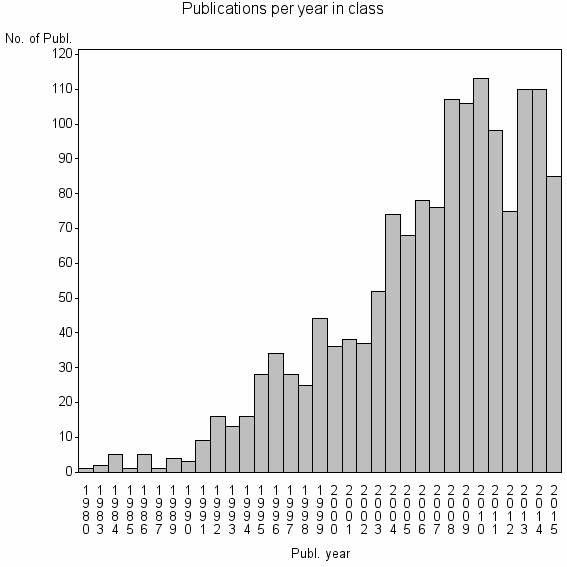
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 5843 | 1498 | 45.6 | 90% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | EG5 | Author keyword | 85 | 74% | 4% | 64 |
| 2 | KINESIN SPINDLE PROTEIN | Author keyword | 73 | 88% | 2% | 35 |
| 3 | KINESIN 5 | Author keyword | 29 | 77% | 1% | 20 |
| 4 | ISPINESIB | Author keyword | 27 | 92% | 1% | 11 |
| 5 | KSP | Author keyword | 25 | 55% | 2% | 31 |
| 6 | MITOTIC KINESIN | Author keyword | 23 | 100% | 1% | 10 |
| 7 | TACC3 | Author keyword | 19 | 61% | 1% | 20 |
| 8 | KSP INHIBITORS | Author keyword | 18 | 83% | 1% | 10 |
| 9 | XMAP215 | Author keyword | 14 | 64% | 1% | 14 |
| 10 | POLEWARD FLUX | Author keyword | 14 | 100% | 0% | 7 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | EG5 | 85 | 74% | 4% | 64 | Search EG5 | Search EG5 |
| 2 | KINESIN SPINDLE PROTEIN | 73 | 88% | 2% | 35 | Search KINESIN+SPINDLE+PROTEIN | Search KINESIN+SPINDLE+PROTEIN |
| 3 | KINESIN 5 | 29 | 77% | 1% | 20 | Search KINESIN+5 | Search KINESIN+5 |
| 4 | ISPINESIB | 27 | 92% | 1% | 11 | Search ISPINESIB | Search ISPINESIB |
| 5 | KSP | 25 | 55% | 2% | 31 | Search KSP | Search KSP |
| 6 | MITOTIC KINESIN | 23 | 100% | 1% | 10 | Search MITOTIC+KINESIN | Search MITOTIC+KINESIN |
| 7 | TACC3 | 19 | 61% | 1% | 20 | Search TACC3 | Search TACC3 |
| 8 | KSP INHIBITORS | 18 | 83% | 1% | 10 | Search KSP+INHIBITORS | Search KSP+INHIBITORS |
| 9 | XMAP215 | 14 | 64% | 1% | 14 | Search XMAP215 | Search XMAP215 |
| 10 | POLEWARD FLUX | 14 | 100% | 0% | 7 | Search POLEWARD+FLUX | Search POLEWARD+FLUX |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | EG5 | 101 | 70% | 6% | 84 |
| 2 | CENTROMERE ASSOCIATED KINESIN | 90 | 66% | 6% | 84 |
| 3 | CROSS LINKS MICROTUBULES | 85 | 85% | 3% | 45 |
| 4 | BIPOLAR KINESIN | 72 | 83% | 3% | 40 |
| 5 | MITOTIC SPINDLE | 71 | 16% | 28% | 414 |
| 6 | ISPINESIB | 61 | 95% | 1% | 20 |
| 7 | SPN ANTIGEN | 57 | 95% | 1% | 19 |
| 8 | MONASTROL | 52 | 62% | 4% | 53 |
| 9 | MITOTIC KINESIN EG5 | 41 | 52% | 4% | 56 |
| 10 | HOMOTETRAMERIC KINESIN 5 | 33 | 100% | 1% | 13 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Prime movers: the mechanochemistry of mitotic kinesins | 2014 | 15 | 168 | 53% |
| Mitosis, microtubule dynamics and the evolution of kinesins | 2015 | 2 | 93 | 56% |
| Advances in the discovery of kinesin spindle protein (Eg5) inhibitors as antitumor agents | 2013 | 21 | 153 | 75% |
| Kinesin motor proteins as targets for cancer therapy | 2009 | 83 | 72 | 71% |
| Kinesins and cancer | 2012 | 70 | 194 | 47% |
| Mitotic functions of kinesin-5 | 2010 | 45 | 64 | 84% |
| Mechanisms of Centrosome Separation and Bipolar Spindle Assembly | 2010 | 69 | 101 | 60% |
| Control of Mitotic Spindle Length | 2010 | 66 | 197 | 56% |
| Kinesin-13s in mitosis: Key players in the spatial and temporal organization of spindle microtubules | 2010 | 47 | 88 | 72% |
| Mechanisms of mitotic spindle assembly and function | 2008 | 172 | 239 | 33% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | CHEM CELL BIOL | 9 | 22% | 2.5% | 37 |
| 2 | CELL COMPUTAT BIOL | 7 | 53% | 0.6% | 9 |
| 3 | MOL MOTORS | 5 | 63% | 0.3% | 5 |
| 4 | RIPPEL ELECT MICROSCOPE IL | 3 | 100% | 0.2% | 3 |
| 5 | CELL BIOL BIOPHYS PROGRAM | 3 | 24% | 0.8% | 12 |
| 6 | CELL ARCHITECTURE | 3 | 38% | 0.4% | 6 |
| 7 | CARDIOVASC RANDALL | 3 | 60% | 0.2% | 3 |
| 8 | CHEM GENET INDEPENDENT GRP | 2 | 67% | 0.1% | 2 |
| 9 | CELL GRP | 2 | 17% | 0.7% | 10 |
| 10 | CHROMOSOME ABIL DYNAM | 2 | 26% | 0.4% | 6 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000246409 | ULTRAVIOLET MICROBEAM//CRANE FLY SPERMATOCYTES//SPINDLE FIBRES |
| 2 | 0.0000226082 | KINETOCHORE//SPINDLE ASSEMBLY CHECKPOINT//MAD2 |
| 3 | 0.0000225767 | DYNACTIN//CLIP 170//EB1 |
| 4 | 0.0000180064 | KINESIN//NECK LINKER//MOL MOTORS GRP |
| 5 | 0.0000152591 | CENTROSOME//CENTRIOLE//CENTRIN |
| 6 | 0.0000137896 | CYTOKINESIS//CLEAVAGE FURROW//CENTRALSPINDLIN |
| 7 | 0.0000126698 | AXONS AND DENDRITES//MOTOR ASSISTED TRANSPORT//KINESIN |
| 8 | 0.0000114538 | AURORA A//AURORA KINASES//AURORA KINASE |
| 9 | 0.0000094646 | GRP BIOPHYSSTAT//PROTEIN MICROTUBULES//TUBULIN RINGS |
| 10 | 0.0000069348 | DROSOPHILA MATERNAL EFFECT MUTATION//DROSOPHILA MELANOGASTER EMBRYO//IMAGINAL PRIMORDIA |