% | 0% | 65 |
| 10 | Computer Science, Hardware & Architecture | 148 | 3% | 0% | 24 |
Address terms |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | PL DNA COMP | 289447 | 1% | 91% | 10 |
| 2 | GRP MATH LINGUIST | 277574 | 4% | 24% | 37 |
| 3 | BIOMOL COMP | 88438 | 1% | 56% | 5 |
| 4 | AFFAIR OFF | 79593 | 1% | 50% | 5 |
| 5 | ADV DESIGN INTELLIGENT COMP | 76149 | 2% | 15% | 16 |
| 6 | BIO X DNA COMP CONSORTIUM | 72772 | 0% | 57% | 4 |
| 7 | CHINA DIGITAL EARTH | 63679 | 0% | 100% | 2 |
| 8 | LEIDEN NAT COMP | 63679 | 0% | 100% | 2 |
| 9 | MOL COMP GRP | 63679 | 0% | 100% | 2 |
| 10 | COMPUTAT BIOMODELLING | 57308 | 0% | 60% | 3 |
Journals |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
|---|---|---|---|---|---|
| 1 | BIOSYSTEMS | 64431 | 8% | 3% | 74 |
| 2 | NATURAL COMPUTING | 42646 | 2% | 6% | 22 |
| 3 | JOURNAL OF COMPUTATIONAL AND THEORETICAL NANOSCIENCE | 20218 | 5% | 1% | 45 |
| 4 | IEEE TRANSACTIONS ON NANOBIOSCIENCE | 20162 | 2% | 3% | 21 |
| 5 | INTERNATIONAL JOURNAL OF FOUNDATIONS OF COMPUTER SCIENCE | 13424 | 2% | 2% | 20 |
| 6 | THEORETICAL COMPUTER SCIENCE | 13266 | 7% | 1% | 67 |
| 7 | INTERNATIONAL JOURNAL OF UNCONVENTIONAL COMPUTING | 10864 | 1% | 4% | 9 |
| 8 | LECTURE NOTES IN COMPUTER SCIENCE | 9413 | 17% | 0% | 163 |
| 9 | FUNDAMENTA INFORMATICAE | 9401 | 3% | 1% | 26 |
| 10 | INTERNATIONAL JOURNAL OF INNOVATIVE COMPUTING INFORMATION AND CONTROL | 5974 | 2% | 1% | 22 |
Author Key Words |
| Rank | Term | Chi square | Shr. of publ. in class containing term |
Class's shr. of term's tot. occurrences |
#P with term in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | DNA COMPUTING | 4429240 | 22% | 66% | 210 | Search DNA+COMPUTING | Search DNA+COMPUTING |
| 2 | DNA COMPUTATION | 877381 | 4% | 67% | 41 | Search DNA+COMPUTATION | Search DNA+COMPUTATION |
| 3 | SPLICING SYSTEMS | 868340 | 3% | 91% | 30 | Search SPLICING+SYSTEMS | Search SPLICING+SYSTEMS |
| 4 | ADLEMAN LIPTON MODEL | 509429 | 2% | 100% | 16 | Search ADLEMAN+LIPTON+MODEL | Search ADLEMAN+LIPTON+MODEL |
| 5 | DNA BASED COMPUTING | 447739 | 2% | 94% | 15 | Search DNA+BASED+COMPUTING | Search DNA+BASED+COMPUTING |
| 6 | MOLECULAR COMPUTING | 445694 | 4% | 33% | 42 | Search MOLECULAR+COMPUTING | Search MOLECULAR+COMPUTING |
| 7 | STICKER MODEL | 390028 | 1% | 88% | 14 | Search STICKER+MODEL | Search STICKER+MODEL |
| 8 | EVOLUTIONARY PROCESSOR | 382072 | 1% | 100% | 12 | Search EVOLUTIONARY+PROCESSOR | Search EVOLUTIONARY+PROCESSOR |
| 9 | NP COMPLETE PROBLEM | 366821 | 4% | 30% | 39 | Search NP+COMPLETE+PROBLEM | Search NP+COMPLETE+PROBLEM |
| 10 | HAIRPIN COMPLETION | 318393 | 1% | 100% | 10 | Search HAIRPIN+COMPLETION | Search HAIRPIN+COMPLETION |
