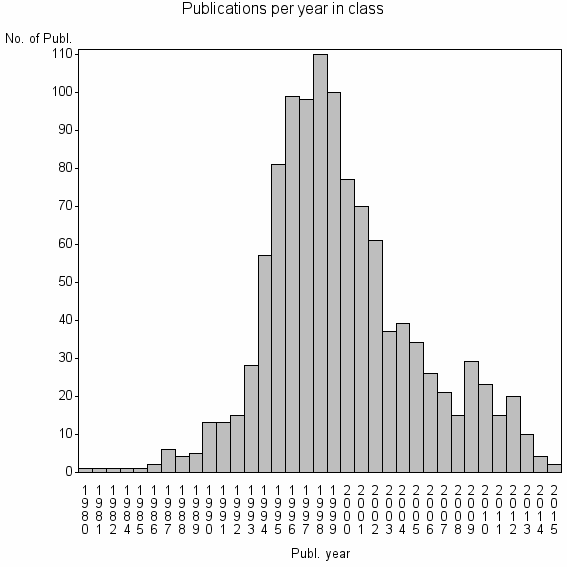
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 9118 | 1119 | 24.5 | 82% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 1339 | 7890 | BIOTECHNIQUES//PCR-METHODS AND APPLICATIONS//DIFFERENTIAL DISPLAY |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | DIFFERENTIAL DISPLAY | Author keyword | 15 | 12% | 11% | 118 |
| 2 | MRNA DIFFERENTIAL DISPLAY | Author keyword | 12 | 25% | 4% | 42 |
| 3 | ANIM NUTR FEED YUNNAN PROV | Address | 11 | 54% | 1% | 14 |
| 4 | MOL GERONTOL UNIT | Address | 4 | 67% | 0% | 4 |
| 5 | SUBTRACTIVE HYBRIDIZATION | Author keyword | 4 | 11% | 3% | 35 |
| 6 | RNA FINGERPRINTING | Author keyword | 4 | 35% | 1% | 9 |
| 7 | REPRESENTATIONAL DIFFERENCE ANALYSIS | Author keyword | 4 | 18% | 2% | 19 |
| 8 | MRNA DIFFERENTIAL DISPLAY PCR | Author keyword | 3 | 100% | 0% | 3 |
| 9 | DIFFERENTIAL DISPLAY REVERSE TRANSCRIPTION POLYMERASE CHAIN REACTION | Author keyword | 3 | 50% | 0% | 4 |
| 10 | SUBTRACTIVE CLONING | Author keyword | 3 | 29% | 1% | 8 |
Web of Science journal categories |
Author Key Words |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | CROSS COMBINATION | 23 | 74% | 2% | 17 |
| 2 | PHOTOACTIVATABLE BIOTIN | 17 | 100% | 1% | 8 |
| 3 | ARBITRARILY PRIMED PCR | 10 | 18% | 5% | 51 |
| 4 | DISPLAY TECHNIQUE | 9 | 83% | 0% | 5 |
| 5 | DDRT PCR | 8 | 70% | 1% | 7 |
| 6 | EXPRESSED MESSENGER RNA | 6 | 80% | 0% | 4 |
| 7 | X LARGE WHITE | 5 | 47% | 1% | 8 |
| 8 | EUKARYOTIC MESSENGER RNA | 5 | 17% | 2% | 26 |
| 9 | COMPETITIVE DNA REASSOCIATION | 4 | 75% | 0% | 3 |
| 10 | AUTOMATED DNA SEQUENCER | 2 | 44% | 0% | 4 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references |
% act. ref. to same field |
|---|---|---|---|---|
| Suppression subtractive hybridization: A versatile method for identifying differentially expressed genes | 1999 | 288 | 20 | 65% |
| RECENT ADVANCES IN DIFFERENTIAL DISPLAY | 1995 | 210 | 30 | 77% |
| Different strategies of differential display: areas of application | 1998 | 58 | 31 | 84% |
| A decade of differential display | 2002 | 73 | 61 | 39% |
| Large-scale tag/PCR-based gene expression profiling | 2014 | 1 | 48 | 38% |
| cDNA representational difference analysis: A sensitive and flexible method for identification of differentially expressed genes | 1999 | 81 | 11 | 45% |
| Differential display technology: a general guide | 2002 | 28 | 29 | 69% |
| Subtractive cloning: Past, present, and future | 1997 | 93 | 70 | 43% |
| RNA FINGERPRINTING AND DIFFERENTIAL DISPLAY USING ARBITRARILY PRIMED PCR | 1995 | 176 | 45 | 47% |
| Principles of differential display | 1999 | 40 | 28 | 68% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | ANIM NUTR FEED YUNNAN PROV | 11 | 54% | 1.3% | 14 |
| 2 | MOL GERONTOL UNIT | 4 | 67% | 0.4% | 4 |
| 3 | ANIM NUTR FEED YUNNAN PROVINCE | 2 | 67% | 0.2% | 2 |
| 4 | GENET CROP BREEDING | 1 | 33% | 0.2% | 2 |
| 5 | ACGT COMPUTAT BIOL BIOINFORMAT UNIT | 1 | 50% | 0.1% | 1 |
| 6 | ALMIRALL PRODESFARMA | 1 | 50% | 0.1% | 1 |
| 7 | BIOTECHNOL CLIMATE CHANGE | 1 | 50% | 0.1% | 1 |
| 8 | CENT HUMAN GENOME | 1 | 50% | 0.1% | 1 |
| 9 | CMIER DEV SYST BIOL | 1 | 50% | 0.1% | 1 |
| 10 | DIS ISTANCE GRP | 1 | 50% | 0.1% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000131730 | ALPHA ZAL//PRAP1//ALPHA ZEARALANOL |
| 2 | 0.0000116042 | SERIAL ANALYSIS OF GENE EXPRESSION//SAGE//GLGI |
| 3 | 0.0000105772 | MKIAA//SMART CDNA LIBRARY//CDNA SEQUENCING |
| 4 | 0.0000090707 | GENOME WALKING//FALSE POSITIVE EXCLUSION//UNKNOWN FLANKING SEQUENCES |
| 5 | 0.0000073579 | HN1//CANC 6401 E THOMAS RD//HORMONAL PROFILING |
| 6 | 0.0000064783 | TAISHO FUNCT GENOM//MULTIPHOTON DETECTION//ADAPTER TAGGED COMPETITIVE PCR |
| 7 | 0.0000063856 | FRAGMENT LENGTH DISTRIBUTION//AGR BIOTECHNOL RD//FLORAL CROP |
| 8 | 0.0000056208 | RNA AMPLIFICATION//LASER CAPTURE MICRODISSECTION//LASER MICRODISSECTION |
| 9 | 0.0000054512 | RNA ISOLATION//RNA EXTRACTION//DNA ISOLATION |
| 10 | 0.0000053420 | PLANT MICROBIAL GENOM//LRT1//NONADDITIVE PROTEIN |