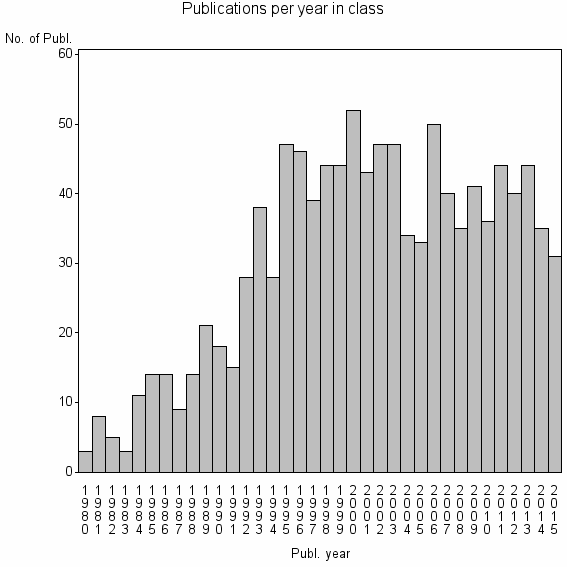
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 9295 | 1101 | 49.7 | 87% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | PKR | Author keyword | 108 | 48% | 15% | 167 |
| 2 | VA RNA | Author keyword | 18 | 89% | 1% | 8 |
| 3 | NF90 | Author keyword | 11 | 67% | 1% | 10 |
| 4 | NC886 | Author keyword | 11 | 100% | 1% | 6 |
| 5 | RNA DEPENDENT PROTEIN KINASE | Author keyword | 8 | 75% | 1% | 6 |
| 6 | PROTEIN KINASE PKR | Author keyword | 7 | 57% | 1% | 8 |
| 7 | EXOGENOUS RNA | Author keyword | 6 | 71% | 0% | 5 |
| 8 | PROTEIN KINASE R | Author keyword | 6 | 35% | 1% | 13 |
| 9 | DSRNA BINDING DOMAIN | Author keyword | 5 | 63% | 0% | 5 |
| 10 | VIRUS CELL INTERACT | Address | 5 | 50% | 1% | 7 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | PKR | 108 | 48% | 15% | 167 | Search PKR | Search PKR |
| 2 | VA RNA | 18 | 89% | 1% | 8 | Search VA+RNA | Search VA+RNA |
| 3 | NF90 | 11 | 67% | 1% | 10 | Search NF90 | Search NF90 |
| 4 | NC886 | 11 | 100% | 1% | 6 | Search NC886 | Search NC886 |
| 5 | RNA DEPENDENT PROTEIN KINASE | 8 | 75% | 1% | 6 | Search RNA+DEPENDENT+PROTEIN+KINASE | Search RNA+DEPENDENT+PROTEIN+KINASE |
| 6 | PROTEIN KINASE PKR | 7 | 57% | 1% | 8 | Search PROTEIN+KINASE+PKR | Search PROTEIN+KINASE+PKR |
| 7 | EXOGENOUS RNA | 6 | 71% | 0% | 5 | Search EXOGENOUS+RNA | Search EXOGENOUS+RNA |
| 8 | PROTEIN KINASE R | 6 | 35% | 1% | 13 | Search PROTEIN+KINASE+R | Search PROTEIN+KINASE+R |
| 9 | DSRNA BINDING DOMAIN | 5 | 63% | 0% | 5 | Search DSRNA+BINDING+DOMAIN | Search DSRNA+BINDING+DOMAIN |
| 10 | DOUBLE STRANDED RNA DEPENDENT PROTEIN KINASE | 5 | 44% | 1% | 8 | Search DOUBLE+STRANDED+RNA+DEPENDENT+PROTEIN+KINASE | Search DOUBLE+STRANDED+RNA+DEPENDENT+PROTEIN+KINASE |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | P68 KINASE | 85 | 86% | 4% | 43 |
| 2 | ACTIVATED P68 KINASE | 53 | 95% | 2% | 18 |
| 3 | CELLULAR ACTIVATOR | 48 | 76% | 3% | 34 |
| 4 | ADENOVIRUS VAI RNA | 46 | 65% | 4% | 44 |
| 5 | INTERFERON ACTION | 44 | 38% | 8% | 91 |
| 6 | PKR | 43 | 32% | 10% | 111 |
| 7 | NUCLEAR FACTOR 90 | 41 | 85% | 2% | 22 |
| 8 | INITIATION FACTOR EIF 2 | 27 | 65% | 2% | 26 |
| 9 | 58 000 DALTON CELLULAR INHIBITOR | 27 | 83% | 1% | 15 |
