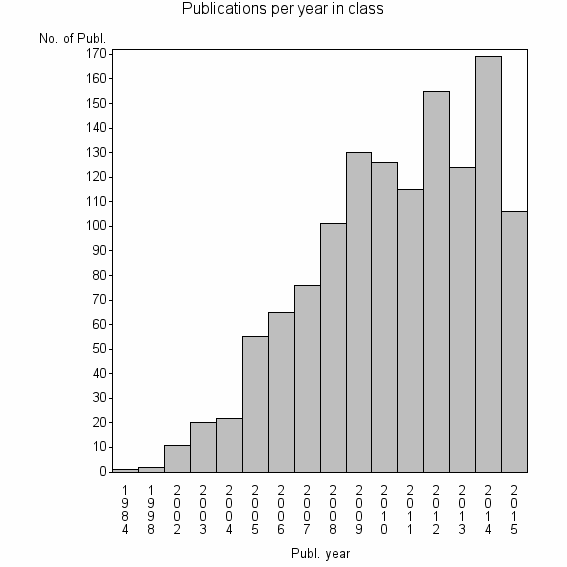
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 7531 | 1278 | 46.7 | 89% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 480 | 14980 | GENETIC EPIDEMIOLOGY//GENOMIC SELECTION//BMC GENETICS |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | EQTL | Author keyword | 24 | 37% | 4% | 53 |
| 2 | GENETICAL GENOMICS | Author keyword | 12 | 50% | 1% | 18 |
| 3 | ALLELE SPECIFIC GENE EXPRESSION | Author keyword | 12 | 86% | 0% | 6 |
| 4 | EXPRESSION QUANTITATIVE TRAIT LOCI | Author keyword | 6 | 40% | 1% | 12 |
| 5 | TRANS REGULATION | Author keyword | 6 | 50% | 1% | 8 |
| 6 | GRONINGEN BIOINFORMAT | Address | 5 | 17% | 2% | 28 |
| 7 | EXPRESSION QUANTITATIVE TRAIT LOCUS | Author keyword | 4 | 47% | 1% | 7 |
| 8 | ALLELE SPECIFIC EXPRESSION | Author keyword | 4 | 21% | 1% | 16 |
| 9 | SYSTEMS GENETICS | Author keyword | 4 | 26% | 1% | 12 |
| 10 | EQTL MAPPING | Author keyword | 3 | 57% | 0% | 4 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | EQTL | 24 | 37% | 4% | 53 | Search EQTL | Search EQTL |
| 2 | GENETICAL GENOMICS | 12 | 50% | 1% | 18 | Search GENETICAL+GENOMICS | Search GENETICAL+GENOMICS |
| 3 | ALLELE SPECIFIC GENE EXPRESSION | 12 | 86% | 0% | 6 | Search ALLELE+SPECIFIC+GENE+EXPRESSION | Search ALLELE+SPECIFIC+GENE+EXPRESSION |
| 4 | EXPRESSION QUANTITATIVE TRAIT LOCI | 6 | 40% | 1% | 12 | Search EXPRESSION+QUANTITATIVE+TRAIT+LOCI | Search EXPRESSION+QUANTITATIVE+TRAIT+LOCI |
| 5 | TRANS REGULATION | 6 | 50% | 1% | 8 | Search TRANS+REGULATION | Search TRANS+REGULATION |
| 6 | EXPRESSION QUANTITATIVE TRAIT LOCUS | 4 | 47% | 1% | 7 | Search EXPRESSION+QUANTITATIVE+TRAIT+LOCUS | Search EXPRESSION+QUANTITATIVE+TRAIT+LOCUS |
| 7 | ALLELE SPECIFIC EXPRESSION | 4 | 21% | 1% | 16 | Search ALLELE+SPECIFIC+EXPRESSION | Search ALLELE+SPECIFIC+EXPRESSION |
| 8 | SYSTEMS GENETICS | 4 | 26% | 1% | 12 | Search SYSTEMS+GENETICS | Search SYSTEMS+GENETICS |
| 9 | EQTL MAPPING | 3 | 57% | 0% | 4 | Search EQTL+MAPPING | Search EQTL+MAPPING |
| 10 | EXPRESSION EVOLUTION | 3 | 57% | 0% | 4 | Search EXPRESSION+EVOLUTION | Search EXPRESSION+EVOLUTION |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | HUMAN GENE EXPRESSION | 96 | 51% | 11% | 135 |
| 2 | REGULATORY VARIATION | 64 | 65% | 5% | 61 |
| 3 | INTEGRATIVE GENOMICS APPROACH | 42 | 74% | 2% | 31 |
| 4 | EQTL | 26 | 87% | 1% | 13 |
| 5 | PHARMACOGENOMIC DISCOVERY | 26 | 87% | 1% | 13 |
| 6 | SEGREGATING MOUSE POPULATION | 15 | 73% | 1% | 11 |
| 7 | ARABINOSIDE CYTOTOXICITY | 12 | 86% | 0% | 6 |
| 8 | TRANSCRIPT LEVEL VARIATION | 11 | 60% | 1% | 12 |
