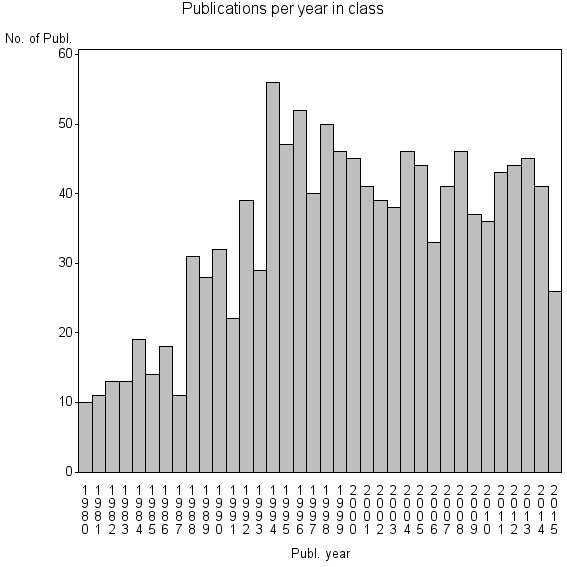
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 8022 | 1226 | 43.1 | 80% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 501 | 14604 | RNA POLYMERASE//SITE SPECIFIC RECOMBINATION//H NS |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | H NS | Author keyword | 51 | 54% | 5% | 67 |
| 2 | HISTONE LIKE PROTEIN | Author keyword | 36 | 66% | 3% | 33 |
| 3 | BACTERIAL CHROMATIN | Author keyword | 23 | 100% | 1% | 10 |
| 4 | PHYSIOL BACTERIENNE | Address | 22 | 81% | 1% | 13 |
| 5 | HU PROTEIN | Author keyword | 19 | 63% | 2% | 19 |
| 6 | NUCLEOID ASSOCIATED PROTEIN | Author keyword | 16 | 64% | 1% | 16 |
| 7 | LEIDEN CHEM CELL OBSERV | Address | 14 | 100% | 1% | 7 |
| 8 | HHA | Author keyword | 11 | 56% | 1% | 14 |
| 9 | INTEGRATION HOST FACTOR | Author keyword | 11 | 40% | 2% | 22 |
| 10 | HU | Author keyword | 10 | 29% | 2% | 28 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | H NS | 51 | 54% | 5% | 67 | Search H+NS | Search H+NS |
| 2 | HISTONE LIKE PROTEIN | 36 | 66% | 3% | 33 | Search HISTONE+LIKE+PROTEIN | Search HISTONE+LIKE+PROTEIN |
| 3 | BACTERIAL CHROMATIN | 23 | 100% | 1% | 10 | Search BACTERIAL+CHROMATIN | Search BACTERIAL+CHROMATIN |
| 4 | HU PROTEIN | 19 | 63% | 2% | 19 | Search HU+PROTEIN | Search HU+PROTEIN |
| 5 | NUCLEOID ASSOCIATED PROTEIN | 16 | 64% | 1% | 16 | Search NUCLEOID+ASSOCIATED+PROTEIN | Search NUCLEOID+ASSOCIATED+PROTEIN |
| 6 | HHA | 11 | 56% | 1% | 14 | Search HHA | Search HHA |
| 7 | INTEGRATION HOST FACTOR | 11 | 40% | 2% | 22 | Search INTEGRATION+HOST+FACTOR | Search INTEGRATION+HOST+FACTOR |
| 8 | HU | 10 | 29% | 2% | 28 | Search HU | Search HU |
| 9 | H NS PROTEIN | 9 | 83% | 0% | 5 | Search H+NS+PROTEIN | Search H+NS+PROTEIN |
| 10 | TRANSCRIPTION FACTOR 1 | 9 | 67% | 1% | 8 | Search TRANSCRIPTION+FACTOR+1 | Search TRANSCRIPTION+FACTOR+1 |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | HISTONE LIKE PROTEINS | 92 | 51% | 10% | 128 |
| 2 | HISTONE LIKE PROTEIN | 46 | 45% | 6% | 76 |
| 3 | IHF | 43 | 51% | 5% | 60 |
| 4 | ESCHERICHIA COLI HU | 41 | 85% | 2% | 22 |
| 5 | BACTERIAL CHROMATIN | 36 | 63% | 3% | 36 |
| 6 | NUCLEOID ASSOCIATED PROTEINS | 36 | 50% | 4% | 52 |
| 7 | NUCLEOID ASSOCIATED PROTEIN | 34 | 48% | 4% | 52 |
| 8 | METHANOTHERMUS FERVIDUS | 33 | 45% | 4% | 55 |
| 9 | SAC7D | 32 | 85% | 1% | 17 |
