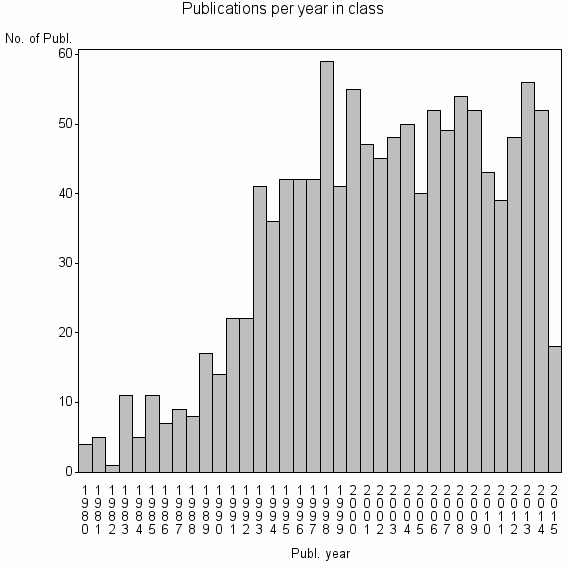
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 8408 | 1187 | 32.3 | 63% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | THREE DIMENSIONAL HOLOGRAPHIC VECTOR OF ATOMIC INTERACTION FIELD 3D HOVAIF | Author keyword | 21 | 90% | 1% | 9 |
| 2 | COMFA | Author keyword | 19 | 13% | 11% | 129 |
| 3 | 3D QSAR | Author keyword | 15 | 11% | 11% | 126 |
| 4 | MULTI DIMENSIONAL QSAR | Author keyword | 14 | 100% | 1% | 7 |
| 5 | MULTIDIMENSIONAL QSAR | Author keyword | 12 | 86% | 1% | 6 |
| 6 | COMSA | Author keyword | 11 | 100% | 1% | 6 |
| 7 | QUANTITATIVE SEQUENCE ACTIVITY MODEL | Author keyword | 11 | 100% | 1% | 6 |
| 8 | 4D QSAR | Author keyword | 10 | 52% | 1% | 14 |
| 9 | QUANTITATIVE STRUCTURE-ACTIVITY RELATIONSHIPS | Journal | 10 | 14% | 6% | 68 |
| 10 | VLIFE MDS | Author keyword | 9 | 83% | 0% | 5 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | THREE DIMENSIONAL HOLOGRAPHIC VECTOR OF ATOMIC INTERACTION FIELD 3D HOVAIF | 21 | 90% | 1% | 9 | Search THREE+DIMENSIONAL+HOLOGRAPHIC+VECTOR+OF+ATOMIC+INTERACTION+FIELD+3D+HOVAIF | Search THREE+DIMENSIONAL+HOLOGRAPHIC+VECTOR+OF+ATOMIC+INTERACTION+FIELD+3D+HOVAIF |
| 2 | COMFA | 19 | 13% | 11% | 129 | Search COMFA | Search COMFA |
| 3 | 3D QSAR | 15 | 11% | 11% | 126 | Search 3D+QSAR | Search 3D+QSAR |
| 4 | MULTI DIMENSIONAL QSAR | 14 | 100% | 1% | 7 | Search MULTI+DIMENSIONAL+QSAR | Search MULTI+DIMENSIONAL+QSAR |
| 5 | MULTIDIMENSIONAL QSAR | 12 | 86% | 1% | 6 | Search MULTIDIMENSIONAL+QSAR | Search MULTIDIMENSIONAL+QSAR |
| 6 | COMSA | 11 | 100% | 1% | 6 | Search COMSA | Search COMSA |
| 7 | QUANTITATIVE SEQUENCE ACTIVITY MODEL | 11 | 100% | 1% | 6 | Search QUANTITATIVE+SEQUENCE+ACTIVITY+MODEL | Search QUANTITATIVE+SEQUENCE+ACTIVITY+MODEL |
| 8 | 4D QSAR | 10 | 52% | 1% | 14 | Search 4D+QSAR | Search 4D+QSAR |
| 9 | VLIFE MDS | 9 | 83% | 0% | 5 | Search VLIFE+MDS | Search VLIFE+MDS |
| 10 | COMSIA | 8 | 11% | 5% | 64 | Search COMSIA | Search COMSIA |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | THERMOLYSIN INHIBITORS | 50 | 81% | 3% | 30 |
| 2 | MOLECULAR FIELD ANALYSIS | 40 | 21% | 15% | 174 |
| 3 | COMFA | 40 | 26% | 11% | 132 |
| 4 | ANALYSIS COMFA | 31 | 52% | 4% | 42 |
| 5 | 4D QSAR | 27 | 92% | 1% | 11 |
| 6 | 3D QSAR | 26 | 20% | 10% | 120 |
| 7 | ALIGNMENT RULES | 26 | 87% | 1% | 13 |
| 8 | PETT DERIVATIVES | 24 | 91% | 1% | 10 |
| 9 | 3 DIMENSIONAL QSAR | 19 | 80% | 1% | 12 |
| 10 | ATOMIC INTERACTION FIELD | 18 | 83% | 1% | 10 |
Journals |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | QUANTITATIVE STRUCTURE-ACTIVITY RELATIONSHIPS | 10 | 14% | 6% | 68 |
