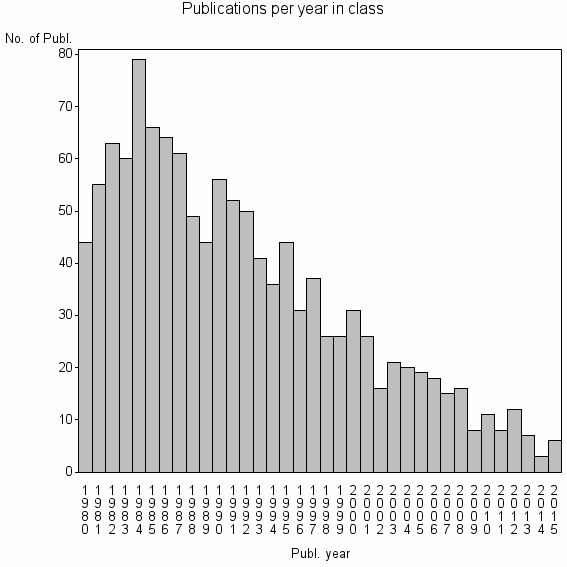
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 8072 | 1221 | 38.7 | 62% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | REPLICATION CONTROL | Author keyword | 21 | 61% | 2% | 22 |
| 2 | PLASMID R6K | Author keyword | 17 | 100% | 1% | 8 |
| 3 | PLASMID RK2 | Author keyword | 11 | 67% | 1% | 10 |
| 4 | RNA II | Author keyword | 11 | 100% | 0% | 6 |
| 5 | PI PROTEIN | Author keyword | 8 | 75% | 0% | 6 |
| 6 | R6K | Author keyword | 8 | 100% | 0% | 5 |
| 7 | PLASMID REPLICATION | Author keyword | 7 | 28% | 2% | 22 |
| 8 | PLASMID COPY NUMBER CONTROL | Author keyword | 7 | 67% | 0% | 6 |
| 9 | REPA PROTEIN | Author keyword | 6 | 80% | 0% | 4 |
| 10 | ROM PROTEIN | Author keyword | 6 | 80% | 0% | 4 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | REPLICATION CONTROL | 21 | 61% | 2% | 22 | Search REPLICATION+CONTROL | Search REPLICATION+CONTROL |
| 2 | PLASMID R6K | 17 | 100% | 1% | 8 | Search PLASMID+R6K | Search PLASMID+R6K |
| 3 | PLASMID RK2 | 11 | 67% | 1% | 10 | Search PLASMID+RK2 | Search PLASMID+RK2 |
| 4 | RNA II | 11 | 100% | 0% | 6 | Search RNA+II | Search RNA+II |
| 5 | PI PROTEIN | 8 | 75% | 0% | 6 | Search PI+PROTEIN | Search PI+PROTEIN |
| 6 | R6K | 8 | 100% | 0% | 5 | Search R6K | Search R6K |
| 7 | PLASMID REPLICATION | 7 | 28% | 2% | 22 | Search PLASMID+REPLICATION | Search PLASMID+REPLICATION |
| 8 | PLASMID COPY NUMBER CONTROL | 7 | 67% | 0% | 6 | Search PLASMID+COPY+NUMBER+CONTROL | Search PLASMID+COPY+NUMBER+CONTROL |
| 9 | REPA PROTEIN | 6 | 80% | 0% | 4 | Search REPA+PROTEIN | Search REPA+PROTEIN |
| 10 | ROM PROTEIN | 6 | 80% | 0% | 4 | Search ROM+PROTEIN | Search ROM+PROTEIN |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | HOST RANGE PLASMID RK2 | 59 | 90% | 2% | 26 |
| 2 | COPY NUMBER CONTROL | 52 | 55% | 5% | 66 |
| 3 | VEGETATIVE REPLICATION | 38 | 86% | 2% | 19 |
| 4 | TRFA | 29 | 77% | 2% | 20 |
| 5 | RK2 REPLICATION | 27 | 92% | 1% | 11 |
| 6 | KEY REGULATORY GENE | 24 | 91% | 1% | 10 |
| 7 | REPZ GENE EXPRESSION | 23 | 86% | 1% | 12 |
| 8 | RSF1040 | 23 | 100% | 1% | 10 |
| 9 | PRIMER TRANSCRIPT | 22 | 49% | 3% | 32 |
| 10 | GAMMA ORIGIN | 21 | 90% | 1% | 9 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Plasmid R6K replication control | 2013 | 4 | 94 | 88% |
