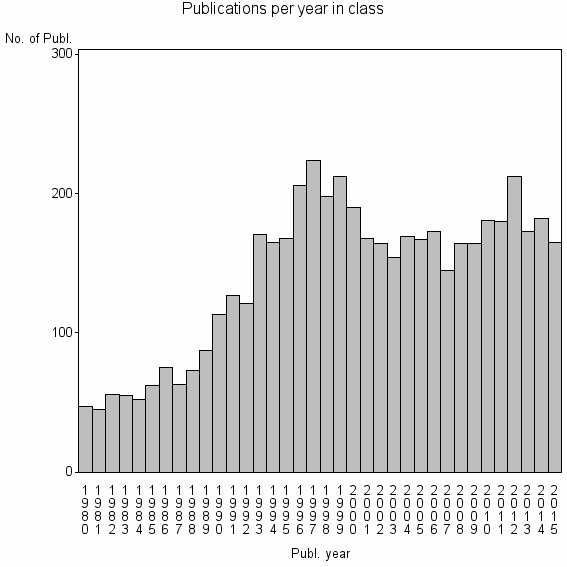
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 1988 | 5071 | 45.5 | 72% |
Classes in level above (level 3) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 66 | 85654 | PHOTOSYSTEM II//PHOTOSYNTHESIS RESEARCH//PHOTOSYSTEM I |
Classes in level below (level 1) |
| ID, lev. below | Publications | Label for level below |
|---|---|---|
| 5350 | 1566 | ELECTRON TRANSFER MATRIX ELEMENT//ELECTRON TRANSFER REACTIVITY//COUPLING MATRIX ELEMENT |
| 10001 | 1036 | COILED COIL//COILED COILS//LEUCINE ZIPPERS |
| 10975 | 951 | PROTEIN DESIGN//FOUR HELIX BUNDLE//TEMPLATE ASSEMBLED SYNTHETIC PROTEIN |
| 15226 | 655 | PROTON COUPLED ELECTRON TRANSFER//CYCLIC VOLTABSORPTOMETRY CVA//UNITE MIXTE RECH UNIV 7591 |
| 15509 | 638 | ARTIFICIAL METALLOENZYMES//ARTIFICIAL METALLOENZYME//HIGH INTENS XRAY DIFFRACT |
| 25763 | 225 | PEPTIDE SELF ASSEMBLED MONOLAYERS//HELICAL PEPTIDE//HELIX PEPTIDE |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | ARTIFICIAL METALLOENZYMES | Author keyword | 47 | 86% | 0% | 24 |
| 2 | COILED COIL | Author keyword | 41 | 24% | 3% | 147 |
| 3 | PROTEIN DESIGN | Author keyword | 34 | 19% | 3% | 157 |
| 4 | PROTEIN DE NOVO DESIGN | Author keyword | 27 | 92% | 0% | 11 |
| 5 | ELECTRON TRANSFER MATRIX ELEMENT | Author keyword | 23 | 100% | 0% | 10 |
| 6 | ELECTRON TRANSFER REACTIVITY | Author keyword | 21 | 85% | 0% | 11 |
| 7 | DE NOVO DESIGN | Author keyword | 20 | 28% | 1% | 61 |
| 8 | PROTON COUPLED ELECTRON TRANSFER | Author keyword | 18 | 36% | 1% | 39 |
| 9 | COILED COILS | Author keyword | 14 | 33% | 1% | 36 |
| 10 | FOUR HELIX BUNDLE | Author keyword | 14 | 35% | 1% | 32 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | ARTIFICIAL METALLOENZYMES | 47 | 86% | 0% | 24 | Search ARTIFICIAL+METALLOENZYMES | Search ARTIFICIAL+METALLOENZYMES |
| 2 | COILED COIL | 41 | 24% | 3% | 147 | Search COILED+COIL | Search COILED+COIL |
| 3 | PROTEIN DESIGN | 34 | 19% | 3% | 157 | Search PROTEIN+DESIGN | Search PROTEIN+DESIGN |
| 4 | PROTEIN DE NOVO DESIGN | 27 | 92% | 0% | 11 | Search PROTEIN+DE+NOVO+DESIGN | Search PROTEIN+DE+NOVO+DESIGN |
| 5 | ELECTRON TRANSFER MATRIX ELEMENT | 23 | 100% | 0% | 10 | Search ELECTRON+TRANSFER+MATRIX+ELEMENT | Search ELECTRON+TRANSFER+MATRIX+ELEMENT |
| 6 | ELECTRON TRANSFER REACTIVITY | 21 | 85% | 0% | 11 | Search ELECTRON+TRANSFER+REACTIVITY | Search ELECTRON+TRANSFER+REACTIVITY |
| 7 | DE NOVO DESIGN | 20 | 28% | 1% | 61 | Search DE+NOVO+DESIGN | Search DE+NOVO+DESIGN |
| 8 | PROTON COUPLED ELECTRON TRANSFER | 18 | 36% | 1% | 39 | Search PROTON+COUPLED+ELECTRON+TRANSFER | Search PROTON+COUPLED+ELECTRON+TRANSFER |
| 9 | COILED COILS | 14 | 33% | 1% | 36 | Search COILED+COILS | Search COILED+COILS |
| 10 | FOUR HELIX BUNDLE | 14 | 35% | 1% | 32 | Search FOUR+HELIX+BUNDLE | Search FOUR+HELIX+BUNDLE |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | GCN4 LEUCINE ZIPPER | 172 | 54% | 4% | 219 |
