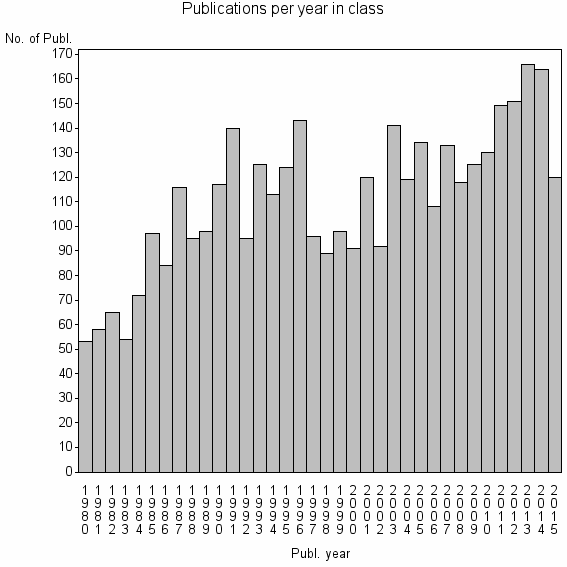
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 2342 | 3993 | 35.6 | 65% |
Classes in level above (level 3) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 295 | 37066 | CORYNEBACTERIUM GLUTAMICUM//ACTA CRYSTALLOGRAPHICA SECTION D-BIOLOGICAL CRYSTALLOGRAPHY//DIHYDRODIPICOLINATE SYNTHASE |
Classes in level below (level 1) |
| ID, lev. below | Publications | Label for level below |
|---|---|---|
| 4524 | 1697 | CORYNEBACTERIUM GLUTAMICUM//GENOMFOR//BIOTECHNOL 1 |
| 9486 | 1083 | ANTHRANILATE SYNTHASE//CHORISMATE MUTASE//PREPHENATE DEHYDROGENASE |
| 17198 | 548 | SHIKIMATE KINASE//SHIKIMATE PATHWAY//CHORISMATE SYNTHASE |
| 21363 | 363 | TYRR//SHIKIMIC ACID PRODUCTION//CIS CIS MUCONIC ACID |
| 23104 | 302 | KDO8P SYNTHASE//3 DEOXY D ARABINO HEPTULOSONATE 7 PHOSPHATE SYNTHASE//KDO8PS |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | CORYNEBACTERIUM GLUTAMICUM | Author keyword | 526 | 68% | 12% | 462 |
| 2 | SHIKIMATE PATHWAY | Author keyword | 90 | 59% | 3% | 102 |
| 3 | SHIKIMATE KINASE | Author keyword | 52 | 87% | 1% | 26 |
| 4 | ANTHRANILATE SYNTHASE | Author keyword | 42 | 70% | 1% | 35 |
| 5 | CHORISMATE MUTASE | Author keyword | 42 | 60% | 1% | 46 |
| 6 | GENOMFOR | Address | 40 | 70% | 1% | 33 |
| 7 | 3 DEOXY D ARABINO HEPTULOSONATE 7 PHOSPHATE SYNTHASE | Author keyword | 38 | 89% | 0% | 17 |
| 8 | BIOTECHNOL 1 | Address | 34 | 34% | 2% | 82 |
| 9 | AMINO ACID PRODUCTION | Author keyword | 30 | 62% | 1% | 31 |
| 10 | CHAIR GENET PROKARYOTES | Address | 29 | 72% | 1% | 23 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | CORYNEBACTERIUM GLUTAMICUM | 526 | 68% | 12% | 462 | Search CORYNEBACTERIUM+GLUTAMICUM | Search CORYNEBACTERIUM+GLUTAMICUM |
| 2 | SHIKIMATE PATHWAY | 90 | 59% | 3% | 102 | Search SHIKIMATE+PATHWAY | Search SHIKIMATE+PATHWAY |
| 3 | SHIKIMATE KINASE | 52 | 87% | 1% | 26 | Search SHIKIMATE+KINASE | Search SHIKIMATE+KINASE |
| 4 | ANTHRANILATE SYNTHASE | 42 | 70% | 1% | 35 | Search ANTHRANILATE+SYNTHASE | Search ANTHRANILATE+SYNTHASE |
| 5 | CHORISMATE MUTASE | 42 | 60% | 1% | 46 | Search CHORISMATE+MUTASE | Search CHORISMATE+MUTASE |
| 6 | 3 DEOXY D ARABINO HEPTULOSONATE 7 PHOSPHATE SYNTHASE | 38 | 89% | 0% | 17 | Search 3+DEOXY+D+ARABINO+HEPTULOSONATE+7+PHOSPHATE+SYNTHASE | Search 3+DEOXY+D+ARABINO+HEPTULOSONATE+7+PHOSPHATE+SYNTHASE |
| 7 | AMINO ACID PRODUCTION | 30 | 62% | 1% | 31 | Search AMINO+ACID+PRODUCTION | Search AMINO+ACID+PRODUCTION |
| 8 | L LYSINE PRODUCTION | 28 | 81% | 0% | 17 | Search L+LYSINE+PRODUCTION | Search L+LYSINE+PRODUCTION |
| 9 | CHORISMATE | 27 | 72% | 1% | 21 | Search CHORISMATE | Search CHORISMATE |
| 10 | CHORISMATE SYNTHASE | 25 | 77% | 0% | 17 | Search CHORISMATE+SYNTHASE | Search CHORISMATE+SYNTHASE |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | BREVIBACTERIUM LACTOFERMENTUM | 190 | 76% | 3% | 134 |
| 2 | BREVIBACTERIUM FLAVUM | 158 | 79% | 3% | 100 |
| 3 | 3 DEOXY D ARABINO HEPTULOSONATE 7 PHOSPHATE SYNTHASE | 140 | 71% | 3% | 113 |