| Control of plasmid DNA replication by iterons: no longer paradoxical | 2000 | 111 | 66 | 71% |
| Plasmid R1 - Replication and its control | 2006 | 38 | 112 | 50% |
| COMPLETE NUCLEOTIDE-SEQUENCE OF BIRMINGHAM INCP-ALPHA PLASMIDS - COMPILATION AND COMPARATIVE-ANALYSIS | 1994 | 268 | 169 | 56% |
| ANTISENSE RNA | 1991 | 240 | 106 | 49% |
| ANTISENSE RNA CONTROL IN BACTERIA, PHAGES, AND PLASMIDS | 1994 | 292 | 171 | 47% |
| IDENTIFICATION AND CLASSIFICATION OF BACTERIAL PLASMIDS | 1988 | 271 | 99 | 79% |
| Replication and partitioning of the broad-host-range plasmid RK2 | 2010 | 10 | 164 | 70% |
| Twenty years of the pPS10 replicon: insights on the molecular mechanism for the activation of DNA replication in iteron-containing bacterial plasmids | 2004 | 39 | 51 | 67% |
| Antisense RNAs in bacteria and their genetic elements | 2002 | 128 | 201 | 47% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | SCI BIOL GENET | 4 | 75% | 0.2% | 3 |
| 2 | GOSNIIGENETIKA | 1 | 50% | 0.1% | 1 |
| 3 | IHNESTR 73 | 1 | 50% | 0.1% | 1 |
| 4 | MAX DELBRUCK MOL MED BERLIN BUCH | 1 | 50% | 0.1% | 1 |
| 5 | MOLEC MICROBIOL BIOTECHNOL | 0 | 33% | 0.1% | 1 |
| 6 | ARBEITSGRP GENET | 0 | 18% | 0.2% | 2 |
| 7 | UNITE POSTULANTE PLAST GENOME BACTERIEN | 0 | 17% | 0.1% | 1 |
| 8 | HALL ATWATER SHANKLIN S | 0 | 14% | 0.1% | 1 |
| 9 | INITIAT BIOINFORMAT EVOLUTIONARY STUDIES IBEST | 0 | 14% | 0.1% | 1 |
| 10 | SUAGM | 0 | 14% | 0.1% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000231256 | DNAA PROTEIN//DNAA//ORIC |
| 2 | 0.0000229964 | ROLLING CIRCLE REPLICATION//PLASMID REPLICATION CONTROL//CRYPTIC PLASMID |
| 3 | 0.0000208302 | BACTERIAL CONJUGATION//COUPLING PROTEIN//R391 |
| 4 | 0.0000190214 | PARB//DNA SEGREGATION//FTSK |
| 5 | 0.0000125983 | TOXIN ANTITOXIN//TOXIN ANTITOXIN SYSTEM//TOXIN ANTITOXIN SYSTEMS |
| 6 | 0.0000123279 | TELLURITE RESISTANCE//TELLURITE REDUCTION//TELLURITE |
| 7 | 0.0000112072 | POLYLINKER//STREPTOMYCIN SENSITIVITY//INSERTIONAL INACTIVATION |
| 8 | 0.0000089650 | PLASMID TRANSFER//GENETIC BIOAUGMENTATION//GENETICALLY ENGINEERED MICROORGANISMS |
| 9 | 0.0000083711 | SKIRBALL PROGRAM MOL PATHOGENESIS//BACTERIOPHAGE P2//RETRON |
| 10 | 0.0000081492 | DNA TRANSPOSITION//PHAGE MU//TN10 |