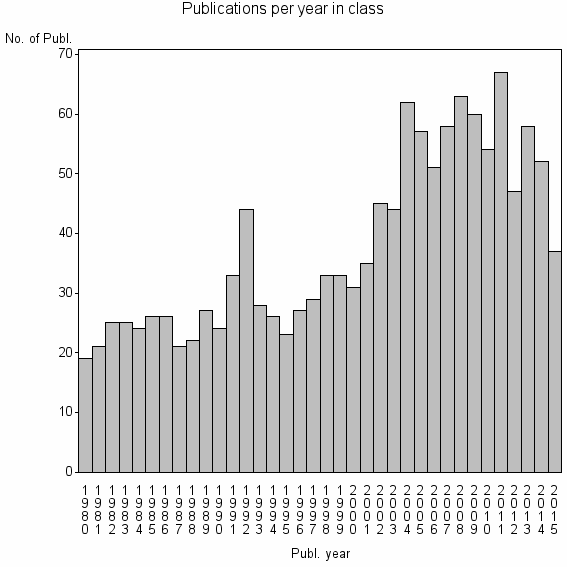
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 6872 | 1357 | 42.8 | 77% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 2019 | 4960 | PHOSPHOTRANSFERASE SYSTEM//LACTOSE PERMEASE//BIOL BAKTERIENPHYSIOL |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | BIOL BAKTERIENPHYSIOL | Address | 41 | 81% | 2% | 25 |
| 2 | PERIPLASMIC BINDING PROTEIN | Author keyword | 26 | 51% | 3% | 37 |
| 3 | GLUTAMINE BINDING PROTEIN | Author keyword | 24 | 91% | 1% | 10 |
| 4 | AG BAKTERIENPHYSIOL | Address | 17 | 100% | 1% | 8 |
| 5 | PERIPLASMIC BINDING PROTEINS | Author keyword | 16 | 60% | 1% | 18 |
| 6 | MALTOSE TRANSPORT | Author keyword | 15 | 68% | 1% | 13 |
| 7 | LMRA | Author keyword | 10 | 57% | 1% | 12 |
| 8 | MSBA | Author keyword | 10 | 58% | 1% | 11 |
| 9 | GGBP | Author keyword | 9 | 83% | 0% | 5 |
| 10 | GLUCOSE GALACTOSE BINDING PROTEIN | Author keyword | 9 | 83% | 0% | 5 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | PERIPLASMIC BINDING PROTEIN | 26 | 51% | 3% | 37 | Search PERIPLASMIC+BINDING+PROTEIN | Search PERIPLASMIC+BINDING+PROTEIN |
| 2 | GLUTAMINE BINDING PROTEIN | 24 | 91% | 1% | 10 | Search GLUTAMINE+BINDING+PROTEIN | Search GLUTAMINE+BINDING+PROTEIN |
| 3 | PERIPLASMIC BINDING PROTEINS | 16 | 60% | 1% | 18 | Search PERIPLASMIC+BINDING+PROTEINS | Search PERIPLASMIC+BINDING+PROTEINS |
| 4 | MALTOSE TRANSPORT | 15 | 68% | 1% | 13 | Search MALTOSE+TRANSPORT | Search MALTOSE+TRANSPORT |
| 5 | LMRA | 10 | 57% | 1% | 12 | Search LMRA | Search LMRA |
| 6 | MSBA | 10 | 58% | 1% | 11 | Search MSBA | Search MSBA |
| 7 | GGBP | 9 | 83% | 0% | 5 | Search GGBP | Search GGBP |
| 8 | GLUCOSE GALACTOSE BINDING PROTEIN | 9 | 83% | 0% | 5 | Search GLUCOSE+GALACTOSE+BINDING+PROTEIN | Search GLUCOSE+GALACTOSE+BINDING+PROTEIN |
| 9 | SOLUTE BINDING PROTEIN | 5 | 50% | 1% | 7 | Search SOLUTE+BINDING+PROTEIN | Search SOLUTE+BINDING+PROTEIN |
| 10 | HISTIDINE PERMEASE | 4 | 67% | 0% | 4 | Search HISTIDINE+PERMEASE | Search HISTIDINE+PERMEASE |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | HISTIDINE PERMEASE | 59 | 67% | 4% | 53 |
| 2 | MALTOSE TRANSPORTER | 57 | 62% | 4% | 58 |
| 3 | MALTOSE TRANSPORT | 47 | 47% | 5% | 74 |
| 4 | MEMBRANE COMPONENTS | 32 | 50% | 3% | 46 |
| 5 | PERIPLASMIC HISTIDINE PERMEASE | 27 | 92% | 1% | 11 |
| 6 | ACTIVE TRANSPORT | 24 | 14% | 12% | 157 |
| 7 | BTUCD F | 24 | 91% | 1% | 10 |
| 8 | MALK SUBUNIT | 22 | 81% | 1% | 13 |
| 9 | ESCHERICHIA COLI MSBA | 20 | 58% | 2% | 23 |