| 2 | PERSPECTIVES IN DRUG DISCOVERY AND DESIGN | 2 | 12% | 1% | 17 |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| 3D-QSAR in Drug Design - A Review | 2010 | 112 | 63 | 46% |
| Multi-dimensional QSAR in drug discovery | 2007 | 48 | 27 | 48% |
| How to Recognize and Workaround Pitfalls in QSAR Studies: A Critical Review | 2009 | 71 | 48 | 27% |
| QSAR and 3D QSAR in drug design .1. methodology | 1997 | 191 | 15 | 53% |
| The CoMFA steroids as a benchmark dataset for development of 3D QSAR methods | 1998 | 59 | 20 | 90% |
| How to Generate Reliable and Predictive CoMFA Models | 2011 | 13 | 89 | 38% |
| 4D-QSAR: Perspectives in Drug Design | 2010 | 28 | 39 | 31% |
| Receptor Dependent Multidimensional QSAR for Modeling Drug - Receptor Interactions | 2009 | 16 | 102 | 44% |
| Comparative molecular similarity indices analysis: CoMSIA | 1998 | 86 | 11 | 45% |
| QSAR Modeling for Quinoxaline Derivatives using Genetic Algorithm and Simulated Annealing Based Feature Selection | 2009 | 26 | 23 | 30% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | MOL MODELING DESIGN MC781 | 6 | 71% | 0.4% | 5 |
| 2 | GRP PHARMACOCHEM | 2 | 30% | 0.5% | 6 |
| 3 | MALMO OFF | 1 | 50% | 0.2% | 2 |
| 4 | STATE CHEMOBIOSENSORS CHEMOBIOMETR MOST | 1 | 100% | 0.2% | 2 |
| 5 | TECHNOL LIFE SCI BIOTECHNOL | 1 | 100% | 0.2% | 2 |
| 6 | PHARMACEUT CHEM UNIT | 1 | 33% | 0.2% | 2 |
| 7 | BIOMED ENGN EDUC MINIST CHONGQING CITY | 1 | 50% | 0.1% | 1 |
| 8 | COMP CHEM UNIT | 1 | 50% | 0.1% | 1 |
| 9 | COMPUTAT BIOL MOL BIOPHYS | 1 | 50% | 0.1% | 1 |
| 10 | ENVIROMENTAL COMPUTAT CHEM GRP | 1 | 50% | 0.1% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000230926 | CP MLR//HAM 3//COMBINATORIAL PROTOCOL IN MULTIPLE LINEAR REGRESSION |
| 2 | 0.0000198666 | OPTIMAL DESCRIPTOR//CORAL SOFTWARE//MIA QSAR |
| 3 | 0.0000107460 | B IT//LIFE SCI INFORMAT//LIMES PROGRAM UNIT CHEM BIOL MED CHEM |
| 4 | 0.0000106680 | JOURNAL OF CHEMICAL INFORMATION AND MODELING//VIRTUAL SCREENING//JOURNAL OF COMPUTER-AIDED MOLECULAR DESIGN |
| 5 | 0.0000103691 | SERAMS//MATH CHEM//ESTROGEN RECEPTOR LIGANDS |
| 6 | 0.0000102702 | QUANTUM SIMILARITY//VECTOR SEMISPACES//MATH CHEM UNIT |
| 7 | 0.0000102585 | TOMOCOMD CARDD SOFTWARE//CHEM BIOACT//UNIDAD INVEST DISENO FARMACOS CONECTIVIDAD MOL |
| 8 | 0.0000098684 | SAR AND QSAR IN ENVIRONMENTAL RESEARCH//TETRAHYMENA PYRIFORMIS//EXCESS TOXICITY |
| 9 | 0.0000070129 | LINEAR SOLVATION ENERGY THEORY//N OCTANOL WATER PARTITION COEFFICIENT LGKOW//TLSER |
| 10 | 0.0000062029 | INFINITE DILUTION NMR//KLOPMAN ATOMIC SOFTNESS//LOCALIZED DILUTION SHIFTS |