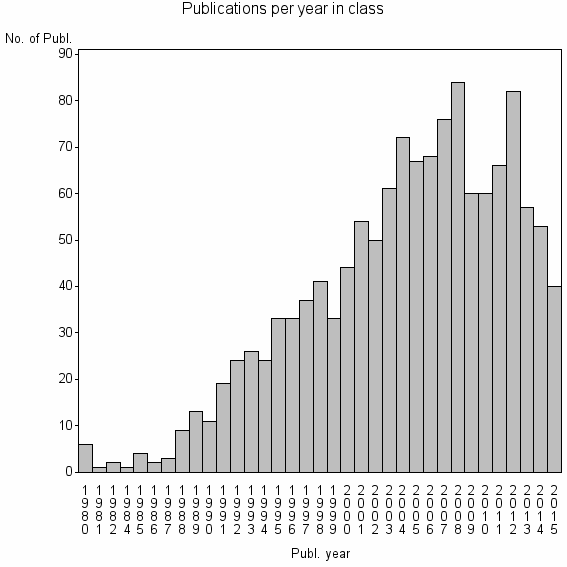
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 7216 | 1316 | 47.6 | 82% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 395 | 16247 | CYTOCHROME P450//CYP2D6//PREGNANE X RECEPTOR |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | SITE OF METABOLISM | Author keyword | 13 | 80% | 1% | 8 |
| 2 | ACTIVE SITE TOPOLOGY | Author keyword | 8 | 75% | 0% | 6 |
| 3 | COMPUTAT CHEM MOL STRUCT | Address | 6 | 80% | 0% | 4 |
| 4 | ATYPICAL KINETICS | Author keyword | 6 | 58% | 1% | 7 |
| 5 | MOLEC EXPTL MED BIOCHEM NX4 | Address | 6 | 100% | 0% | 4 |
| 6 | TYPE II BINDING | Author keyword | 5 | 63% | 0% | 5 |
| 7 | 1 PYRENEBUTANOL | Author keyword | 4 | 67% | 0% | 4 |
| 8 | CYP2B4 | Author keyword | 4 | 50% | 0% | 6 |
| 9 | DRUG METAB PHARMACOKINET BIOANALYT CHEM | Address | 4 | 50% | 0% | 6 |
| 10 | HUMAN DRUG METABOLISM | Author keyword | 4 | 56% | 0% | 5 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | SITE OF METABOLISM | 13 | 80% | 1% | 8 | Search SITE+OF+METABOLISM | Search SITE+OF+METABOLISM |
| 2 | ACTIVE SITE TOPOLOGY | 8 | 75% | 0% | 6 | Search ACTIVE+SITE+TOPOLOGY | Search ACTIVE+SITE+TOPOLOGY |
| 3 | ATYPICAL KINETICS | 6 | 58% | 1% | 7 | Search ATYPICAL+KINETICS | Search ATYPICAL+KINETICS |
| 4 | TYPE II BINDING | 5 | 63% | 0% | 5 | Search TYPE+II+BINDING | Search TYPE+II+BINDING |
| 5 | 1 PYRENEBUTANOL | 4 | 67% | 0% | 4 | Search 1+PYRENEBUTANOL | Search 1+PYRENEBUTANOL |
| 6 | CYP2B4 | 4 | 50% | 0% | 6 | Search CYP2B4 | Search CYP2B4 |
| 7 | HUMAN DRUG METABOLISM | 4 | 56% | 0% | 5 | Search HUMAN+DRUG+METABOLISM | Search HUMAN+DRUG+METABOLISM |
| 8 | BURIED ACTIVE SITE | 3 | 100% | 0% | 3 | Search BURIED+ACTIVE+SITE | Search BURIED+ACTIVE+SITE |
| 9 | IN SILICO ADMET | 3 | 45% | 0% | 5 | Search IN+SILICO+ADMET | Search IN+SILICO+ADMET |
| 10 | 2C9 | 3 | 60% | 0% | 3 | Search 2C9 | Search 2C9 |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | COMBINED PROTEIN | 67 | 87% | 3% | 33 |
| 2 | 3A4 | 47 | 32% | 9% | 122 |
| 3 | SUBSTRATE RECOGNITION SITES | 35 | 59% | 3% | 39 |
| 4 | POSSIBLE PATHWAYS | 33 | 100% | 1% | 13 |
| 5 | SMARTCYP | 31 | 92% | 1% | 12 |
| 6 | COMPETITIVE CYP2C9 INHIBITORS | 28 | 81% | 1% | 17 |
| 7 | 3A4 INHIBITORS | 27 | 92% | 1% | 11 |
| 8 | 2B1 | 27 | 62% | 2% | 28 |
| 9 | HUMAN CYTOCHROME P4502C9 | 27 | 53% | 3% | 36 |
