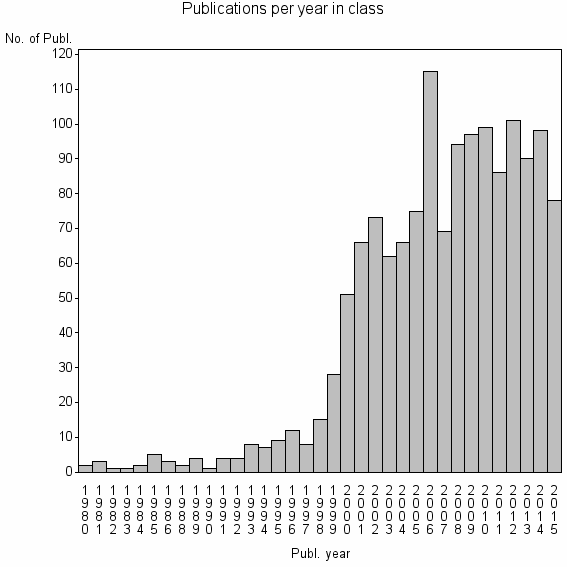
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 6286 | 1439 | 52.1 | 91% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 302 | 17871 | TRANSLESION SYNTHESIS//DNA POLYMERASE//RECA PROTEIN |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | TRANSLESION SYNTHESIS | Author keyword | 128 | 61% | 9% | 136 |
| 2 | DNA POLYMERASE ETA | Author keyword | 58 | 73% | 3% | 44 |
| 3 | DNA DAMAGE TOLERANCE | Author keyword | 57 | 79% | 3% | 37 |
| 4 | REV1 | Author keyword | 49 | 78% | 2% | 32 |
| 5 | TRANSLESION DNA SYNTHESIS | Author keyword | 48 | 56% | 4% | 58 |
| 6 | REV3 | Author keyword | 36 | 79% | 2% | 23 |
| 7 | POLYMERASE ETA | Author keyword | 35 | 89% | 1% | 16 |
| 8 | Y FAMILY POLYMERASE | Author keyword | 34 | 93% | 1% | 13 |
| 9 | POL KAPPA | Author keyword | 33 | 100% | 1% | 13 |
| 10 | DNA POLYMERASE IOTA | Author keyword | 31 | 82% | 1% | 18 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | TRANSLESION SYNTHESIS | 128 | 61% | 9% | 136 | Search TRANSLESION+SYNTHESIS | Search TRANSLESION+SYNTHESIS |
| 2 | DNA POLYMERASE ETA | 58 | 73% | 3% | 44 | Search DNA+POLYMERASE+ETA | Search DNA+POLYMERASE+ETA |
| 3 | DNA DAMAGE TOLERANCE | 57 | 79% | 3% | 37 | Search DNA+DAMAGE+TOLERANCE | Search DNA+DAMAGE+TOLERANCE |
| 4 | REV1 | 49 | 78% | 2% | 32 | Search REV1 | Search REV1 |
| 5 | TRANSLESION DNA SYNTHESIS | 48 | 56% | 4% | 58 | Search TRANSLESION+DNA+SYNTHESIS | Search TRANSLESION+DNA+SYNTHESIS |
| 6 | REV3 | 36 | 79% | 2% | 23 | Search REV3 | Search REV3 |
| 7 | POLYMERASE ETA | 35 | 89% | 1% | 16 | Search POLYMERASE+ETA | Search POLYMERASE+ETA |
| 8 | Y FAMILY POLYMERASE | 34 | 93% | 1% | 13 | Search Y+FAMILY+POLYMERASE | Search Y+FAMILY+POLYMERASE |
| 9 | POL KAPPA | 33 | 100% | 1% | 13 | Search POL+KAPPA | Search POL+KAPPA |
| 10 | DNA POLYMERASE IOTA | 31 | 82% | 1% | 18 | Search DNA+POLYMERASE+IOTA | Search DNA+POLYMERASE+IOTA |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | THYMINE THYMINE DIMER | 293 | 81% | 12% | 176 |
| 2 | TRANSLESION SYNTHESIS | 287 | 52% | 27% | 388 |
| 3 | ESCHERICHIA COLI DINB | 212 | 90% | 7% | 94 |
| 4 | POL ETA | 159 | 80% | 7% | 98 |
| 5 | REV1 PROTEIN | 155 | 85% | 6% | 81 |
| 6 | LESION BYPASS | 128 | 60% | 10% | 139 |
| 7 | PIGMENTOSUM VARIANT CELLS | 125 | 83% | 5% | 70 |
| 8 | POSTREPLICATION REPAIR | 113 | 52% | 11% | 154 |
