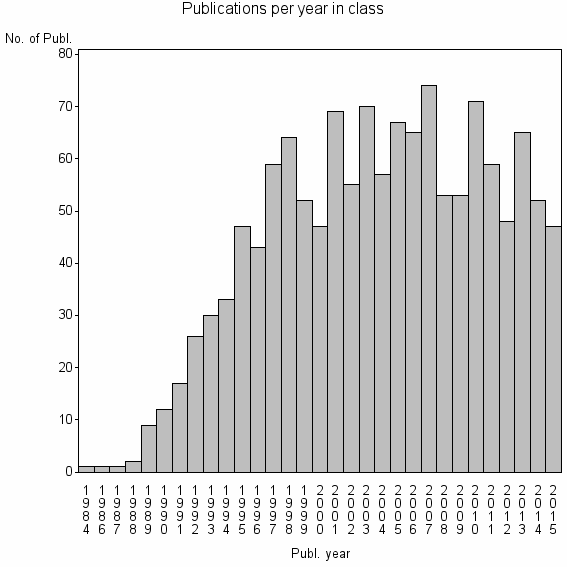
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 6945 | 1349 | 47.5 | 93% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | C EBP BETA | Author keyword | 28 | 30% | 6% | 80 |
| 2 | C EBP DELTA | Author keyword | 25 | 60% | 2% | 28 |
| 3 | C EBP | Author keyword | 20 | 25% | 5% | 69 |
| 4 | C EBP ALPHA | Author keyword | 20 | 25% | 5% | 68 |
| 5 | MOL BIOL CANC GENET | Address | 20 | 100% | 1% | 9 |
| 6 | CCAAT ENHANCER BINDING PROTEINS | Author keyword | 17 | 62% | 1% | 18 |
| 7 | CEBPD | Author keyword | 15 | 73% | 1% | 11 |
| 8 | CCAAT ENHANCER BINDING PROTEIN | Author keyword | 14 | 31% | 3% | 37 |
| 9 | CELL SIGNALING CANC GRP | Address | 11 | 65% | 1% | 11 |
| 10 | EUKARYOT TRANSCRIPT REGULAT SECT | Address | 11 | 67% | 1% | 10 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | C EBP BETA | 28 | 30% | 6% | 80 | Search C+EBP+BETA | Search C+EBP+BETA |
| 2 | C EBP DELTA | 25 | 60% | 2% | 28 | Search C+EBP+DELTA | Search C+EBP+DELTA |
| 3 | C EBP | 20 | 25% | 5% | 69 | Search C+EBP | Search C+EBP |
| 4 | C EBP ALPHA | 20 | 25% | 5% | 68 | Search C+EBP+ALPHA | Search C+EBP+ALPHA |
| 5 | CCAAT ENHANCER BINDING PROTEINS | 17 | 62% | 1% | 18 | Search CCAAT+ENHANCER+BINDING+PROTEINS | Search CCAAT+ENHANCER+BINDING+PROTEINS |
| 6 | CEBPD | 15 | 73% | 1% | 11 | Search CEBPD | Search CEBPD |
| 7 | CCAAT ENHANCER BINDING PROTEIN | 14 | 31% | 3% | 37 | Search CCAAT+ENHANCER+BINDING+PROTEIN | Search CCAAT+ENHANCER+BINDING+PROTEIN |
| 8 | CCAAT ENHANCER BINDING PROTEIN DELTA | 9 | 83% | 0% | 5 | Search CCAAT+ENHANCER+BINDING+PROTEIN+DELTA | Search CCAAT+ENHANCER+BINDING+PROTEIN+DELTA |
| 9 | CEBPA | 8 | 38% | 1% | 18 | Search CEBPA | Search CEBPA |
| 10 | CCAAT ENHANCER BINDING PROTEIN BETA | 6 | 33% | 1% | 16 | Search CCAAT+ENHANCER+BINDING+PROTEIN+BETA | Search CCAAT+ENHANCER+BINDING+PROTEIN+BETA |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | BINDING PROTEIN ALPHA | 48 | 28% | 11% | 149 |
| 2 | SERUM ALBUMIN GENE | 38 | 66% | 3% | 35 |
| 3 | C EBP BETA | 34 | 19% | 12% | 163 |
| 4 | NF IL6 | 29 | 43% | 4% | 52 |
| 5 | ENHANCER BINDING PROTEIN | 24 | 15% | 11% | 148 |
| 6 | 3 C EBP ISOFORMS | 24 | 53% | 2% | 31 |
| 7 | BINDING PROTEIN BETA | 23 | 23% | 7% | 89 |
| 8 | TRANSCRIPTIONAL ACTIVATOR PROTEIN | 21 | 38% | 3% | 44 |
| 9 | C EBP ALPHA | 21 | 13% | 11% | 142 |
| 10 | GRANULOCYTIC DIFFERENTIATION | 16 | 18% | 6% | 78 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| CCAAT/enhancer-binding proteins: structure, function and regulation | 2002 | 736 | 195 | 75% |
