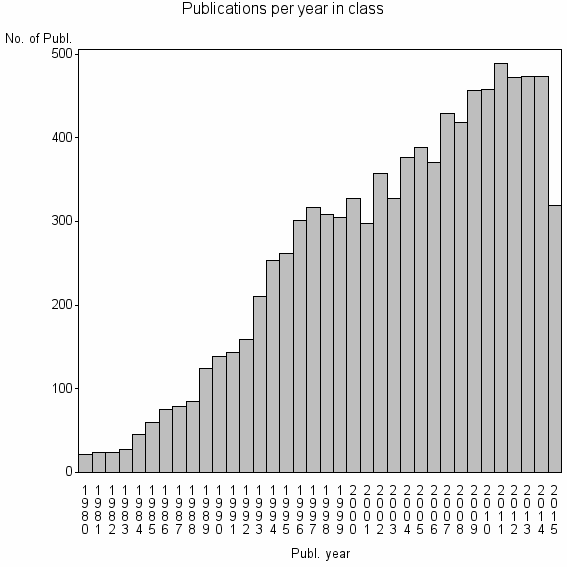
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 1084 | 9394 | 43.3 | 83% |
Classes in level above (level 3) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 136 | 62081 | RNA-A PUBLICATION OF THE RNA SOCIETY//RIBOZYME//NUCLEIC ACIDS RESEARCH |
Classes in level below (level 1) |
| ID, lev. below | Publications | Label for level below |
|---|---|---|
| 3040 | 2004 | RNA SECONDARY STRUCTURE//RNA SECONDARY STRUCTURE PREDICTION//RNA STRUCTURE PREDICTION |
| 3111 | 1980 | HAMMERHEAD RIBOZYME//HAIRPIN RIBOZYME//RIBOZYME |
| 3858 | 1821 | GROUP I INTRON//SELF SPLICING//RIBOZYME |
| 8044 | 1224 | HFQ//S RNA//RYHB |
| 12378 | 846 | PROTEIN SPLICING//INTEIN//HOMING ENDONUCLEASE |
| 12424 | 843 | RIBOSWITCH//RIBOSWITCHES//STREPTOMYCES DAVAWENSIS |
| 20449 | 398 | Q BETA REPLICASE//POLONIES//RNA PHAGE |
| 23870 | 278 | BASE PAIR KINETICS//GNRA TETRALOOPS//RAAA HAIRPIN MOTIF |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | RIBOZYME | Author keyword | 390 | 53% | 6% | 523 |
| 2 | GROUP I INTRON | Author keyword | 215 | 70% | 2% | 180 |
| 3 | RIBOSWITCH | Author keyword | 200 | 72% | 2% | 155 |
| 4 | HFQ | Author keyword | 191 | 79% | 1% | 123 |
| 5 | RNA FOLDING | Author keyword | 185 | 57% | 2% | 216 |
| 6 | SELF SPLICING | Author keyword | 161 | 92% | 1% | 65 |
| 7 | CATALYTIC RNA | Author keyword | 159 | 67% | 2% | 142 |
| 8 | PROTEIN SPLICING | Author keyword | 148 | 79% | 1% | 96 |
| 9 | HAMMERHEAD RIBOZYME | Author keyword | 127 | 68% | 1% | 112 |
| 10 | RNA CATALYSIS | Author keyword | 113 | 71% | 1% | 92 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | RIBOZYME | 390 | 53% | 6% | 523 | Search RIBOZYME | Search RIBOZYME |
| 2 | GROUP I INTRON | 215 | 70% | 2% | 180 | Search GROUP+I+INTRON | Search GROUP+I+INTRON |
| 3 | RIBOSWITCH | 200 | 72% | 2% | 155 | Search RIBOSWITCH | Search RIBOSWITCH |
| 4 | HFQ | 191 | 79% | 1% | 123 | Search HFQ | Search HFQ |
| 5 | RNA FOLDING | 185 | 57% | 2% | 216 | Search RNA+FOLDING | Search RNA+FOLDING |
| 6 | SELF SPLICING | 161 | 92% | 1% | 65 | Search SELF+SPLICING | Search SELF+SPLICING |
| 7 | CATALYTIC RNA | 159 | 67% | 2% | 142 | Search CATALYTIC+RNA | Search CATALYTIC+RNA |
| 8 | PROTEIN SPLICING | 148 | 79% | 1% | 96 | Search PROTEIN+SPLICING | Search PROTEIN+SPLICING |
| 9 | HAMMERHEAD RIBOZYME | 127 | 68% | 1% | 112 | Search HAMMERHEAD+RIBOZYME | Search HAMMERHEAD+RIBOZYME |
| 10 | RNA CATALYSIS | 113 | 71% | 1% | 92 | Search RNA+CATALYSIS | Search RNA+CATALYSIS |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | TETRAHYMENA RIBOZYME | 563 | 74% | 4% | 421 |
| 2 | SELF CLEAVAGE | 467 | 69% | 4% | 396 |
| 3 | HAIRPIN RIBOZYME | 376 | 69% | 3% | 325 |
| 4 | SM LIKE PROTEIN | 326 | 85% | 2% | 173 |
| 5 | HAMMERHEAD RIBOZYME | 288 | 47% | 5% | 457 |
