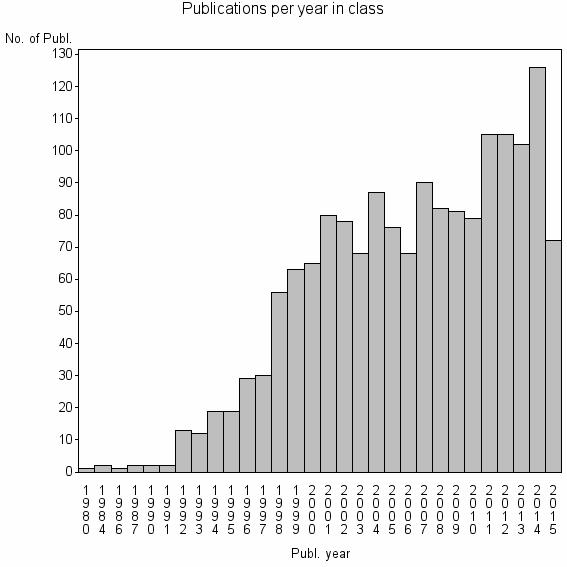
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 5046 | 1615 | 53.0 | 94% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | SWI SNF | Author keyword | 108 | 55% | 8% | 135 |
| 2 | BRG1 | Author keyword | 56 | 57% | 4% | 67 |
| 3 | ISWI | Author keyword | 56 | 74% | 3% | 42 |
| 4 | CHROMATIN REMODELING | Author keyword | 48 | 23% | 11% | 185 |
| 5 | ARID1B | Author keyword | 33 | 100% | 1% | 13 |
| 6 | SWI SNF COMPLEX | Author keyword | 28 | 62% | 2% | 29 |
| 7 | RSF 1 | Author keyword | 27 | 92% | 1% | 11 |
| 8 | BRAHMA | Author keyword | 19 | 60% | 1% | 21 |
| 9 | BRM | Author keyword | 17 | 44% | 2% | 30 |
| 10 | BAF57 | Author keyword | 15 | 88% | 0% | 7 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | SWI SNF | 108 | 55% | 8% | 135 | Search SWI+SNF | Search SWI+SNF |
| 2 | BRG1 | 56 | 57% | 4% | 67 | Search BRG1 | Search BRG1 |
| 3 | ISWI | 56 | 74% | 3% | 42 | Search ISWI | Search ISWI |
| 4 | CHROMATIN REMODELING | 48 | 23% | 11% | 185 | Search CHROMATIN+REMODELING | Search CHROMATIN+REMODELING |
| 5 | ARID1B | 33 | 100% | 1% | 13 | Search ARID1B | Search ARID1B |
| 6 | SWI SNF COMPLEX | 28 | 62% | 2% | 29 | Search SWI+SNF+COMPLEX | Search SWI+SNF+COMPLEX |
| 7 | RSF 1 | 27 | 92% | 1% | 11 | Search RSF+1 | Search RSF+1 |
| 8 | BRAHMA | 19 | 60% | 1% | 21 | Search BRAHMA | Search BRAHMA |
| 9 | BRM | 17 | 44% | 2% | 30 | Search BRM | Search BRM |
| 10 | BAF57 | 15 | 88% | 0% | 7 | Search BAF57 | Search BAF57 |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | SWI SNF COMPLEX | 159 | 45% | 16% | 265 |
| 2 | ISWI | 108 | 64% | 7% | 106 |
| 3 | BRG1 | 107 | 68% | 6% | 93 |
| 4 | YEAST SWI SNF COMPLEX | 100 | 60% | 7% | 109 |
| 5 | SWI SNF | 86 | 53% | 7% | 114 |
| 6 | ISW2 NUCLEOSOME COMPLEX | 81 | 96% | 2% | 25 |
| 7 | MAMMALIAN SWI SNF COMPLEXES | 75 | 63% | 5% | 75 |
| 8 | EXTRANUCLEOSOMAL DNA | 69 | 91% | 2% | 29 |
| 9 | CHROMATIN REMODELING COMPLEX | 65 | 24% | 14% | 231 |
| 10 | MULTISUBUNIT COMPLEX | 64 | 57% | 5% | 75 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| The Biology of Chromatin Remodeling Complexes | 2009 | 630 | 179 | 59% |
| Chromatin remodelling during development | 2010 | 324 | 97 | 55% |
| SWI/SNF nucleosome remodellers and cancer | 2011 | 263 | 140 | 56% |
| ATP-dependent chromatin remodeling: genetics, genomics and mechanisms | 2011 | 241 | 245 | 55% |
| Cooperation between complexes that regulate chromatin structure and transcription | 2002 | 985 | 86 | 62% |
| Mechanisms and Functions of ATP-Dependent Chromatin-Remodeling Enzymes | 2013 | 66 | 89 | 66% |
| The SWI/SNF complex and cancer | 2009 | 210 | 205 | 74% |
| Chromatin remodelling: the industrial revolution of DNA around histones | 2006 | 287 | 129 | 63% |
| Regulating the Chromatin Landscape: Structural and Mechanistic Perspectives | 2014 | 14 | 158 | 62% |
| ATP-dependent nucleosomere modeling | 2002 | 460 | 167 | 73% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | MOL CELL BIOL MOL BIOLAOBA KU | 11 | 100% | 0.4% | 6 |
| 2 | GENOME ABIL CHROMATIN REMODELING SECT | 2 | 67% | 0.1% | 2 |
| 3 | SINSHEIMER S 350 | 2 | 67% | 0.1% | 2 |
| 4 | UNIT EPIGENET REGULAT | 2 | 67% | 0.1% | 2 |
| 5 | HOST PARASITE INTERACTMINATO KU | 2 | 33% | 0.3% | 5 |
| 6 | SSP STEM CELL UNIT | 2 | 24% | 0.4% | 7 |
| 7 | ALTHOUSE 306 | 2 | 33% | 0.2% | 4 |
| 8 | IST TELETHON DULBECCO | 1 | 31% | 0.2% | 4 |
| 9 | ONCOL SCI MOL BIOL GENET | 1 | 50% | 0.1% | 2 |
| 10 | SSP STEM CELL UNITHIGASHI KU | 1 | 50% | 0.1% | 2 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000181678 | NUCLEOSOME POSITIONING//NUCLEOSOME//NUCLEOSOME MAPPING |
| 2 | 0.0000147541 | HISTONE CHAPERONE//ASF1//CAF 1 |
| 3 | 0.0000122984 | NUCLEAR ACTIN//NUCLEAR MYOSIN I//NUCLEAR MYOSIN |
| 4 | 0.0000122885 | SCHIMKE IMMUNO OSSEOUS DYSPLASIA//SMARCAL1//SCHIMKE IMMUNOOSSEOUS DYSPLASIA |
| 5 | 0.0000116547 | MTA1//METASTASIS ASSOCIATED GENE 1//MTA3 |
| 6 | 0.0000111357 | MYST//MOZ//HISTONE ACETYLTRANSFERASE |
| 7 | 0.0000104337 | ARID3B//JUMONJI//AT RICH INTERACTION DOMAIN |
| 8 | 0.0000103851 | REPTIN//PONTIN//R2TP |
| 9 | 0.0000097353 | RHABDOID TUMOR//ATYPICAL TERATOID RHABDOID TUMORS//EPITHELIOID SARCOMA |
| 10 | 0.0000096936 | NUCLEAR FACTOR 1//NUCLEAR FACTOR I//DEV GENOM GRP |