| 2 | BIOTIN AVIDIN TECHNOLOGY | 89 | 81% | 1% | 54 |
| 3 | TUNNELING PATHWAYS | 76 | 54% | 2% | 98 |
| 4 | DISTANCE DEPENDENCE | 75 | 27% | 5% | 240 |
| 5 | ARTIFICIAL METALLOENZYMES | 73 | 53% | 2% | 96 |
| 6 | DENOVO DESIGN | 69 | 66% | 1% | 64 |
| 7 | 4 HELIX BUNDLE | 65 | 46% | 2% | 105 |
| 8 | TRANSFER MATRIX ELEMENTS | 61 | 67% | 1% | 55 |
| 9 | 4 HELIX BUNDLE PROTEIN | 58 | 51% | 2% | 82 |
| 10 | DE NOVO DESIGN | 53 | 19% | 5% | 247 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Design of functional metalloproteins | 2009 | 232 | 98 | 61% |
| Electron transfer in proteins | 1996 | 733 | 95 | 68% |
| Proton-coupled electron transfer with photoexcited ruthenium(II), rhenium(I), and iridium(III) complexes | 2015 | 4 | 67 | 76% |
| Artificial metalloenzymes for enantioselective catalysis | 2014 | 23 | 47 | 83% |
| Charge Transfer in Dynamical Biosystems, or The Treachery of (Static) Images | 2015 | 6 | 56 | 52% |
| Understanding Hydrogen Atom Transfer: From Bond Strengths to Marcus Theory | 2011 | 125 | 35 | 51% |
| Coiled coils: New structures and new functions | 1996 | 999 | 48 | 46% |
| Theory of Proton-Coupled Electron Transfer in Energy Conversion Processes | 2009 | 128 | 37 | 73% |
| The design of coiled-coil structures and assemblies | 2005 | 262 | 121 | 77% |
| The structure of alpha-helical coiled coils | 2005 | 330 | 90 | 57% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | BIOL BIOCHEM 2 | 10 | 50% | 0.3% | 15 |
| 2 | JOHNSON FDN | 9 | 13% | 1.2% | 62 |
| 3 | HIGH INTENS XRAY DIFFRACT | 8 | 60% | 0.2% | 9 |
| 4 | UNITE MIXTE RECH UNIV 7591 | 4 | 50% | 0.1% | 6 |
| 5 | BCH DORIGNY | 4 | 31% | 0.2% | 11 |
| 6 | BIOMOL DESIGN DRUG DEV | 4 | 31% | 0.2% | 10 |
| 7 | UM SYLVESTER BRAMAN FAMILY BREAST CANC | 3 | 57% | 0.1% | 4 |
| 8 | FUNAI | 2 | 44% | 0.1% | 4 |
| 9 | KANSAI PHOTON | 2 | 67% | 0.0% | 2 |
| 10 | UNITE MIXTE RECH UNIV | 2 | 40% | 0.1% | 4 |
Related classes at same level (level 2) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000017248 | CERULOPLASMIN//PLASTOCYANIN//BLUE COPPER PROTEIN |
| 2 | 0.0000015230 | ESIPT//DUAL FLUORESCENCE//TICT |
| 3 | 0.0000014793 | CHLOROPEROXIDASE//CYTOCHROME C3//PEROXIDASE |
| 4 | 0.0000013101 | INORGAN CHEM SECT//TERPYRIDINE//ARYLAZOIMIDAZOLE |
| 5 | 0.0000011130 | FOLDAMERS//PEPTAIBOL//BETA PEPTIDES |
| 6 | 0.0000010818 | REACTION CENTER//PHOTOSYNTHETIC REACTION CENTER//MULTIDIMENS SPECT |
| 7 | 0.0000010624 | PROTEINS-STRUCTURE FUNCTION AND BIOINFORMATICS//PROTEIN STRUCTURE PREDICTION//PROTEIN FOLDING |
| 8 | 0.0000008878 | HELSINKI BIOENERGET GRP//CYTOCHROME BC1 COMPLEX//CYTOCHROME C OXIDASE |
| 9 | 0.0000008850 | MG CHLOROPHYLL A//NADH MODELS//NADH MODEL |
| 10 | 0.0000008659 | METHEMOGLOBINEMIA//CYTOCHROME B5//DAPSONE |