| 9 | ERROR PRONE | 112 | 51% | 11% | 158 |
| 10 | DAMAGE INDUCED MUTAGENESIS | 107 | 82% | 4% | 63 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Y-family DNA polymerases and their role in tolerance of cellular DNA damage | 2012 | 143 | 119 | 77% |
| DNA polymerases and cancer | 2011 | 139 | 191 | 64% |
| Eukaryotic Translesion Polymerases and Their Roles and Regulation in DNA Damage Tolerance | 2009 | 197 | 269 | 79% |
| Eukaryotic translesion synthesis DNA polymerases: Specificity of structure and function | 2005 | 534 | 150 | 62% |
| What a difference a decade makes: Insights into translesion DNA synthesis | 2007 | 161 | 144 | 75% |
| Regulation of PCNA-protein interactions for genome stability | 2013 | 46 | 142 | 54% |
| Y-family DNA polymerases in mammalian cells | 2009 | 65 | 183 | 86% |
| DNA polymerase zeta (pol zeta) in higher eukaryotes | 2008 | 79 | 89 | 81% |
| Timing and spacing of ubiquitin-dependent DNA damage bypass | 2011 | 30 | 66 | 74% |
| The fidelity of DNA synthesis by eukaryotic replicative and translesion synthesis polymerases | 2008 | 176 | 124 | 39% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | GENOM INTEGR | 17 | 51% | 1.7% | 24 |
| 2 | SECT DNA REPLICAT REPAIR MUTAGENESIS | 13 | 38% | 1.9% | 28 |
| 3 | SEALY MOL SCI | 12 | 18% | 4.4% | 63 |
| 4 | GENOME ABIL SECT | 4 | 28% | 0.8% | 11 |
| 5 | GENET INFORMAT ANAL | 3 | 100% | 0.2% | 3 |
| 6 | CANC GENOM IN IDUALIZED MED | 3 | 60% | 0.2% | 3 |
| 7 | UPR 3081 | 3 | 60% | 0.2% | 3 |
| 8 | CNRS FRE3211 | 2 | 67% | 0.1% | 2 |
| 9 | GENOME DYNAM GRP | 2 | 67% | 0.1% | 2 |
| 10 | OHIO STATE BIOPHYS PROGRAM | 2 | 21% | 0.6% | 9 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000242385 | DNA POLYMERASE//DNA POLYMERASE BETA//DNA POLYMERASE LAMBDA |
| 2 | 0.0000116339 | CLAMP LOADER//REPLICATION FACTOR C//DNA REPLICAT |
| 3 | 0.0000108044 | SOS MUTAGENESIS//UMUDC//LEXA REPRESSOR |
| 4 | 0.0000084067 | DNA POLYMERASE ALPHA//DNA SYNTHESOME//DNA POLYMERASE DELTA |
| 5 | 0.0000082051 | RAD51//HOMOLOGOUS RECOMBINATION//RAD52 |
| 6 | 0.0000073859 | SOMATIC HYPERMUTATION//CLASS SWITCH RECOMBINATION//ACTIVATION INDUCED CYTIDINE DEAMINASE |
| 7 | 0.0000071441 | COOPERAT LIFE SCI//FRONTIER GENOM DRUG DISCOVERY//DEHYDROALTENUSIN |
| 8 | 0.0000068442 | FANCONI ANEMIA//FANCONI ANAEMIA//FANCD2 |
| 9 | 0.0000068075 | DSK2//UBIQUILIN 1//LUBAC |
| 10 | 0.0000064240 | CHK1//RAD9//CLASPIN |