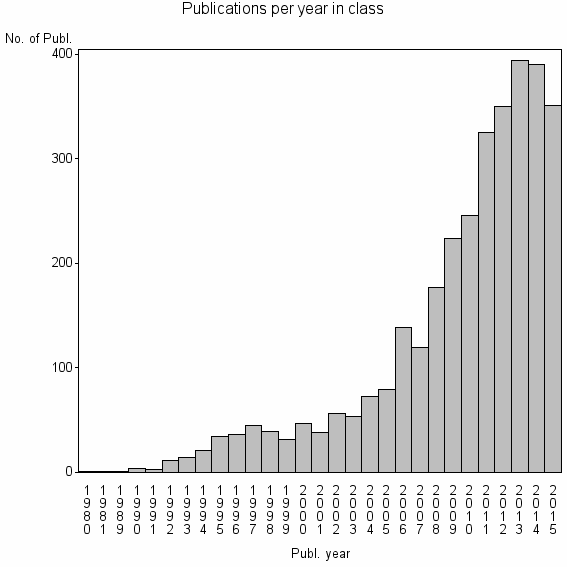
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 510 | 3298 | 42.3 | 90% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | APTAMER | Author keyword | 640 | 53% | 26% | 849 |
| 2 | SELEX | Author keyword | 338 | 63% | 10% | 336 |
| 3 | APTAMERS | Author keyword | 219 | 49% | 10% | 326 |
| 4 | APTASENSOR | Author keyword | 121 | 60% | 4% | 134 |
| 5 | CELL SELEX | Author keyword | 103 | 97% | 1% | 30 |
| 6 | ELECTROCHEMICAL APTASENSOR | Author keyword | 89 | 79% | 2% | 57 |
| 7 | DNA APTAMER | Author keyword | 82 | 68% | 2% | 72 |
| 8 | RNA APTAMER | Author keyword | 25 | 39% | 2% | 52 |
| 9 | APTASENSORS | Author keyword | 23 | 72% | 1% | 18 |
| 10 | IN VITRO SELECTION | Author keyword | 21 | 21% | 3% | 91 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | APTAMER | 640 | 53% | 26% | 849 | Search APTAMER | Search APTAMER |
| 2 | SELEX | 338 | 63% | 10% | 336 | Search SELEX | Search SELEX |
| 3 | APTAMERS | 219 | 49% | 10% | 326 | Search APTAMERS | Search APTAMERS |
| 4 | APTASENSOR | 121 | 60% | 4% | 134 | Search APTASENSOR | Search APTASENSOR |
| 5 | CELL SELEX | 103 | 97% | 1% | 30 | Search CELL+SELEX | Search CELL+SELEX |
| 6 | ELECTROCHEMICAL APTASENSOR | 89 | 79% | 2% | 57 | Search ELECTROCHEMICAL+APTASENSOR | Search ELECTROCHEMICAL+APTASENSOR |
| 7 | DNA APTAMER | 82 | 68% | 2% | 72 | Search DNA+APTAMER | Search DNA+APTAMER |
| 8 | RNA APTAMER | 25 | 39% | 2% | 52 | Search RNA+APTAMER | Search RNA+APTAMER |
| 9 | APTASENSORS | 23 | 72% | 1% | 18 | Search APTASENSORS | Search APTASENSORS |
| 10 | IN VITRO SELECTION | 21 | 21% | 3% | 91 | Search IN+VITRO+SELECTION | Search IN+VITRO+SELECTION |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | SYSTEMATIC EVOLUTION | 447 | 77% | 9% | 301 |
| 2 | DNA APTAMERS | 376 | 79% | 7% | 242 |
| 3 | EXPONENTIAL ENRICHMENT | 374 | 74% | 8% | 274 |
| 4 | RNA APTAMERS | 275 | 72% | 7% | 219 |
| 5 | DNA APTAMER | 236 | 64% | 7% | 233 |
| 6 | IN VITRO SELECTION | 170 | 29% | 15% | 498 |
| 7 | SELEX | 156 | 72% | 4% | 123 |
| 8 | APTASENSOR | 141 | 57% | 5% | 169 |
| 9 | NUCLEIC ACID APTAMERS | 129 | 62% | 4% | 135 |
| 10 | RNA LIGANDS | 88 | 71% | 2% | 72 |
