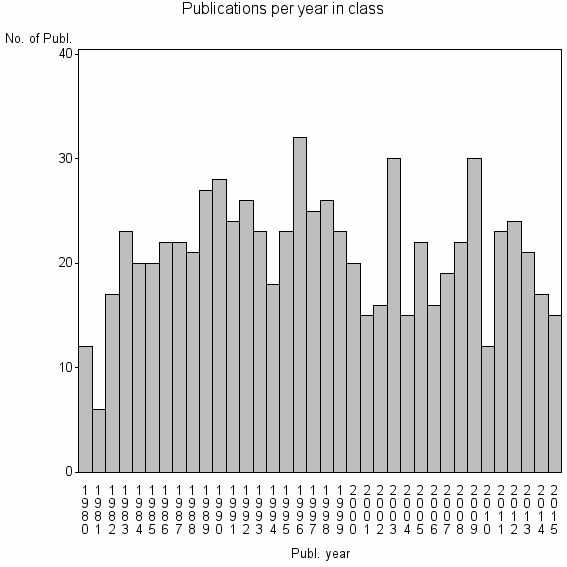
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 3711 | 755 | 32.1 | 60% |
Classes in level above (level 3) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 465 | 22192 | NM23//RIBONUCLEOTIDE REDUCTASE//DEOXYCYTIDINE KINASE |
Classes in level below (level 1) |
| ID, lev. below | Publications | Label for level below |
|---|---|---|
| 14796 | 680 | ADENYLATE KINASE//THYMIDYLATE KINASE//CHIM STRUCT MACROMOL |
| 34228 | 75 | CONDITIONAL MEAN AND VARIANCE//MACRO DATA//SPECTRAL LIKELIHOOD |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | ADENYLATE KINASE | Author keyword | 44 | 35% | 14% | 103 |
| 2 | THYMIDYLATE KINASE | Author keyword | 22 | 66% | 3% | 21 |
| 3 | CHIM STRUCT MACROMOL | Address | 22 | 54% | 4% | 29 |
| 4 | NUCLEOSIDE MONOPHOSPHATE KINASE | Author keyword | 22 | 81% | 2% | 13 |
| 5 | UMP KINASE | Author keyword | 17 | 75% | 2% | 12 |
| 6 | URIDYLATE KINASE | Author keyword | 14 | 100% | 1% | 7 |
| 7 | THYMIDINE MONOPHOSPHATE KINASE | Author keyword | 13 | 80% | 1% | 8 |
| 8 | UMP CMP KINASE | Author keyword | 11 | 78% | 1% | 7 |
| 9 | ENZYMOL PL MICROBIOL | Address | 8 | 75% | 1% | 6 |
| 10 | NUCLEOSIDE MONOPHOSPHATE KINASES | Author keyword | 6 | 80% | 1% | 4 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | ADENYLATE KINASE | 44 | 35% | 14% | 103 | Search ADENYLATE+KINASE | Search ADENYLATE+KINASE |
| 2 | THYMIDYLATE KINASE | 22 | 66% | 3% | 21 | Search THYMIDYLATE+KINASE | Search THYMIDYLATE+KINASE |
| 3 | NUCLEOSIDE MONOPHOSPHATE KINASE | 22 | 81% | 2% | 13 | Search NUCLEOSIDE+MONOPHOSPHATE+KINASE | Search NUCLEOSIDE+MONOPHOSPHATE+KINASE |
| 4 | UMP KINASE | 17 | 75% | 2% | 12 | Search UMP+KINASE | Search UMP+KINASE |
| 5 | URIDYLATE KINASE | 14 | 100% | 1% | 7 | Search URIDYLATE+KINASE | Search URIDYLATE+KINASE |
| 6 | THYMIDINE MONOPHOSPHATE KINASE | 13 | 80% | 1% | 8 | Search THYMIDINE+MONOPHOSPHATE+KINASE | Search THYMIDINE+MONOPHOSPHATE+KINASE |
| 7 | UMP CMP KINASE | 11 | 78% | 1% | 7 | Search UMP+CMP+KINASE | Search UMP+CMP+KINASE |
| 8 | NUCLEOSIDE MONOPHOSPHATE KINASES | 6 | 80% | 1% | 4 | Search NUCLEOSIDE+MONOPHOSPHATE+KINASES | Search NUCLEOSIDE+MONOPHOSPHATE+KINASES |
| 9 | GUANYLATE KINASE | 4 | 33% | 1% | 11 | Search GUANYLATE+KINASE | Search GUANYLATE+KINASE |
| 10 | ADENYLATE KINASE ACTIVITY | 4 | 75% | 0% | 3 | Search ADENYLATE+KINASE+ACTIVITY | Search ADENYLATE+KINASE+ACTIVITY |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | THYMIDINE 5 O MONOPHOSPHATE ANALOGS | 42 | 94% | 2% | 15 |
| 2 | NUCLEOSIDE MONOPHOSPHATE KINASES | 36 | 70% | 4% | 30 |
| 3 | UMP CMP KINASE | 34 | 79% | 3% | 22 |
| 4 | INHIBITOR P1 P5 DIADENOSINE 5 PENTAPHOSPHATE | 30 | 84% | 2% | 16 |
| 5 | ADENOSINE TRIPHOSPHATE TRANSPHOSPHORYLASES | 29 | 88% | 2% | 14 |
| 6 | URIDYLATE KINASE | 26 | 87% | 2% | 13 |
