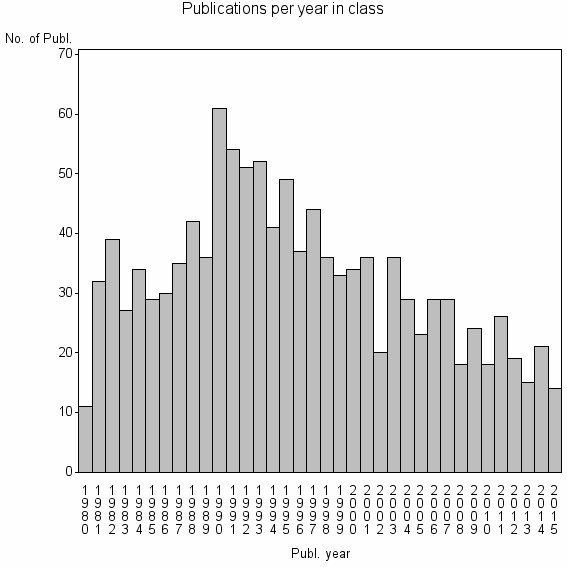
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 8638 | 1164 | 42.3 | 77% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 302 | 17871 | TRANSLESION SYNTHESIS//DNA POLYMERASE//RECA PROTEIN |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | RECA PROTEIN | Author keyword | 103 | 68% | 8% | 90 |
| 2 | RECA | Author keyword | 29 | 26% | 8% | 94 |
| 3 | DNA STRAND EXCHANGE | Author keyword | 10 | 47% | 1% | 16 |
| 4 | STRAND EXCHANGE | Author keyword | 10 | 35% | 2% | 22 |
| 5 | HOMOLOGOUS PAIRING | Author keyword | 7 | 39% | 1% | 14 |
| 6 | HOMOLOGOUS GENETIC RECOMBINATION | Author keyword | 6 | 80% | 0% | 4 |
| 7 | NUCLEOTIDE COFACTOR | Author keyword | 6 | 71% | 0% | 5 |
| 8 | RADA | Author keyword | 4 | 40% | 1% | 8 |
| 9 | RECX | Author keyword | 4 | 46% | 1% | 6 |
| 10 | GENOTOXICOL CYCLE CELLULAIRE | Address | 3 | 57% | 0% | 4 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | RECA PROTEIN | 103 | 68% | 8% | 90 | Search RECA+PROTEIN | Search RECA+PROTEIN |
| 2 | RECA | 29 | 26% | 8% | 94 | Search RECA | Search RECA |
| 3 | DNA STRAND EXCHANGE | 10 | 47% | 1% | 16 | Search DNA+STRAND+EXCHANGE | Search DNA+STRAND+EXCHANGE |
| 4 | STRAND EXCHANGE | 10 | 35% | 2% | 22 | Search STRAND+EXCHANGE | Search STRAND+EXCHANGE |
| 5 | HOMOLOGOUS PAIRING | 7 | 39% | 1% | 14 | Search HOMOLOGOUS+PAIRING | Search HOMOLOGOUS+PAIRING |
| 6 | HOMOLOGOUS GENETIC RECOMBINATION | 6 | 80% | 0% | 4 | Search HOMOLOGOUS+GENETIC+RECOMBINATION | Search HOMOLOGOUS+GENETIC+RECOMBINATION |
| 7 | NUCLEOTIDE COFACTOR | 6 | 71% | 0% | 5 | Search NUCLEOTIDE+COFACTOR | Search NUCLEOTIDE+COFACTOR |
| 8 | RADA | 4 | 40% | 1% | 8 | Search RADA | Search RADA |
| 9 | RECX | 4 | 46% | 1% | 6 | Search RECX | Search RECX |
| 10 | FLUORESCENT DYE LABELED OLIGONUCLEOTIDE | 3 | 100% | 0% | 3 | Search FLUORESCENT+DYE+LABELED+OLIGONUCLEOTIDE | Search FLUORESCENT+DYE+LABELED+OLIGONUCLEOTIDE |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | PROMOTED ATP HYDROLYSIS | 68 | 100% | 2% | 22 |
| 2 | NUCLEOPROTEIN FILAMENTS | 59 | 56% | 6% | 73 |
| 3 | HOMOLOGOUS RECOGNITION | 53 | 95% | 2% | 18 |
| 4 | DNA STRAND EXCHANGE | 51 | 40% | 9% | 100 |
| 5 | 3 STRANDED DNA | 42 | 64% | 4% | 41 |
| 6 | ESCHERICHIA COLI RECA | 40 | 30% | 9% | 109 |
| 7 | GENETIC RECOMBINATION | 38 | 18% | 17% | 195 |
| 8 | HETEROLOGOUS INSERT | 26 | 100% | 1% | 11 |