| 4 | OXYGEN DEPRIVATION CONDITIONS | 140 | 84% | 2% | 77 |
| 5 | 3 DEOXY D MANNO OCTULOSONATE 8 PHOSPHATE SYNTHASE | 94 | 89% | 1% | 42 |
| 6 | LYSINE PRODUCTION | 79 | 73% | 2% | 61 |
| 7 | 3 DEHYDROQUINASE | 75 | 91% | 1% | 31 |
| 8 | PREPHENATE | 71 | 74% | 1% | 52 |
| 9 | ATCC 13032 | 64 | 100% | 1% | 21 |
| 10 | SHIKIMATE PATHWAY | 64 | 36% | 4% | 145 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Bio-based production of chemicals, materials and fuels - Corynebacterium glutamicum as versatile cell factory | 2012 | 52 | 73 | 88% |
| The Shikimate Pathway and Aromatic Amino Acid Biosynthesis in Plants | 2012 | 78 | 191 | 60% |
| The complete Corynebacterium glutamicum ATCC 13032 genome sequence and its impact on the production of L-aspartate-derived amino acids and vitamins | 2003 | 432 | 99 | 76% |
| The shikimate pathway | 1999 | 384 | 211 | 58% |
| Bio-based production of organic acids with Corynebacterium glutamicum | 2013 | 22 | 88 | 64% |
| Systems and synthetic metabolic engineering for amino acid production - the heartbeat of industrial strain development | 2012 | 34 | 71 | 77% |
| Towards bacterial strains overproducing L-tryptophan and other aromatics by metabolic engineering | 2006 | 97 | 90 | 90% |
| The Corynebacterium glutamicum genome: features and impacts on biotechnological processes | 2003 | 272 | 78 | 71% |
| Advanced Biotechnology: Metabolically Engineered Cells for the Bio-Based Production of Chemicals and Fuels, Materials, and Health-Care Products | 2015 | 3 | 377 | 34% |
| Sigma factors and promoters in Corynebacterium glutamicum | 2011 | 34 | 103 | 80% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | GENOMFOR | 40 | 70% | 0.8% | 33 |
| 2 | BIOTECHNOL 1 | 34 | 34% | 2.1% | 82 |
| 3 | CHAIR GENET PROKARYOTES | 29 | 72% | 0.6% | 23 |
| 4 | IBG BIOTECHNOL 1 | 17 | 36% | 1.0% | 38 |
| 5 | BIOL CEBITEC | 17 | 100% | 0.2% | 8 |
| 6 | BIO GEOWISSEN | 11 | 38% | 0.6% | 23 |
| 7 | FOR UNGSZENTRUM BIOTECHNOL 1 | 11 | 37% | 0.6% | 24 |
| 8 | FINE CHEM BIOTECHNOL | 10 | 63% | 0.3% | 10 |
| 9 | GENOMFOR SYSTEMBIOL | 7 | 56% | 0.2% | 9 |
| 10 | ZENTRUM GENOMFOR | 6 | 71% | 0.1% | 5 |
Related classes at same level (level 2) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000029797 | DIHYDRODIPICOLINATE SYNTHASE//ASPARTATE KINASE//METHIONINE DEPENDENCE |
| 2 | 0.0000013405 | ALANINE RACEMASE//TYROSINE PHENOL LYASE//MURD |
| 3 | 0.0000013012 | PHOSPHOTRANSFERASE SYSTEM//LACTOSE PERMEASE//BIOL BAKTERIENPHYSIOL |
| 4 | 0.0000010996 | NIFA PROTEIN//GLNK//SIGMA54 |
| 5 | 0.0000010654 | BMC SYSTEMS BIOLOGY//FLUX BALANCE ANALYSIS//SYNTHETIC BIOLOGY |
| 6 | 0.0000010511 | HYDANTOINASE//D HYDANTOINASE//DIRECTED EVOLUTION |
| 7 | 0.0000009197 | ACETIC ACID BACTERIA//PYRROLOQUINOLINE QUINONE//GLUCONOBACTER |
| 8 | 0.0000008305 | O ANTIGEN//BACTERIAL POLYSACCHARIDE STRUCTURE//O SPECIFIC POLYSACCHARIDE |
| 9 | 0.0000008257 | BIOTECHNOLOGY AND BIOENGINEERING//MAMMALIAN CELL CULTURE//HYBRIDOMA |
| 10 | 0.0000008162 | GALACTOSEMIA//GALACTOSE 1 PHOSPHATE URIDYLTRANSFERASE//GALACTOSE 1 PHOSPHATE |