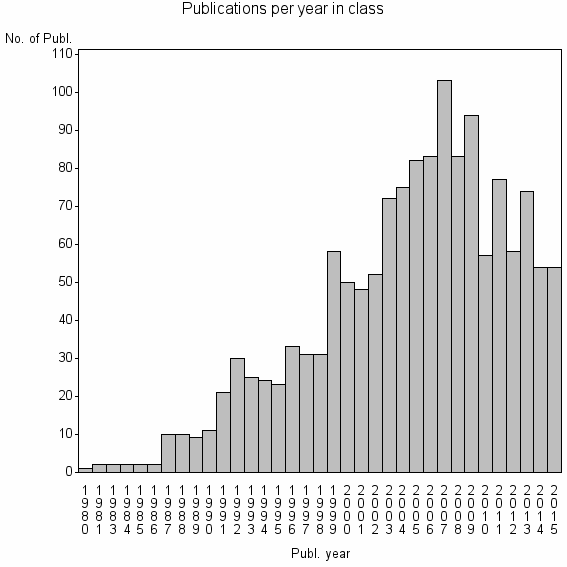
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 6257 | 1443 | 37.4 | 76% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 42 | 31225 | PROTEINS-STRUCTURE FUNCTION AND BIOINFORMATICS//PROTEIN STRUCTURE PREDICTION//PROTEIN FOLDING |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | BAKER CHEM CHEM BIOL | Address | 25 | 27% | 5% | 77 |
| 2 | REPLICA EXCHANGE METHOD | Author keyword | 20 | 58% | 2% | 23 |
| 3 | GENERALIZED ENSEMBLE ALGORITHM | Author keyword | 19 | 80% | 1% | 12 |
| 4 | MULTICANONICAL | Author keyword | 19 | 76% | 1% | 13 |
| 5 | MULTICANONICAL SIMULATION | Author keyword | 17 | 79% | 1% | 11 |
| 6 | UNRES FORCE FIELD | Author keyword | 15 | 88% | 0% | 7 |
| 7 | GENERALIZED ENSEMBLE SIMULATIONS | Author keyword | 13 | 80% | 1% | 8 |
| 8 | MULTICANONICAL ALGORITHM | Author keyword | 11 | 65% | 1% | 11 |
| 9 | MULTICANONICAL MOLECULAR DYNAMICS | Author keyword | 11 | 67% | 1% | 10 |
| 10 | WANG LANDAU SAMPLING | Author keyword | 11 | 54% | 1% | 14 |
Web of Science journal categories |
Author Key Words |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | MULTICANONICAL ENSEMBLE | 85 | 69% | 5% | 73 |
| 2 | SPIN GLASS SIMULATIONS | 74 | 79% | 3% | 48 |
| 3 | ISOLATED C PEPTIDE | 55 | 70% | 3% | 46 |
| 4 | HIERARCHICAL DESIGN | 52 | 80% | 2% | 32 |
| 5 | MULTIPLE MINIMA PROBLEM | 48 | 40% | 7% | 96 |
| 6 | MULTICANONICAL ALGORITHMS | 46 | 78% | 2% | 31 |
| 7 | MULTICANONICAL ALGORITHM | 46 | 76% | 2% | 32 |
| 8 | MULTIBARIC MULTITHERMAL ENSEMBLE | 38 | 86% | 1% | 19 |
| 9 | HYDROGEN BOND INTERACTIONS | 29 | 26% | 7% | 97 |
| 10 | PROTEIN STRUCTURE SIMULATIONS | 25 | 65% | 2% | 24 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Parallel tempering: Theory, applications, and new perspectives | 2005 | 373 | 90 | 38% |
| Equilibrium Sampling in Biomolecular Simulations | 2011 | 59 | 126 | 39% |
| Advanced replica-exchange sampling to study the flexibility and plasticity of peptides and proteins | 2013 | 12 | 89 | 54% |
| Computational techniques for efficient conformational sampling of proteins | 2008 | 66 | 41 | 51% |
| Generalized-ensemble algorithms for molecular simulations of biopolymers | 2001 | 441 | 118 | 61% |
| Density of States-Based Molecular Simulations | 2012 | 9 | 114 | 43% |
| New Monte Carlo algorithms for protein folding | 1999 | 165 | 71 | 51% |
| Atomic simulations of protein folding, using the replica exchange algorithm | 2004 | 71 | 46 | 41% |
| Protein-folding dynamics: Overview of molecular simulation techniques | 2007 | 143 | 132 | 12% |
| Applications of the Wang-Landau Algorithm to Phase Transitions of a Single Polymer Chain | 2013 | 2 | 35 | 43% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | BAKER CHEM CHEM BIOL | 25 | 27% | 5.3% | 77 |
| 2 | ADV SIMULAT IAS 2 | 6 | 80% | 0.3% | 4 |
| 3 | THEORET SCI NTZ | 6 | 30% | 1.2% | 18 |
| 4 | SOFT MATTER SYST GRP | 5 | 60% | 0.4% | 6 |
| 5 | FESTKORPERFOR IFF 2 | 4 | 75% | 0.2% | 3 |
| 6 | FUNCT MOL SCI | 4 | 11% | 2.2% | 32 |
| 7 | JOHN VON NEUMAN COMP | 3 | 100% | 0.2% | 3 |
| 8 | PROT INFORMAT | 3 | 28% | 0.7% | 10 |
| 9 | JOHN VON NEUMANN COMP | 3 | 12% | 1.5% | 22 |
| 10 | TASK | 3 | 60% | 0.2% | 3 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000193027 | MARKOV STATE MODELS//TRANSITION PATH SAMPLING//TRANSITION PATH THEORY |
| 2 | 0.0000154812 | HP MODEL//STATISTICAL POTENTIALS//PROTEIN STRUCTURE PREDICTION |
| 3 | 0.0000138391 | BIOINFORMAT TELEMED//HYDROPHOBICITY DEFICIENCY//HELIX INTERACTIONS |
| 4 | 0.0000130059 | BETA HAIRPIN//SIDE CHAIN INTERACTIONS//TRP CAGE |
| 5 | 0.0000125384 | LOOP PREDICTION//LOOP MODELING//STEPWISE FOLDING |
| 6 | 0.0000124089 | ENVIRONM IND CHEM//ONE STEP PERTURBATION//SAMPL4 |
| 7 | 0.0000104194 | PERTURBED MATRIX METHOD//ANHARMONIC SOLID//CHEM QUANTUM PHYS GRP |
| 8 | 0.0000103349 | PROTEIN FOLDING//PHI VALUE//CONTACT ORDER |
| 9 | 0.0000084183 | NAKA WORKS//CYCLOALCANES//CONFORMATIONAL POPULATION |
| 10 | 0.0000081377 | PARTITION FUNCTION ZEROS//ANTIFERROMAGNETIC POTTS MODEL//GRP MODELIZAC SIMULAC NUMER MATEMAT IND |