| The CCAAT/enhancer (C/EBP) family of basic-leucine zipper (bZIP) transcription factors is a multifaceted highly-regulated system for gene regulation | 2011 | 77 | 133 | 57% |
| Biological role of the CCAAT enhancer-binding protein family of transcription factors | 1998 | 561 | 85 | 79% |
| The C/EBP family of transcription factors: a paradigm for interaction between gene expression and proliferation control | 2007 | 144 | 61 | 67% |
| Regulation of C/EBP beta and resulting functions in cells of the monocytic lineage | 2012 | 28 | 158 | 59% |
| C/EBP alpha mutations in acute myeloid leukaemias | 2004 | 144 | 52 | 83% |
| The role of C/EBP isoforms in the control of inflammatory and native immunity functions | 1998 | 433 | 66 | 70% |
| Dysregulation of the C/EBP alpha Differentiation Pathway in Human Cancer | 2009 | 82 | 105 | 64% |
| Upstream open reading frames: Molecular switches in (patho)physiology | 2010 | 54 | 86 | 51% |
| Translational control of C/EBP alpha and C/EBP beta isoform expression | 2000 | 309 | 99 | 45% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | MOL BIOL CANC GENET | 20 | 100% | 0.7% | 9 |
| 2 | CELL SIGNALING CANC GRP | 11 | 65% | 0.8% | 11 |
| 3 | EUKARYOT TRANSCRIPT REGULAT SECT | 11 | 67% | 0.7% | 10 |
| 4 | STATE CELL GENE THER Y | 7 | 67% | 0.4% | 6 |
| 5 | EUKARYOT TRANSCRIPT REGULAT GRP | 6 | 71% | 0.4% | 5 |
| 6 | OHIO STATE COMPREHENS CANC | 5 | 47% | 0.6% | 8 |
| 7 | PATHOL MOL HUMAN GENET | 4 | 75% | 0.2% | 3 |
| 8 | SECT HEMATOL WWW428A | 4 | 75% | 0.2% | 3 |
| 9 | MOL MECH DEV GRP | 3 | 100% | 0.2% | 3 |
| 10 | GRP RECH BIOL DEV | 2 | 44% | 0.3% | 4 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000161417 | PU1//SPI 1 PU1//SFPI1 PU1 |
| 2 | 0.0000134715 | SALICYLIDENEAMINO 2 THIOPHENOL//GAMABUFOTALIN//INFLAMMATION CONTROL |
| 3 | 0.0000104474 | ADIPOGENESIS//PREADIPOCYTE//ADIPOCYTE DIFFERENTIATION |
| 4 | 0.0000083255 | BIOL MOLEC RUE CHEVAUX 67//HEPATOCYTE NUCLEAR FACTOR I//HEREDITARY PERSISTENCE OF ALPHA FETOPROTEIN |
| 5 | 0.0000079520 | TRIBBLES//TRB3//TRIB3 |
| 6 | 0.0000062361 | FLT3//FLT3 ITD//NPM1 |
| 7 | 0.0000060808 | PHOSPHOENOLPYRUVATE CARBOXYKINASE//GLYCERONEOGENESIS//PHOSPHOENOLPYRUVATE CARBOXYKINASE GENE |
| 8 | 0.0000050829 | EPIMORPHIN//MORPHOREGULAT//VERTEBROGENESIS |
| 9 | 0.0000046551 | DRS GENE//HEPATIC LEUKEMIA FACTOR//E2A HLF |
| 10 | 0.0000044493 | GFI1B//GFI1//ZELLBIOL TUMORFOR |