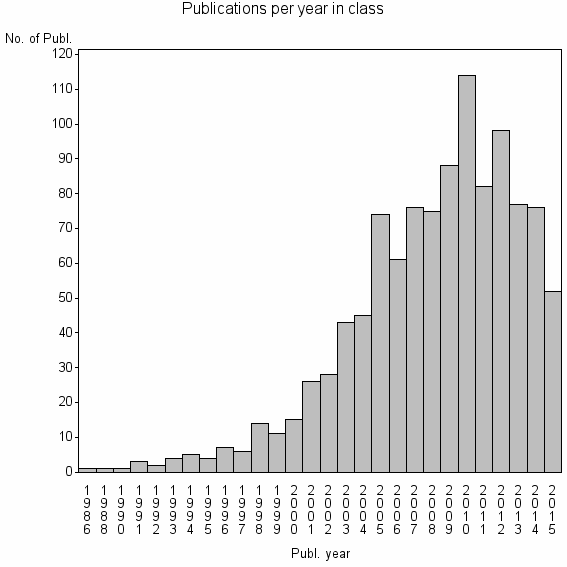
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 9414 | 1089 | 43.4 | 85% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 42 | 31225 | PROTEINS-STRUCTURE FUNCTION AND BIOINFORMATICS//PROTEIN STRUCTURE PREDICTION//PROTEIN FOLDING |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | COMPLEX SCI SOFTWARE | Address | 15 | 82% | 1% | 9 |
| 2 | 3D ZERNIKE DESCRIPTOR | Author keyword | 12 | 86% | 1% | 6 |
| 3 | PROTEIN SURFACE SHAPE | Author keyword | 12 | 86% | 1% | 6 |
| 4 | STRUCTURE BASED FUNCTION PREDICTION | Author keyword | 9 | 83% | 0% | 5 |
| 5 | PROTEIN FUNCTION PREDICTION | Author keyword | 9 | 22% | 3% | 34 |
| 6 | FUNCTIONAL SITE PREDICTION | Author keyword | 6 | 53% | 1% | 8 |
| 7 | THEMATICS | Author keyword | 6 | 47% | 1% | 9 |
| 8 | GLOBAL COMPUTAT CHEM | Address | 5 | 63% | 0% | 5 |
| 9 | LIGAND BINDING SITE PREDICTION | Author keyword | 5 | 63% | 0% | 5 |
| 10 | FUNCTIONAL RESIDUES | Author keyword | 5 | 44% | 1% | 8 |
Web of Science journal categories |
Author Key Words |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | FUNCTIONAL SITES | 38 | 42% | 6% | 70 |
| 2 | 3D TEMPLATES | 35 | 76% | 2% | 25 |
| 3 | FUNCTIONALLY IMPORTANT RESIDUES | 27 | 63% | 2% | 27 |
| 4 | 3D COORDINATE TEMPLATES | 25 | 58% | 3% | 29 |
| 5 | SIDE CHAIN PATTERNS | 24 | 91% | 1% | 10 |
| 6 | FUNCTION ANNOTATION | 15 | 68% | 1% | 13 |
| 7 | LIGAND BINDING SITES | 15 | 22% | 6% | 60 |
| 8 | FUNCTION INFERENCE | 14 | 100% | 1% | 7 |
| 9 | ENZYME ACTIVE SITES | 12 | 37% | 2% | 27 |
| 10 | COORDINATE TEMPLATES | 11 | 100% | 1% | 6 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Structure-based druggability assessment - identifying suitable targets for small molecule therapeutics | 2011 | 39 | 49 | 57% |
| Druggable pockets and binding site centric chemical space: a paradigm shift in drug discovery | 2010 | 88 | 112 | 51% |
| ConSurf: Using Evolutionary Data to Raise Testable Hypotheses about Protein Function | 2013 | 35 | 43 | 23% |
| Predicting protein function from sequence and structure | 2007 | 156 | 96 | 32% |
| Predicting protein function from sequence and structural data | 2005 | 152 | 48 | 50% |
| Searching for functional sites in protein structures | 2004 | 107 | 39 | 59% |
| Protein function annotation by homology-based inference | 2009 | 70 | 72 | 36% |
| Toward mechanistic classification of enzyme functions | 2011 | 15 | 53 | 45% |
| Pocket-Based Drug Design: Exploring Pocket Space | 2013 | 9 | 84 | 42% |
| Exploring the structure and function paradigm | 2008 | 36 | 75 | 51% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | COMPLEX SCI SOFTWARE | 15 | 82% | 0.8% | 9 |
| 2 | GLOBAL COMPUTAT CHEM | 5 | 63% | 0.5% | 5 |
| 3 | MOL BIOINFORMAT | 4 | 21% | 1.4% | 15 |
| 4 | STUDY SYST BIOL | 3 | 19% | 1.4% | 15 |
| 5 | CIBR COMPUTAT INTEGRAT BIOMED | 2 | 67% | 0.2% | 2 |
| 6 | GENOM BIOCHIM METAB | 2 | 67% | 0.2% | 2 |
| 7 | UMR GENOM METAB 8030 | 2 | 67% | 0.2% | 2 |
| 8 | MARKEY STRUCT BIOL | 2 | 11% | 1.3% | 14 |
| 9 | CHANGSHU | 1 | 31% | 0.4% | 4 |
| 10 | COMPUTAT CHEM CHEMINFORMAT | 1 | 50% | 0.2% | 2 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000224003 | CAPRI//PROTEIN PROTEIN DOCKING//PROTEIN DOCKING |
| 2 | 0.0000219066 | PROTEIN STRUCTURE PREDICTION//FOLD RECOGNITION//CASP |
| 3 | 0.0000205996 | MANDELATE RACEMASE//ENOLASE SUPERFAMILY//MANDELATE RACEMASE EC 5122 |
| 4 | 0.0000180181 | ZINC PROTEINS//DRUG DESIGN BIOTECHNOL//METALLOPROTEOME |
| 5 | 0.0000137003 | INTERACTION PROPENSITY//RNA BINDING RESIDUES//COMPUTAT INTELLIGENCE LEARNING DISCOVERY |
| 6 | 0.0000122981 | EVOLUTIONARY BIOINFORMAT//CLOSED LOOPS//SKEW PAIRS |
| 7 | 0.0000120002 | EMBL OUTSTN//PROT INFORMAT OURCE//HLTH SCI TECHNOL ICIR |
| 8 | 0.0000112225 | JOURNAL OF CHEMICAL INFORMATION AND MODELING//VIRTUAL SCREENING//JOURNAL OF COMPUTER-AIDED MOLECULAR DESIGN |
| 9 | 0.0000095753 | POLYPHARMACOLOGY//NETWORK PHARMACOLOGY//DRUG REPOSITIONING |
| 10 | 0.0000093986 | BIOMAG BANK//PDBJ//RCSB PROT DATA BANK |