| 7 | GTP AMP PHOSPHOTRANSFERASE | 26 | 80% | 2% | 16 |
| 8 | AZT ACTIVATION | 22 | 68% | 3% | 19 |
| 9 | SUBSTRATE AMP | 17 | 100% | 1% | 8 |
| 10 | CMP KINASE | 14 | 65% | 2% | 13 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Drug Design and Identification of Potent Leads Against Mycobacterium tuberculosis Thymidine Monophosphate Kinase | 2012 | 15 | 45 | 69% |
| The many isoforms of human adenylate kinases | 2014 | 5 | 59 | 42% |
| Adenylate Kinase and AMP Signaling Networks: Metabolic Monitoring, Signal Communication and Body Energy Sensing | 2009 | 77 | 234 | 15% |
| Pyrimidine Salvage Pathway in Mycobacterium tuberculosis | 2011 | 11 | 115 | 28% |
| Human and viral nucleoside/nucleotide kinases involved in antiviral drug activation: Structural and catalytic properties | 2010 | 25 | 179 | 19% |
| Bacterial monophosphate kinases - Biochemical properties and regulatory mechanisms | 2007 | 0 | 26 | 81% |
| Evolution and classification of P-loop kinases and related proteins | 2003 | 125 | 169 | 12% |
| Structural requirements for efficient phosphorylation of nucleotide analogs by human thymidylate kinase | 2004 | 22 | 36 | 28% |
| Human adenylate kinases - classification, structure, physiological and pathological importance | 2015 | 0 | 59 | 46% |
| Adenylate kinases | 1997 | 0 | 91 | 92% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | CHIM STRUCT MACROMOL | 22 | 54% | 3.8% | 29 |
| 2 | ENZYMOL PL MICROBIOL | 8 | 75% | 0.8% | 6 |
| 3 | URA2128 | 2 | 44% | 0.5% | 4 |
| 4 | MOL PHARMACOL DRUG DISCOVERY SECT | 2 | 67% | 0.3% | 2 |
| 5 | FRE 2852 CNRS | 1 | 100% | 0.3% | 2 |
| 6 | FFW | 1 | 21% | 0.7% | 5 |
| 7 | ENZYMOL MOL FONCT | 1 | 27% | 0.4% | 3 |
| 8 | UNITE CHIM BIOCATALYSE | 1 | 15% | 0.7% | 5 |
| 9 | BIOCIENCIAS RIO CLARO BIOL | 1 | 50% | 0.1% | 1 |
| 10 | CNRS URA 09 | 1 | 50% | 0.1% | 1 |
Related classes at same level (level 2) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000026165 | NM23//NUCLEOSIDE DIPHOSPHATE KINASE//ING4 |
| 2 | 0.0000016703 | TRIOSEPHOSPHATE ISOMERASE//PYRUVATE KINASE DEFICIENCY//PYRUVATE KINASE |
| 3 | 0.0000012801 | DEOXYCYTIDINE KINASE//PURINE NUCLEOSIDE PHOSPHORYLASE//NUCLEOSIDE TRANSPORT |
| 4 | 0.0000012202 | LESCH NYHAN SYNDROME//TIAZOFURIN//LESCH NYHAN DISEASE |
| 5 | 0.0000011969 | RIBONUCLEOTIDE REDUCTASE//TRIMIDOX//DIDOX |
| 6 | 0.0000011867 | F 1 ATPASE//ATP SYNTHASE//V ATPASE |
| 7 | 0.0000011123 | CREATINE//MAGNETIC RESONANCE SPECTROSCOPY//ARGININE KINASE |
| 8 | 0.0000009309 | PEPTIDE DEFORMYLASE//ENOYL ACP REDUCTASE//PLATENSIMYCIN |
| 9 | 0.0000009261 | ACTA CRYSTALLOGRAPHICA SECTION D-BIOLOGICAL CRYSTALLOGRAPHY//ACTA CRYSTALLOGRAPHICA SECTION A//JOURNAL OF APPLIED CRYSTALLOGRAPHY |
| 10 | 0.0000008066 | PROTEINS-STRUCTURE FUNCTION AND BIOINFORMATICS//PROTEIN STRUCTURE PREDICTION//PROTEIN FOLDING |