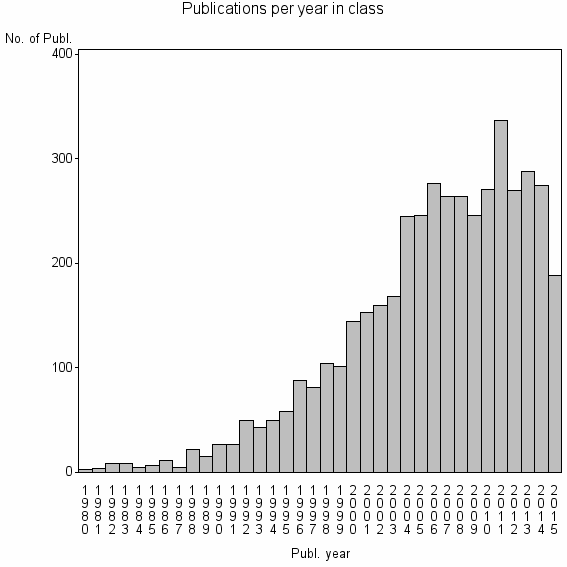
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 2168 | 4502 | 43.2 | 86% |
Classes in level above (level 3) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 460 | 22518 | BACTERIORHODOPSIN//PHOTOCYCLE//PHYTOCHROME |
Classes in level below (level 1) |
| ID, lev. below | Publications | Label for level below |
|---|---|---|
| 3492 | 1893 | TRANSMEMBRANE HELIX//MEMBRANE PROTEIN FOLDING//TRANSMEMBRANE HELICES |
| 10565 | 987 | CELL FREE PROTEIN SYNTHESIS//RIBOSOME DISPLAY//S30 EXTRACT |
| 12019 | 873 | TEV PROTEASE//SOLUBILITY ENHANCER//FUSION TAG |
| 13805 | 749 | AMPHIPOLS//UMR 7099//AMPHIPOL |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | CELL FREE PROTEIN SYNTHESIS | Author keyword | 172 | 64% | 4% | 170 |
| 2 | MEMBRANE PROTEIN | Author keyword | 43 | 12% | 7% | 324 |
| 3 | RIBOSOME DISPLAY | Author keyword | 36 | 48% | 1% | 54 |
| 4 | TRANSMEMBRANE HELIX | Author keyword | 35 | 44% | 1% | 61 |
| 5 | MEMBRANE PROTEIN FOLDING | Author keyword | 34 | 54% | 1% | 44 |
| 6 | AMPHIPOLS | Author keyword | 32 | 80% | 0% | 20 |
| 7 | S30 EXTRACT | Author keyword | 30 | 100% | 0% | 12 |
| 8 | MRNA DISPLAY | Author keyword | 28 | 61% | 1% | 30 |
| 9 | UMR 7099 | Address | 27 | 46% | 1% | 44 |
| 10 | CELL FREE EXPRESSION | Author keyword | 25 | 53% | 1% | 33 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | CELL FREE PROTEIN SYNTHESIS | 172 | 64% | 4% | 170 | Search CELL+FREE+PROTEIN+SYNTHESIS | Search CELL+FREE+PROTEIN+SYNTHESIS |
| 2 | MEMBRANE PROTEIN | 43 | 12% | 7% | 324 | Search MEMBRANE+PROTEIN | Search MEMBRANE+PROTEIN |
| 3 | RIBOSOME DISPLAY | 36 | 48% | 1% | 54 | Search RIBOSOME+DISPLAY | Search RIBOSOME+DISPLAY |
| 4 | TRANSMEMBRANE HELIX | 35 | 44% | 1% | 61 | Search TRANSMEMBRANE+HELIX | Search TRANSMEMBRANE+HELIX |
| 5 | MEMBRANE PROTEIN FOLDING | 34 | 54% | 1% | 44 | Search MEMBRANE+PROTEIN+FOLDING | Search MEMBRANE+PROTEIN+FOLDING |
| 6 | AMPHIPOLS | 32 | 80% | 0% | 20 | Search AMPHIPOLS | Search AMPHIPOLS |
| 7 | S30 EXTRACT | 30 | 100% | 0% | 12 | Search S30+EXTRACT | Search S30+EXTRACT |
| 8 | MRNA DISPLAY | 28 | 61% | 1% | 30 | Search MRNA+DISPLAY | Search MRNA+DISPLAY |
| 9 | CELL FREE EXPRESSION | 25 | 53% | 1% | 33 | Search CELL+FREE+EXPRESSION | Search CELL+FREE+EXPRESSION |
| 10 | MEMBRANE PROTEINS | 23 | 11% | 5% | 205 | Search MEMBRANE+PROTEINS | Search MEMBRANE+PROTEINS |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | TRANSMEMBRANE ALPHA HELICES | 73 | 44% | 3% | 124 |
| 2 | HYDROPHOBIC MISMATCH | 54 | 36% | 3% | 121 |
| 3 | AMPHIPOLS | 53 | 85% | 1% | 28 |
| 4 | KNOWLEDGE BASED SCALE | 52 | 100% | 0% | 18 |