Core articles |
The table includes core articles in the class. The following variables is taken into account for the relevance score of an article in a cluster c: (1) Number of references referring to publications in the class. (2) Share of total number of active references referring to publications in the class. (3) Age of the article. New articles get higher score than old articles. (4) Citation rate, normalized to year. |
| Rank | Reference | # ref. in cl. |
Shr. of ref. in cl. |
Citations |
|---|---|---|---|---|
| 1 | XU, J , TAN, GJ , (2007) A REVIEW ON DNA COMPUTING MODELS.JOURNAL OF COMPUTATIONAL AND THEORETICAL NANOSCIENCE. VOL. 4. ISSUE 7-8. P. 1219 -1230 | 31 | 89% | 15 |
| 2 | WANG, ZC , TAN, J , HUANG, DM , REN, YC , JI, ZW , (2014) A BIOLOGICAL ALGORITHM TO SOLVE THE ASSIGNMENT PROBLEM BASED ON DNA MOLECULES COMPUTATION.APPLIED MATHEMATICS AND COMPUTATION. VOL. 244. ISSUE . P. 183 -190 | 23 | 96% | 4 |
| 3 | SANCHES, CAA , SOMA, NY , (2016) A GENERAL RESOLUTION OF INTRACTABLE PROBLEMS IN POLYNOMIAL TIME THROUGH DNA COMPUTING.BIOSYSTEMS. VOL. 150. ISSUE . P. 119 -131 | 20 | 95% | 0 |
| 4 | WANG, ZC , HUANG, DM , TAN, J , LIU, TF , ZHAO, K , LI, L , (2015) A PARALLEL ALGORITHM FOR SOLVING THE N-QUEENS PROBLEM BASED ON INSPIRED COMPUTATIONAL MODEL.BIOSYSTEMS. VOL. 131. ISSUE . P. 22 -29 | 25 | 76% | 1 |
| 5 | NING, JG , LI, YM , YU, W , (2015) MOLECULAR STICKER MODEL STIMULATION ON SILICON FOR A MAXIMUM CLIQUE PROBLEM.INTERNATIONAL JOURNAL OF MOLECULAR SCIENCES. VOL. 16. ISSUE 6. P. 13474 -13489 | 22 | 85% | 0 |
| 6 | WANG, ZC , HUANG, DM , MENG, HJ , TANG, CP , (2013) A NEW FAST ALGORITHM FOR SOLVING THE MINIMUM SPANNING TREE PROBLEM BASED ON DNA MOLECULES COMPUTATION.BIOSYSTEMS. VOL. 114. ISSUE 1. P. 1-7 | 21 | 91% | 1 |
| 7 | WANG, ZC , QIN, JF , JI, ZW , HUANG, DM , LI, L , (2015) SOLVING THE MAXIMUM WEIGHTED CLIQUE PROBLEM BASED ON PARALLEL BIOLOGICAL COMPUTING MODEL.MATHEMATICAL PROBLEMS IN ENGINEERING. VOL. . ISSUE . P. - | 19 | 95% | 0 |
| 8 | WANG, ZC , PU, J , CAO, LL , TAN, J , (2015) A PARALLEL BIOLOGICAL OPTIMIZATION ALGORITHM TO SOLVE THE UNBALANCED ASSIGNMENT PROBLEM BASED ON DNA MOLECULAR COMPUTING.INTERNATIONAL JOURNAL OF MOLECULAR SCIENCES. VOL. 16. ISSUE 10. P. 25338 -25352 | 21 | 84% | 0 |
| 9 | WANG, ZC , JI, ZW , SU, ZY , ZHAO, K , WANG, XM , (2016) SOLVING THE MAXIMAL MATCHING PROBLEM WITH DNA MOLECULES IN ADLEMAN-LIPTON MODEL.INTERNATIONAL JOURNAL OF BIOMATHEMATICS. VOL. 9. ISSUE 2. P. - | 19 | 86% | 0 |
| 10 | SHAO, XG , JIANG, HY , CAI, WS , (2002) ADVANCES IN BIOMOLECULAR COMPUTING.PROGRESS IN CHEMISTRY. VOL. 14. ISSUE 1. P. 37-46 | 24 | 89% | 30 |
Classes with closest relation at Level 1 |