| 6 | SELF CLEAVAGE REACTION | 256 | 100% | 1% | 62 |
| 7 | ESCHERICHIA COLI HFQ | 222 | 91% | 1% | 92 |
| 8 | GROUP I INTRON | 194 | 51% | 3% | 271 |
| 9 | SMALL NONCODING RNAS | 186 | 62% | 2% | 192 |
| 10 | GROUP I RIBOZYME | 186 | 71% | 2% | 151 |
Journals |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | RNA-A PUBLICATION OF THE RNA SOCIETY | 45 | 12% | 4% | 348 |
| 2 | RNA BIOLOGY | 15 | 13% | 1% | 104 |
| 3 | RNA | 14 | 14% | 1% | 97 |
| 4 | ANTISENSE RESEARCH AND DEVELOPMENT | 1 | 10% | 0% | 7 |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Expanded sequence dependence of thermodynamic parameters improves prediction of RNA secondary structure | 1999 | 2499 | 93 | 74% |
| A Decade of Riboswitches | 2013 | 133 | 73 | 88% |
| Regulatory RNAs in Bacteria | 2009 | 566 | 116 | 71% |
| Regulation by Small RNAs in Bacteria: Expanding Frontiers | 2011 | 250 | 106 | 84% |
| Hfq and its constellation of RNA | 2011 | 239 | 146 | 79% |
| Regulatory small RNAs from the 3 ' regions of bacterial mRNAs | 2015 | 4 | 57 | 81% |
| Riboswitch engineering - making the all-important second and third steps | 2015 | 4 | 49 | 80% |
| Prospects for Riboswitch Discovery and Analysis | 2011 | 125 | 100 | 86% |
| Bacterial Small RNA-based Negative Regulation: Hfq and Its Accomplices | 2013 | 52 | 56 | 77% |
| Riboswitches: Emerging themes in RNA structure and function | 2008 | 193 | 85 | 85% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | GRP ECOL GENET | 31 | 74% | 0.2% | 23 |
| 2 | RNA GRP | 19 | 37% | 0.4% | 42 |
| 3 | RNA BIOL GRP | 19 | 47% | 0.3% | 30 |
| 4 | SINGLE MOL ANAL GRP | 15 | 49% | 0.2% | 23 |
| 5 | GRP ECOL GENET RIZOSFERA | 15 | 77% | 0.1% | 10 |
| 6 | GRP BIOINFORMAT COMPUTAT BIOL | 15 | 88% | 0.1% | 7 |
| 7 | NONCODING RNA TECHNOL HLTH | 11 | 31% | 0.3% | 31 |
| 8 | TC JENKINS BIOPHYS | 10 | 22% | 0.4% | 41 |
| 9 | HUMAN GENET MOL PEDIAT DIS | 9 | 45% | 0.2% | 15 |
| 10 | INTERDISCIPLINARY BIOINFORMAT | 8 | 16% | 0.5% | 49 |
Related classes at same level (level 2) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000016766 | RNASE P//AMINOACYL TRNA SYNTHETASE//TRNA |
| 2 | 0.0000012993 | RIBOSOME//TRANS TRANSLATION//TMRNA |
| 3 | 0.0000012843 | Z DNA//JOURNAL OF BIOMOLECULAR STRUCTURE & DYNAMICS//B Z TRANSITION |
| 4 | 0.0000011903 | ORIGINS OF LIFE AND EVOLUTION OF BIOSPHERES//ORIGINS OF LIFE AND EVOLUTION OF THE BIOSPHERE//ORIGIN OF LIFE |
| 5 | 0.0000009579 | APTAMER//G QUADRUPLEX//SELEX |
| 6 | 0.0000007445 | PHOSPHOLIPIDOSIS//2 DEOXYSTREPTAMINE//AMINOGLYCOSIDES |
| 7 | 0.0000007326 | CAS9//GENOME EDITING//CRISPR |
| 8 | 0.0000007147 | RNA POLYMERASE//SITE SPECIFIC RECOMBINATION//H NS |
| 9 | 0.0000006595 | JOURNAL OF MAGNETIC RESONANCE//JOURNAL OF BIOMOLECULAR NMR//RESIDUAL DIPOLAR COUPLINGS |
| 10 | 0.0000006422 | MIMIC HYDROLASE//METALLOMICELLE//P NITROPHENYL PICOLINATE |