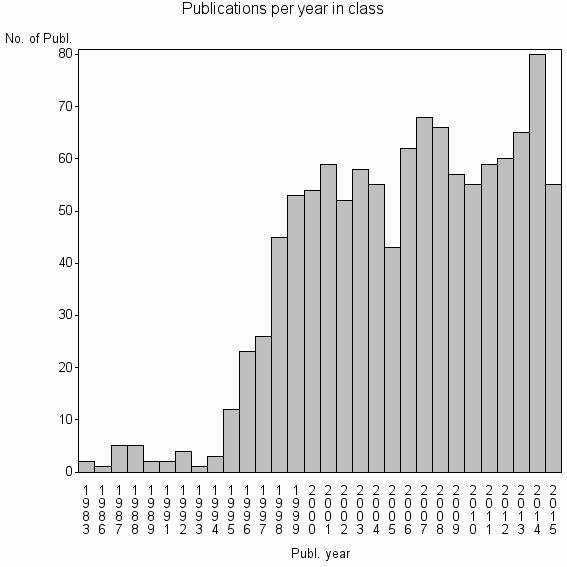
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 8974 | 1132 | 51.8 | 94% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | MYST | Author keyword | 34 | 93% | 1% | 13 |
| 2 | MOZ | Author keyword | 27 | 76% | 2% | 19 |
| 3 | HISTONE ACETYLTRANSFERASE | Author keyword | 24 | 28% | 7% | 75 |
| 4 | SAGA COMPLEX | Author keyword | 19 | 76% | 1% | 13 |
| 5 | GCN5 | Author keyword | 17 | 39% | 3% | 34 |
| 6 | BRPF1 | Author keyword | 12 | 86% | 1% | 6 |
| 7 | TIP60 | Author keyword | 9 | 30% | 2% | 25 |
| 8 | PCAF | Author keyword | 8 | 30% | 2% | 21 |
| 9 | T816 | Author keyword | 6 | 58% | 1% | 7 |
| 10 | SAGA | Author keyword | 6 | 24% | 2% | 21 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | MYST | 34 | 93% | 1% | 13 | Search MYST | Search MYST |
| 2 | MOZ | 27 | 76% | 2% | 19 | Search MOZ | Search MOZ |
| 3 | HISTONE ACETYLTRANSFERASE | 24 | 28% | 7% | 75 | Search HISTONE+ACETYLTRANSFERASE | Search HISTONE+ACETYLTRANSFERASE |
| 4 | SAGA COMPLEX | 19 | 76% | 1% | 13 | Search SAGA+COMPLEX | Search SAGA+COMPLEX |
| 5 | GCN5 | 17 | 39% | 3% | 34 | Search GCN5 | Search GCN5 |
| 6 | BRPF1 | 12 | 86% | 1% | 6 | Search BRPF1 | Search BRPF1 |
| 7 | TIP60 | 9 | 30% | 2% | 25 | Search TIP60 | Search TIP60 |
| 8 | PCAF | 8 | 30% | 2% | 21 | Search PCAF | Search PCAF |
| 9 | T816 | 6 | 58% | 1% | 7 | Search T816 | Search T816 |
| 10 | SAGA | 6 | 24% | 2% | 21 | Search SAGA | Search SAGA |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | GCN5 | 39 | 52% | 5% | 53 |
| 2 | CHROMATIN TRANSCRIPTION | 28 | 55% | 3% | 35 |
| 3 | PCAF | 27 | 53% | 3% | 36 |
| 4 | PUTATIVE TRANSCRIPTIONAL ADAPTER | 27 | 74% | 2% | 20 |
| 5 | CBP | 24 | 18% | 11% | 125 |
| 6 | LYSINE ACETYLTRANSFERASES | 24 | 91% | 1% | 10 |
| 7 | NUCLEOSOME ACETYLATION | 24 | 45% | 3% | 39 |
| 8 | ACIDIC ACTIVATION DOMAINS | 22 | 49% | 3% | 33 |
| 9 | MOZ | 22 | 60% | 2% | 24 |
| 10 | YEAST GCN5P | 21 | 54% | 2% | 27 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| 50 years of protein acetylation: from gene regulation to epigenetics, metabolism and beyond | 2015 | 6 | 93 | 17% |
| The diverse functions of histone acetyltransferase complexes | 2003 | 303 | 67 | 55% |
| Histone acetyltransferases | 2001 | 980 | 264 | 36% |
| ATAC-king the complexity of SAGA during evolution | 2012 | 32 | 131 | 50% |
| Histone acetyl transferases as emerging drug targets | 2009 | 102 | 63 | 49% |
| The MOZ Histone Acetyltransferase in Epigenetic Signaling and Disease | 2014 | 5 | 38 | 53% |
| Small Molecule Inhibitors of Histone Acetyltransferases as Epigenetic Tools and Drug Candidates | 2012 | 22 | 105 | 56% |
| Functions of site-specific histone acerylation and deacetylation | 2007 | 560 | 138 | 15% |
| Histone acetyltransferase complexes: one size doesn't fit all | 2007 | 345 | 114 | 24% |
| Structure and functions of the GNAT superfamily of acetyltransferases | 2005 | 240 | 73 | 30% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | TRANSCRIPT DIS | 4 | 22% | 1.4% | 16 |
| 2 | BIOCRYSTALLOG COMPUTAT MOL BIOL | 3 | 50% | 0.4% | 5 |
| 3 | ALTHOUSE 306 | 3 | 42% | 0.4% | 5 |
| 4 | PHARMACEUT GENE MODULAT | 2 | 24% | 0.7% | 8 |
| 5 | CHARLIE E MIDT BIOMED SCI | 2 | 67% | 0.2% | 2 |
| 6 | ONCOGENOM AIRC | 2 | 67% | 0.2% | 2 |
| 7 | HOTEL DIEU QUEBEC CHUQ | 2 | 23% | 0.7% | 8 |
| 8 | PROGRAM GENE EXP S REGULAT | 2 | 24% | 0.6% | 7 |
| 9 | CANC UK STEM CELL HAEMATOPOIESIS GRP | 1 | 38% | 0.3% | 3 |
| 10 | GRP GENE ENVIRONM BIOL | 1 | 100% | 0.2% | 2 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000151114 | NUT MIDLINE CARCINOMA//BROMODOMAIN//BRD4 |
| 2 | 0.0000125085 | ING4//ING1//P33ING1 |
| 3 | 0.0000123232 | RUBINSTEIN TAYBI SYNDROME//CORNEAL KELOID//TRISOMY 16Q |
| 4 | 0.0000117356 | HISTONE CHAPERONE//ASF1//CAF 1 |
| 5 | 0.0000111357 | SWI SNF//BRG1//ISWI |
| 6 | 0.0000110800 | AIB1//STEROID HORMONES SECT//RIP140 |
| 7 | 0.0000108751 | HISTONE DEMETHYLASE//HISTONE DEMETHYLATION//LSD1 |
| 8 | 0.0000103986 | TFIID//TFIIA//MOT1 |
| 9 | 0.0000100689 | CITED2//CITED//MELANOCYTE SPECIFIC GENE MSG1 |
| 10 | 0.0000079973 | REPTIN//PONTIN//R2TP |