| 10 | PROTEIN KINASE PKR | 24 | 22% | 9% | 97 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| The dsRNA protein kinase PKR: Virus and cell control | 2007 | 240 | 193 | 61% |
| PKR; a sentinel kinase for cellular stress | 1999 | 535 | 74 | 78% |
| Impact of protein kinase PKR in cell biology: from antiviral to antiproliferative action | 2006 | 296 | 381 | 47% |
| The impact of PKR activation: from neurodegeneration to cancer | 2014 | 12 | 127 | 58% |
| The Role of Protein Kinase R in the Interferon Response | 2011 | 49 | 132 | 67% |
| The double-stranded RNA-dependent protein kinase PKR: Structure and function | 1997 | 406 | 205 | 78% |
| When two strands are better than one: The mediators and modulators of the cellular responses to double-stranded RNA | 1996 | 404 | 147 | 54% |
| Structure and function of the protein kinase R | 2007 | 102 | 233 | 72% |
| Inhibition of PKR by RNA and DNA viruses | 2006 | 106 | 125 | 53% |
| Activation of PKR: an open and shut case? | 2007 | 50 | 38 | 87% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | VIRUS CELL INTERACT | 5 | 50% | 0.6% | 7 |
| 2 | GRP BRAIN AGING EA 3808 | 4 | 75% | 0.3% | 3 |
| 3 | ANALYT ULTRACENTRIFUGAT IL | 4 | 32% | 0.9% | 10 |
| 4 | GRP BRAIN AGING | 4 | 41% | 0.6% | 7 |
| 5 | CLIN MEMORY | 2 | 67% | 0.2% | 2 |
| 6 | INMUNOPATOL SIDA | 2 | 67% | 0.2% | 2 |
| 7 | MANITOBA GRP PROT STRUCT FUNCT | 2 | 67% | 0.2% | 2 |
| 8 | PROGRAM BIOCHEM CELLULAR DEV BIOL | 2 | 67% | 0.2% | 2 |
| 9 | SERV HISTOL BIOL VIEILLISSEMENT | 2 | 67% | 0.2% | 2 |
| 10 | PROGRAM MOL VIROL | 2 | 31% | 0.5% | 5 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000165934 | EIF3//GENE REGULAT DEV//EIF1 |
| 2 | 0.0000131057 | RNASE L//2 5A//SECT BIOMED CHEM |
| 3 | 0.0000125696 | ADAR//RNA EDITING//ADAR2 |
| 4 | 0.0000066952 | IRF 1//IRF 2//INTERFERON REGULATORY FACTOR 1 |
| 5 | 0.0000064494 | O I//VIRAL MICRORNAS//VIRAL MIRNA |
| 6 | 0.0000064385 | MX PROTEIN//MX GENE//MXA PROTEIN |
| 7 | 0.0000059451 | RIG I//IRF3//MDA5 |
| 8 | 0.0000051653 | MRNA DELIVERY//MRNA TRANSFECTION//RIBOL |
| 9 | 0.0000046826 | INTERFERON RECEPTOR//IFNAR2//IFNAR1 |
| 10 | 0.0000043684 | IRES//INTERNAL RIBOSOME ENTRY SITE IRES//INTERNAL RIBOSOME ENTRY SITE |