Journals |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | NUCLEIC ACID THERAPEUTICS | 6 | 16% | 1% | 32 |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Aptamers as a replacement for antibodies in enzyme-linked immunosorbent assay | 2015 | 10 | 104 | 64% |
| Aptamers as therapeutics | 2010 | 404 | 152 | 66% |
| Aptamer in Bioanalytical Applications | 2011 | 201 | 156 | 78% |
| Functional Nucleic Acid Sensors | 2009 | 703 | 593 | 44% |
| Aptamers from Cell-Based Selection for Bioanalytical Applications | 2013 | 86 | 122 | 47% |
| Recent advances in aptasensors based on graphene and graphene-like nanomaterials | 2015 | 6 | 158 | 49% |
| Applications of Aptamers as Sensors | 2009 | 264 | 142 | 71% |
| Aptamers Generated from Cell-SELEX for Molecular Medicine: A Chemical Biology Approach | 2010 | 182 | 51 | 78% |
| Aptamers Facilitating Amplified Detection of Biomolecules | 2015 | 5 | 194 | 55% |
| Therapeutic RNA aptamers in clinical trials | 2013 | 47 | 63 | 70% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | BIO NANO INTER E | 19 | 28% | 1.7% | 57 |
| 2 | SHANDS CANC | 11 | 13% | 2.4% | 80 |
| 3 | MOL MED PATHOL BIOCHEM | 11 | 78% | 0.2% | 7 |
| 4 | UF GENET | 10 | 22% | 1.2% | 41 |
| 5 | RNA SCI TECHNOL | 9 | 83% | 0.2% | 5 |
| 6 | PROGRAM UNIT CHEM BIOL MED CHEM | 6 | 38% | 0.4% | 12 |
| 7 | DUKE TRANSLAT | 5 | 36% | 0.4% | 12 |
| 8 | BIOQUIM INVEST | 4 | 47% | 0.2% | 7 |
| 9 | BIOLMOL SCI BIOMED | 4 | 75% | 0.1% | 3 |
| 10 | BIONANOTECHNOL MOL ENGN HUNAN PROV | 4 | 16% | 0.7% | 23 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000129602 | ROLLING CIRCLE AMPLIFICATION//HYBRIDIZATION CHAIN REACTION//DNAZYME |
| 2 | 0.0000101163 | DNAZYME//DEOXYRIBOZYME//CATALYTIC DNA |
| 3 | 0.0000075159 | L DNA//HETEROCHIRAL DNA//UMR CNRS UMII 5625 |
| 4 | 0.0000074595 | RIBOSWITCH//RIBOSWITCHES//STREPTOMYCES DAVAWENSIS |
| 5 | 0.0000065247 | IMMUNO PCR//LEHRSTUHL BIOL CHEM MIKROSTRUKTURTECH//REAL TIME IMMUNO PCR |
| 6 | 0.0000064427 | ABASIC SITE//REGULATORY BIOORGAN CHEM//MISMATCH BINDING LIGAND |
| 7 | 0.0000057288 | INT NANOTECHNOL//CIGMH//DNA MATERIALS |
| 8 | 0.0000056273 | COLORIMETRIC DETECTION//MERCURY ION//MERCURY IONS |
| 9 | 0.0000047302 | ELECTROCHEMICAL IMMUNOSENSOR//IMMUNOSENSOR//ELECTROCHEMICAL IMMUNOASSAY |
| 10 | 0.0000043495 | ELECTROCHEMICAL DNA BIOSENSOR//DNA BIOSENSOR//GENOSENSOR |