| 10 | RS PREDICTOR | 26 | 100% | 1% | 11 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Structure-based ligand design to overcome CYP inhibition in drug discovery projects | 2014 | 4 | 43 | 86% |
| The ins and outs of cytochrome P450s | 2007 | 127 | 31 | 52% |
| State-of-the-art tools for computational site of metabolism predictions: Comparative analysis, mechanistical insights, and future applications | 2007 | 70 | 37 | 81% |
| Structure-based Drug Metabolism Predictions for Drug Design | 2010 | 63 | 88 | 48% |
| Structural Diversity of Eukaryotic Membrane Cytochrome P450s | 2013 | 18 | 96 | 54% |
| Cytochrome P450 and chemical toxicology | 2008 | 435 | 143 | 14% |
| In silico site of metabolism prediction of cytochrome P450-mediated biotransformations | 2011 | 24 | 66 | 65% |
| Conformational diversity and ligand tunnels of mammalian cytochrome P450s | 2013 | 9 | 77 | 71% |
| Dynamics and Hydration of the Active Sites of Mammalian Cytochromes P450 Probed by Molecular Dynamics Simulations | 2012 | 16 | 85 | 65% |
| Cytochrome P450 in silico: An integrative modeling approach | 2005 | 136 | 191 | 52% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | COMPUTAT CHEM MOL STRUCT | 6 | 80% | 0.3% | 4 |
| 2 | MOLEC EXPTL MED BIOCHEM NX4 | 6 | 100% | 0.3% | 4 |
| 3 | DRUG METAB PHARMACOKINET BIOANALYT CHEM | 4 | 50% | 0.5% | 6 |
| 4 | CHEM PHARMACOCHEM | 3 | 40% | 0.5% | 6 |
| 5 | LHASA LTD | 3 | 42% | 0.4% | 5 |
| 6 | MEM 255 | 2 | 67% | 0.2% | 2 |
| 7 | CHEM PROTEOM IL | 2 | 50% | 0.2% | 3 |
| 8 | DISCOVERY DMPK BIOANALYT CHEM | 2 | 28% | 0.4% | 5 |
| 9 | MOL INFORMAT STRUCT DESIGN | 1 | 38% | 0.2% | 3 |
| 10 | DDS DMPK | 1 | 100% | 0.2% | 2 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000263137 | PUTIDAREDOXIN//CYTOCHROME P450CAM//CYP106A2 |
| 2 | 0.0000225062 | ANTLEY BIXLER SYNDROME//CYTOCHROME P450 OXIDOREDUCTASE//NADPH CYTOCHROME P450 REDUCTASE |
| 3 | 0.0000209519 | CYP3A//COCKTAIL//MECHANISM BASED INHIBITION |
| 4 | 0.0000146093 | COMPOUND I//LISE MEITNER MINERVA COMPUTAT QUANTUM CHEM//CPD I |
| 5 | 0.0000138519 | N ALKYLPROTOPORPHYRIN IX//1 AMINOBENZOTRIAZOLE//CHICK EMBRYO LIVER |
| 6 | 0.0000137100 | PHARMACOKINET BIOANAL//CYP2D19//PROPRANOLOL ENANTIOMER |
| 7 | 0.0000101594 | CYP2A6//CYP2A5//CYP2A13 |
| 8 | 0.0000087491 | CYP1B1//CYP2S1//CYTOCHROME P4501B1 |
| 9 | 0.0000081405 | REACTIVE METABOLITE//MASS DEFECT FILTER//TIENILIC ACID |
| 10 | 0.0000079039 | CYP2C12//ONE DAY TREATMENT//S BIOCHEM |