| 10 | SSO7D | 27 | 83% | 1% | 15 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Integrated circuits: how transcriptional silencing and counter-silencing facilitate bacterial evolution | 2015 | 2 | 74 | 55% |
| H-NS: A universal regulator for a dynamic genome | 2004 | 270 | 104 | 65% |
| Bacterial nucleoid-associated proteins, nucleoid structure and gene expression | 2010 | 195 | 127 | 42% |
| New insights into transcriptional regulation by H-NS | 2008 | 103 | 65 | 63% |
| IHF and HU: flexible architects of bent DNA | 2004 | 175 | 60 | 82% |
| The role of nucleoid-associated proteins in the organization and compaction of bacterial chromatin | 2005 | 168 | 96 | 55% |
| Anti-silencing: overcoming H-NS-mediated repression of transcription in Gram-negative enteric bacteria | 2008 | 90 | 139 | 52% |
| H-NS, the genome sentinel | 2007 | 136 | 83 | 55% |
| Functional Evolution of Bacterial Histone-Like HU Proteins | 2011 | 26 | 104 | 70% |
| Effects of nucleoid-associated proteins on bacterial chromosome structure and gene expression | 2010 | 85 | 54 | 57% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | PHYSIOL BACTERIENNE | 22 | 81% | 1.1% | 13 |
| 2 | LEIDEN CHEM CELL OBSERV | 14 | 100% | 0.6% | 7 |
| 3 | AUTOMAT INTELLIGENT SCI | 2 | 67% | 0.2% | 2 |
| 4 | FRE2326 | 2 | 67% | 0.2% | 2 |
| 5 | PROGRAM BIOTECHNOL BIOENGN | 2 | 40% | 0.3% | 4 |
| 6 | PLASMA MEMBRANE NUCL SIGNALING | 2 | 15% | 0.8% | 10 |
| 7 | CELL OBSERV | 1 | 50% | 0.2% | 2 |
| 8 | CNRS URA 409 | 1 | 100% | 0.2% | 2 |
| 9 | PROGRAM BIOTECHNOL SCI ENGN | 1 | 100% | 0.2% | 2 |
| 10 | DYNAM EVOLUT EXP S GENOMES MICROORGANISME | 1 | 30% | 0.2% | 3 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000133615 | PPGPP//STRINGENT RESPONSE//RPOS |
| 2 | 0.0000127822 | PARB//DNA SEGREGATION//FTSK |
| 3 | 0.0000114512 | AIST TSUKUBA 6 10//LEUCINE RESPONSIVE REGULATORY PROTEIN//ASNC |
| 4 | 0.0000093600 | DNA GYRASE//GYRASE//REVERSE GYRASE |
| 5 | 0.0000092286 | SITE SPECIFIC RECOMBINATION//SERINE RECOMBINASE//PHI C31 INTEGRASE |
| 6 | 0.0000081034 | EPIDEMIOLOGICAL SIGNIFICANCE//SIMILARITY IDENTIFICATION//AIRBORNE ESCHERICHIA COLI |
| 7 | 0.0000071774 | ENVZ//OMPR//ENVZ OSMOSENSOR |
| 8 | 0.0000068165 | DNAA PROTEIN//DNAA//ORIC |
| 9 | 0.0000063264 | CYCLIN B HOMOLOGUE//CRYPTHECODINIUM COHNII//ANABAENA SPECIES |
| 10 | 0.0000061510 | PROGRAM COMPUTAT GENOM//CIFN//PROGRAMA GENOM COMPUTAC |