| 9 | ETOPOSIDE INDUCED CYTOTOXICITY | 11 | 67% | 1% | 10 |
| 10 | CISPLATIN INDUCED CYTOTOXICITY | 10 | 52% | 1% | 14 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| The role of regulatory variation in complex traits and disease | 2015 | 5 | 218 | 59% |
| Systems genetics approaches to understand complex traits | 2014 | 44 | 119 | 39% |
| Mapping complex disease traits with global gene expression | 2009 | 306 | 98 | 42% |
| Molecular networks as sensors and drivers of common human diseases | 2009 | 279 | 46 | 43% |
| Methods of integrating data to uncover genotype-phenotype interactions | 2015 | 9 | 84 | 19% |
| Genetics of human gene expression: mapping DNA variants that influence gene expression | 2009 | 114 | 70 | 64% |
| Genome-wide allele-specific analysis: insights into regulatory variation | 2010 | 84 | 32 | 69% |
| Genetics of global gene expression | 2006 | 248 | 99 | 45% |
| From genome to function by studying eQTLs | 2014 | 9 | 70 | 70% |
| Genome-Wide Significant Loci: How Important Are They? Systems Genetics to Understand Heritability of Coronary Artery Disease and Other Common Complex Disorders | 2015 | 3 | 84 | 40% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | GRONINGEN BIOINFORMAT | 5 | 17% | 2.2% | 28 |
| 2 | UMR598 | 3 | 60% | 0.2% | 3 |
| 3 | BIOINFORMAT EXPERTISE | 2 | 67% | 0.2% | 2 |
| 4 | HUMAN CENTRIFUGE MED TRAINING | 2 | 67% | 0.2% | 2 |
| 5 | MOL NEUROSCI NEURODEGENERAT DIS | 2 | 67% | 0.2% | 2 |
| 6 | CARDIOVASC EPIDEMIOL HUMAN GENOM BRANCH | 2 | 30% | 0.5% | 6 |
| 7 | COMM CLIN PHARMACOL PHARMACOGENOM | 2 | 12% | 1.2% | 15 |
| 8 | SECT GENET MED | 2 | 28% | 0.4% | 5 |
| 9 | ICAHN GENOM MULTISCALE BIOL | 2 | 14% | 0.8% | 10 |
| 10 | COMM CLIN PHARMACOL PHARMACOGEN | 2 | 14% | 0.8% | 10 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000125366 | NSY MOUSE//NEW ZEALAND OBESE MOUSE//MOUSE GENOM OURCE |
| 2 | 0.0000124559 | PLANT MICROBIAL GENOM//LRT1//NONADDITIVE PROTEIN |
| 3 | 0.0000109493 | RARE VARIANTS//GENETIC EPIDEMIOLOGY//RARE VARIANT ANALYSIS |
| 4 | 0.0000095315 | SEX BIASED GENE EXPRESSION//SEXUAL ANTAGONISM//INTRALOCUS SEXUAL CONFLICT |
| 5 | 0.0000080119 | INTERVAL MAPPING//SYSTEMS MAPPING//SELECTIVE GENOTYPING |
| 6 | 0.0000072939 | RNA SEQ//DE NOVO ASSEMBLY//TRANSCRIPTOME ASSEMBLY |
| 7 | 0.0000069391 | CHIP SEQ//PEAK CALLING//DNASE SEQ |
| 8 | 0.0000063171 | MERIEUX//BLOOD TRANSCRIPTOME//DUTCH NUTRIGENOM CONSORTIUM |
| 9 | 0.0000060360 | DIFFERENTIAL GENE EXPRESSION DETECTION//CANCER OUTLIER PROFILE ANALYSIS//BREAST CANC DIAG TREATMENT HENAN |
| 10 | 0.0000058611 | NETWORK INFERENCE//GENE REGULATORY NETWORKS//GENE NETWORK INFERENCE |