| 5 | SYNTHESIS SYSTEM | 49 | 45% | 2% | 81 |
| 6 | FREE PROTEIN SYNTHESIS | 47 | 43% | 2% | 83 |
| 7 | DRIVE ASSOCIATION | 46 | 78% | 1% | 31 |
| 8 | FREE TRANSLATION SYSTEM | 45 | 56% | 1% | 55 |
| 9 | FREE TRANSLATION | 44 | 71% | 1% | 36 |
| 10 | GXXXG MOTIF | 44 | 46% | 2% | 71 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Mechanisms of Integral Membrane Protein Insertion and Folding | 2015 | 5 | 149 | 42% |
| Cell-free protein synthesis: Applications come of age | 2012 | 64 | 86 | 73% |
| Amphipols From A to Z | 2011 | 84 | 85 | 64% |
| Amphipols for Each Season | 2014 | 12 | 132 | 70% |
| Transmembrane helix dimerization: Beyond the search for sequence motifs | 2012 | 40 | 122 | 84% |
| Membrane protein assembly into Nanodiscs | 2010 | 143 | 55 | 47% |
| Membrane protein folding and stability: Physical principles | 1999 | 1077 | 162 | 43% |
| Enhancement of soluble protein expression through the use of fusion tags | 2006 | 179 | 48 | 75% |
| Solving the membrane protein folding problem | 2005 | 264 | 83 | 77% |
| Amphipols, Nanodiscs, and Fluorinated Surfactants: Three Nonconventional Approaches to Studying Membrane Proteins in Aqueous Solutions | 2010 | 83 | 134 | 56% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | UMR 7099 | 27 | 46% | 1.0% | 44 |
| 2 | FRC 550 | 19 | 80% | 0.3% | 12 |
| 3 | PHYSICOCHIM MOL MEMBRANES BIOL | 13 | 61% | 0.3% | 14 |
| 4 | ICTAS | 9 | 29% | 0.6% | 25 |
| 5 | SCI BIO OURCES PROGRAM | 8 | 100% | 0.1% | 5 |
| 6 | UMR 7099FRC 550 | 8 | 100% | 0.1% | 5 |
| 7 | MEMBRANE PROT CRYSTALLOG GRP | 8 | 38% | 0.4% | 16 |
| 8 | STOCKHOLM BIOINFORMAT | 7 | 18% | 0.8% | 36 |
| 9 | BIOL PHYSICOCHIM PROT MEMBRANAI | 7 | 64% | 0.2% | 7 |
| 10 | PROT ENGN SECT | 7 | 46% | 0.2% | 11 |
Related classes at same level (level 2) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000019976 | RGS PROTEINS//G PROTEIN COUPLED RECEPTOR//RGS |
| 2 | 0.0000017129 | PROTEIN TRANSLOCATION//PROTEIN IMPORT//SECA |
| 3 | 0.0000015764 | PHOSPHOTRANSFERASE SYSTEM//LACTOSE PERMEASE//BIOL BAKTERIENPHYSIOL |
| 4 | 0.0000015560 | AUTOTRANSPORTER//PORIN//BACTERIAL CHEMOTAXIS |
| 5 | 0.0000013077 | PROTEINS-STRUCTURE FUNCTION AND BIOINFORMATICS//PROTEIN STRUCTURE PREDICTION//PROTEIN FOLDING |
| 6 | 0.0000013031 | PHAGE DISPLAY//CATALYTIC ANTIBODIES//CATALYTIC ANTIBODY |
| 7 | 0.0000011754 | ANTIMICROBIAL PEPTIDE//ANTIMICROBIAL PEPTIDES//DEFENSINS |
| 8 | 0.0000011186 | MONOOLEIN//CUBOSOMES//PHYTANTRIOL |
| 9 | 0.0000010591 | MOLTEN GLOBULE//APOMYOGLOBIN//PROTEIN FORMULATION |
| 10 | 0.0000010085 | CHEMISTRY AND PHYSICS OF LIPIDS//LIPOSOME//BIOCHIMICA ET BIOPHYSICA ACTA-BIOMEMBRANES |