| 10 | MALTOSE BINDING | 20 | 38% | 3% | 41 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| ABC transporters: the power to change | 2009 | 304 | 66 | 68% |
| Structure, function, and evolution of bacterial ATP-binding cassette systems | 2008 | 386 | 520 | 39% |
| Structural diversity of ABC transporters | 2014 | 11 | 93 | 83% |
| A structural classification of substrate-binding proteins | 2010 | 103 | 49 | 63% |
| Structure and mechanism of ABC transporter proteins | 2007 | 241 | 31 | 65% |
| Structure and mechanism of ATP-binding cassette transporters | 2009 | 138 | 31 | 55% |
| ATP-binding cassette transporters in bacteria | 2004 | 329 | 121 | 49% |
| ABC TRANSPORTERS - FROM MICROORGANISMS TO MAN | 1992 | 2732 | 215 | 30% |
| Atomic structure and specificity of bacterial periplasmic receptors for active transport and chemotaxis: Variation of common themes | 1996 | 335 | 34 | 74% |
| Perspectives on the structure-function of ABC transporters: The Switch and Constant Contact Models | 2012 | 29 | 131 | 70% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | BIOL BAKTERIENPHYSIOL | 41 | 81% | 1.8% | 25 |
| 2 | AG BAKTERIENPHYSIOL | 17 | 100% | 0.6% | 8 |
| 3 | BIOL PHYSIOL MIKROORGANISMEN | 6 | 100% | 0.3% | 4 |
| 4 | MOL SENSING | 4 | 27% | 0.9% | 12 |
| 5 | BIO N210 | 3 | 40% | 0.4% | 6 |
| 6 | URA 1444 | 2 | 28% | 0.4% | 5 |
| 7 | FAIRCHILD 702A | 1 | 31% | 0.3% | 4 |
| 8 | DRDC BIOPHYS MOL CELLULAIRE | 1 | 100% | 0.1% | 2 |
| 9 | LEHRSTUHL TECH MIKROBIOL WEIHENSTEPHAN | 1 | 100% | 0.1% | 2 |
| 10 | UNITE PROGRAMMAT MOL TOXICOL GENET | 1 | 18% | 0.5% | 7 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000086969 | ANALYT BIOSENSORS GRP//INAABG//GEAS |
| 2 | 0.0000075516 | DOMAIN INSERTION//PROTEIN SWITCH//BIFUNCTIONAL FUSION ENZYME |
| 3 | 0.0000073276 | LACTOSE PERMEASE//PHYSIOL MICROBIOL//MELIBIOSE CARRIER |
| 4 | 0.0000071730 | P GLYCOPROTEIN//MULTIDRUG RESISTANCE//PSC 833 |
| 5 | 0.0000068740 | RIBOSOME MODULATION FACTOR//AMINE PHARMA//POLYAMINE MODULON |
| 6 | 0.0000064289 | BETA ROLL//ESCHERICHIA COLI HEMOLYSIN//C TERMINAL SECRETION SIGNAL |
| 7 | 0.0000058728 | MULTIXENOBIOTIC RESISTANCE//MOL ECOTOXICOL//MXR |
| 8 | 0.0000056761 | CYCLIC AMP RECEPTOR PROTEIN//CATABOLITE ACTIVATOR PROTEIN CAP//CAMP RECEPTOR PROTEIN |
| 9 | 0.0000056465 | CFTR//CYSTIC FIBROSIS TRANSMEMBRANE CONDUCTANCE REGULATOR//GREGORY FLEMING JAMES CYST FIBROSIS |
| 10 | 0.0000055952 | PHO REGULON//C P LYASE//CARBON PHOSPHORUS BOND |