| 9 | COLI RECA PROTEIN | 23 | 41% | 4% | 43 |
| 10 | STABLE COMPLEXES | 22 | 63% | 2% | 22 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Motoring along with the bacterial RecA protein | 2007 | 91 | 102 | 72% |
| The bacterial REcA protein and the recombinational DNA repair of stalled replication forks | 2002 | 239 | 145 | 70% |
| HOMOLOGOUS PAIRING AND DNA STRAND-EXCHANGE PROTEINS | 1994 | 241 | 207 | 66% |
| RecA protein: Structure, function, and role in recombinational DNA repair | 1997 | 304 | 374 | 50% |
| The bacterial RecA protein as a motor protein | 2003 | 121 | 114 | 68% |
| Structure and mechanism of Escherichia coli RecA ATPase | 2005 | 55 | 56 | 75% |
| BIOCHEMISTRY OF GENETIC-RECOMBINATION - ENERGETICS AND MECHANISM OF DNA STRAND EXCHANGE | 1991 | 222 | 113 | 87% |
| THE RECA PROTEIN - STRUCTURE AND FUNCTION | 1990 | 373 | 243 | 71% |
| HELICAL INTERACTIONS IN HOMOLOGOUS PAIRING AND STRAND EXCHANGE DRIVEN BY RECA PROTEIN | 1991 | 173 | 57 | 89% |
| Molecular design and functional organization of the RecA protein | 2003 | 74 | 178 | 61% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | GENOTOXICOL CYCLE CELLULAIRE | 3 | 57% | 0.3% | 4 |
| 2 | EDUC BIOPHYS | 2 | 40% | 0.3% | 4 |
| 3 | MOL RADIAT BIOPHYS | 2 | 12% | 1.2% | 14 |
| 4 | HUMAN GENE 2 | 1 | 100% | 0.2% | 2 |
| 5 | AGR LIFE SCI BIOCHEM | 1 | 19% | 0.4% | 5 |
| 6 | UNITE MIXTE RECH 216 | 1 | 33% | 0.2% | 2 |
| 7 | YASHIMA SUPERSTRUCT HELIX PROJECT | 1 | 33% | 0.2% | 2 |
| 8 | FRE 2230 | 1 | 17% | 0.3% | 4 |
| 9 | UMR 2027 | 1 | 11% | 0.5% | 6 |
| 10 | ANAL ULTRASTRUCT BATIMENT BIOL | 1 | 50% | 0.1% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000255217 | SOS MUTAGENESIS//UMUDC//LEXA REPRESSOR |
| 2 | 0.0000237209 | HOLLIDAY JUNCTION//RECBCD ENZYME//RECBCD |
| 3 | 0.0000200920 | KIN17 PROTEIN//GENET RADIOSENSIBIL//GENET RADIOSENSIBILITE |
| 4 | 0.0000172280 | RAD51//HOMOLOGOUS RECOMBINATION//RAD52 |
| 5 | 0.0000128451 | REPLICATION PROTEIN A//SINGLE STRANDED DNA BINDING PROTEIN//REPLICATION PROTEIN A RPA |
| 6 | 0.0000110531 | SINGLE STRANDED OLIGONUCLEOTIDE//GENE REPAIR//SINGLE STRANDED OLIGONUCLEOTIDES |
| 7 | 0.0000086129 | OBSTET GYNECOLSEALY STRUCT BIOL//PRIMASE//HELICASE |
| 8 | 0.0000080667 | DEINOCOCCUS RADIODURANS//DEINOCOCCUS//DEINOCOCCACEAE |
| 9 | 0.0000069342 | WARWICK ANALYT SCI//GELRED//DNA TOPOISOMERS |
| 10 | 0.0000054008 | BMSE PROGRAM//MAGNETIC TWEEZERS//SMALL BIOSYST |